4RPT
 
 | |
6PCU
 
 | |
7K6V
 
 | |
5D9Y
 
 | | Crystal structure of TET2-5fC complex | | Descriptor: | DNA (5'-D(*AP*CP*TP*GP*TP*(5FC)P*GP*AP*AP*GP*CP*T)-3'), DNA (5'-D(*AP*GP*CP*TP*TP*CP*GP*AP*CP*AP*GP*T)-3'), FE (III) ION, ... | | Authors: | Hu, L, Cheng, J, Rao, Q, Li, Z, Li, J, Xu, Y. | | Deposit date: | 2015-08-19 | | Release date: | 2015-11-04 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.971 Å) | | Cite: | Structural insight into substrate preference for TET-mediated oxidation.
Nature, 527, 2015
|
|
5DEU
 
 | | Crystal structure of TET2-5hmC complex | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, CHLORIDE ION, DNA (5'-D(*AP*CP*CP*AP*CP*(5HC)P*GP*GP*TP*GP*GP*T)-3'), ... | | Authors: | Hu, L, Cheng, J, Rao, Q, Li, Z, Li, J, Xu, Y. | | Deposit date: | 2015-08-26 | | Release date: | 2015-11-04 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.801 Å) | | Cite: | Structural insight into substrate preference for TET-mediated oxidation.
Nature, 527, 2015
|
|
4DS0
 
 | | Cell attachment protein VP8* of a human rotavirus specifically interacts with A-type histo-blood group antigen | | Descriptor: | Outer capsid protein VP4, alpha-L-fucopyranose-(1-2)-[2-acetamido-2-deoxy-alpha-D-galactopyranose-(1-3)]beta-D-galactopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose | | Authors: | Hu, L, Crawford, S.E, Czako, R, Cortes-Penfield, N.W, Smith, D.F, Le Pendu, J, Estes, M.K, Prasad, B.V.V. | | Deposit date: | 2012-02-17 | | Release date: | 2012-04-11 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.56 Å) | | Cite: | Cell attachment protein VP8* of a human rotavirus specifically interacts with A-type histo-blood group antigen.
Nature, 485, 2012
|
|
4DRV
 
 | | Cell attachment protein VP8* of a human rotavirus specifically interacts with A-type histo-blood group antigen | | Descriptor: | Outer capsid protein VP4, alpha-L-fucopyranose-(1-2)-[2-acetamido-2-deoxy-alpha-D-galactopyranose-(1-3)]alpha-D-galactopyranose | | Authors: | Hu, L, Crawford, S.E, Czako, R, Cortes-Penfield, N.W, Smith, D.F, Le Pendu, J, Estes, M.K, Prasad, B.V.V. | | Deposit date: | 2012-02-17 | | Release date: | 2012-04-11 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.56 Å) | | Cite: | Cell attachment protein VP8* of a human rotavirus specifically interacts with A-type histo-blood group antigen.
Nature, 485, 2012
|
|
4DRR
 
 | | Cell attachment protein VP8* of a human rotavirus specifically interacts with A-type histo-blood group antigen | | Descriptor: | Outer capsid protein VP4, SODIUM ION | | Authors: | Hu, L, Crawford, S.E, Czako, R, Cortes-Penfield, N.W, Smith, D.F, Le Pendu, J, Estes, M.K, Prasad, B.V.V. | | Deposit date: | 2012-02-17 | | Release date: | 2012-04-11 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Cell attachment protein VP8* of a human rotavirus specifically interacts with A-type histo-blood group antigen.
Nature, 485, 2012
|
|
7RSW
 
 | | Crystal structure of group B human rotavirus VP8* | | Descriptor: | Outer capsid protein VP4, peptide | | Authors: | Hu, L, Salmen, W, Sankaran, B, Prasad, B.V. | | Deposit date: | 2021-08-11 | | Release date: | 2022-07-20 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (1.32 Å) | | Cite: | Novel fold of rotavirus glycan-binding domain predicted by AlphaFold2 and determined by X-ray crystallography.
Commun Biol, 5, 2022
|
|
4NM6
 
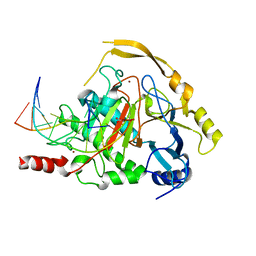 | | Crystal structure of TET2-DNA complex | | Descriptor: | 5'-D(*AP*CP*CP*AP*CP*(5CM)P*GP*GP*TP*GP*GP*T)-3', FE (II) ION, Methylcytosine dioxygenase TET2, ... | | Authors: | Hu, L, Li, Z, Cheng, J, Rao, Q, Gong, W, Liu, M, Wang, P, Xu, Y. | | Deposit date: | 2013-11-14 | | Release date: | 2013-12-18 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.026 Å) | | Cite: | Crystal Structure of TET2-DNA Complex: Insight into TET-Mediated 5mC Oxidation.
Cell(Cambridge,Mass.), 155, 2013
|
|
4YG0
 
