6EXE
 
 | |
6EXC
 
 | |
6EXA
 | |
6EXD
 
 | |
6EXB
 
 | |
6SZ9
 
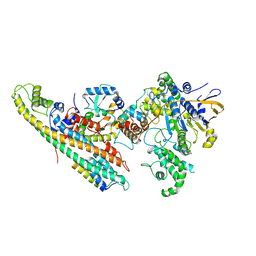 | | Type IV Coupling Complex (T4CC) from L. pneumophila. | | Descriptor: | DotY, DotZ, IcmJ (DotN), ... | | Authors: | Mace, K, Meir, A, Lukoyanova, N, Waksman, G. | | Deposit date: | 2019-10-02 | | Release date: | 2020-05-13 | | Last modified: | 2022-11-23 | | Method: | ELECTRON MICROSCOPY (3.7 Å) | | Cite: | Mechanism of effector capture and delivery by the type IV secretion system from Legionella pneumophila.
Nat Commun, 11, 2020
|
|
7OVB
 
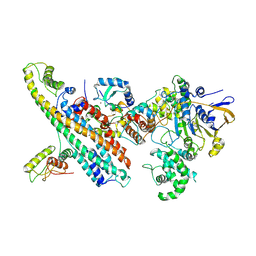 | | L. pneumophila Type IV Coupling Complex (T4CC) with density for DotY N-terminal and middle domains | | Descriptor: | DotY, DotZ, IcmJ (DotN), ... | | Authors: | Mace, K, Meir, A, Lukoyanova, N, Waksman, G. | | Deposit date: | 2021-06-14 | | Release date: | 2021-12-01 | | Last modified: | 2022-02-23 | | Method: | ELECTRON MICROSCOPY (3.61 Å) | | Cite: | Proteins DotY and DotZ modulate the dynamics and localization of the type IVB coupling complex of Legionella pneumophila.
Mol.Microbiol., 117, 2022
|
|
8FAZ
 
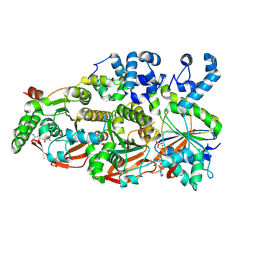 | | Cryo-EM structure of the human BCDX2 complex | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, DNA repair protein RAD51 homolog 2, DNA repair protein RAD51 homolog 3, ... | | Authors: | Jia, L, Wasmuth, E.V, Ruben, E.A, Sung, P, Rawal, Y, Greene, E.C, Meir, A, Olsen, S.K. | | Deposit date: | 2022-11-29 | | Release date: | 2023-06-21 | | Last modified: | 2023-08-02 | | Method: | ELECTRON MICROSCOPY (2.3 Å) | | Cite: | Structural insights into BCDX2 complex function in homologous recombination.
Nature, 619, 2023
|
|
8GBJ
 
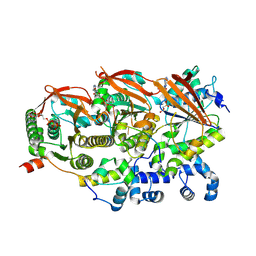 | | Cryo-EM structure of a human BCDX2/ssDNA complex | | Descriptor: | DNA (5'-D(P*CP*CP*CP*CP*CP*C)-3'), DNA repair protein RAD51 homolog 2, DNA repair protein RAD51 homolog 3, ... | | Authors: | Jia, L, Wasmuth, E.V, Ruben, E.A, Sung, P, Rawal, Y, Greene, E.C, Meir, A, Olsen, S.K. | | Deposit date: | 2023-02-26 | | Release date: | 2023-06-21 | | Last modified: | 2023-08-02 | | Method: | ELECTRON MICROSCOPY (3.11 Å) | | Cite: | Structural insights into BCDX2 complex function in homologous recombination.
Nature, 619, 2023
|
|
3EW2
 
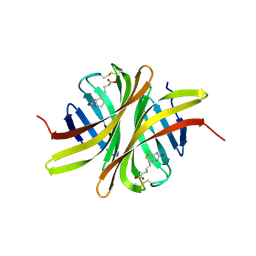 | | Crystal structure of rhizavidin-biotin complex | | Descriptor: | BIOTIN, rhizavidin | | Authors: | Livnah, O, Meir, A. | | Deposit date: | 2008-10-14 | | Release date: | 2008-12-23 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal structure of rhizavidin: insights into the enigmatic high-affinity interaction of an innate biotin-binding protein dimer.
J.Mol.Biol., 386, 2009
|
|
3EW1
 
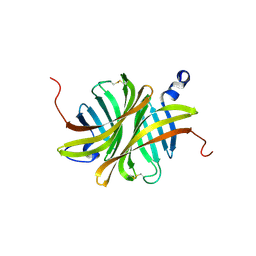 | | Crystal structure of rhizavidin | | Descriptor: | rhizavidin | | Authors: | Livnah, O, Meir, A. | | Deposit date: | 2008-10-14 | | Release date: | 2008-12-23 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Crystal structure of rhizavidin: insights into the enigmatic high-affinity interaction of an innate biotin-binding protein dimer.
J.Mol.Biol., 386, 2009
|
|
4GGZ
 | | The structure of bradavidin2-biotin complex | | Descriptor: | BIOTIN, Bradavidin 2 | | Authors: | Livnah, O, Meir, A. | | Deposit date: | 2012-08-07 | | Release date: | 2013-06-19 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | The highly dynamic oligomeric structure of bradavidin II is unique among avidin proteins.
Protein Sci., 22, 2013
|
|
4GGR
 
 | | The structure of apo bradavidin2 (Form A) | | Descriptor: | Bradavidin 2 | | Authors: | Livnah, O, Meir, A. | | Deposit date: | 2012-08-07 | | Release date: | 2013-06-19 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | The highly dynamic oligomeric structure of bradavidin II is unique among avidin proteins.
Protein Sci., 22, 2013
|
|
4GGT
 
 | | Structure of apo Bradavidin2 (Form B) | | Descriptor: | Bradavidin 2 | | Authors: | Livnah, O, Meir, A. | | Deposit date: | 2012-08-07 | | Release date: | 2013-06-19 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.693 Å) | | Cite: | The highly dynamic oligomeric structure of bradavidin II is unique among avidin proteins.
Protein Sci., 22, 2013
|
|
3SZJ
 
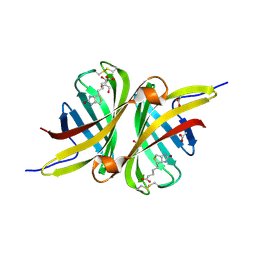 | | Structure of the shwanavidin-biotin complex | | Descriptor: | ACETIC ACID, Avidin/streptavidin, BIOTIN, ... | | Authors: | Livnah, O, Meir, A. | | Deposit date: | 2011-07-19 | | Release date: | 2012-04-11 | | Last modified: | 2012-06-13 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Structural Adaptation of a Thermostable Biotin-binding Protein in a Psychrophilic Environment.
J.Biol.Chem., 287, 2012
|
|
3T2W
 
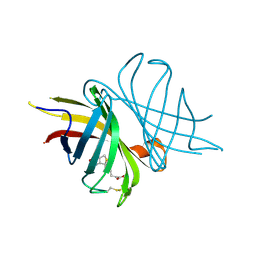 | |
3SZI
 
 | | Structure of apo shwanavidin (P21 form) | | Descriptor: | Avidin/streptavidin, FORMIC ACID | | Authors: | Livnah, O, Meir, A. | | Deposit date: | 2011-07-19 | | Release date: | 2012-04-25 | | Last modified: | 2012-06-13 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Structural Adaptation of a Thermostable Biotin-binding Protein in a Psychrophilic Environment.
J.Biol.Chem., 287, 2012
|
|
3T2X
 
 | |
3SZH
 
 | | Crystal structure of apo shwanavidin (P1 form) | | Descriptor: | Avidin/streptavidin | | Authors: | Livnah, O, Meir, A. | | Deposit date: | 2011-07-19 | | Release date: | 2012-04-11 | | Last modified: | 2012-06-13 | | Method: | X-RAY DIFFRACTION (1.07 Å) | | Cite: | Structural Adaptation of a Thermostable Biotin-binding Protein in a Psychrophilic Environment.
J.Biol.Chem., 287, 2012
|
|