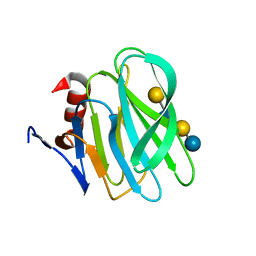 | | Structural basis of glycan recognition in neonate-specific rotaviruses | | Descriptor: | Outer capsid protein VP4, beta-D-galactopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-3)-beta-D-galactopyranose-(1-4)-beta-D-glucopyranose | | Authors: | Hu, L, Prasad, B.V.V. | | Deposit date: | 2015-02-25 | | Release date: | 2015-09-30 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.285 Å) | | Cite: | Structural basis of glycan specificity in neonate-specific bovine-human reassortant rotavirus.
Nat Commun, 6, 2015
|
|
4YG3
 
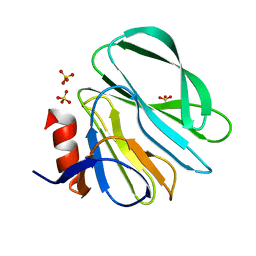 | |
4YFW
 
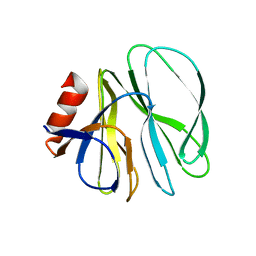 | |
4YG6
 
 | | Structural basis of glycan recognition in neonate-specific rotaviruses | | Descriptor: | Outer capsid protein VP8*, PHOSPHATE ION, beta-D-galactopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Hu, L, Prasad, B.V.V. | | Deposit date: | 2015-02-25 | | Release date: | 2015-09-30 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.46 Å) | | Cite: | Structural basis of glycan specificity in neonate-specific bovine-human reassortant rotavirus.
Nat Commun, 6, 2015
|
|
4YE2
 
 | |
4YFZ
 
 | | Structural basis of glycan recognition in neonate-specific rotaviruses | | Descriptor: | Outer capsid protein VP4, beta-D-galactopyranose-(1-3)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-3)-beta-D-galactopyranose-(1-4)-beta-D-glucopyranose | | Authors: | Hu, L, Prasad, B.V.V. | | Deposit date: | 2015-02-25 | | Release date: | 2015-09-30 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Structural basis of glycan specificity in neonate-specific bovine-human reassortant rotavirus.
Nat Commun, 6, 2015
|
|
4G0A
 
 | |
4G0J
 
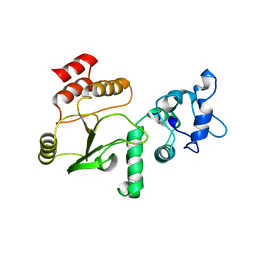 | |
4JW1
 
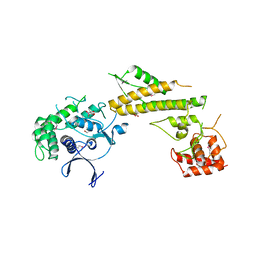 | | Crystal structure of N-terminal 618-residue fragment of LepB from Legionella pneumophila | | Descriptor: | CITRATE ANION, Effector protein B, GLYCEROL | | Authors: | Hu, L, Yao, Q, Zhu, Y, Shao, F. | | Deposit date: | 2013-03-26 | | Release date: | 2013-05-08 | | Last modified: | 2013-08-14 | | Method: | X-RAY DIFFRACTION (3.16 Å) | | Cite: | Structural analyses of Legionella LepB reveal a new GAP fold that catalytically mimics eukaryotic RasGAP
Cell Res., 23, 2013
|
|
3SHB
 
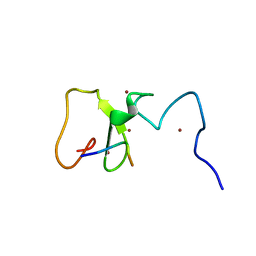 | | Crystal Structure of PHD Domain of UHRF1 | | Descriptor: | E3 ubiquitin-protein ligase UHRF1, Histone H3 peptide, ZINC ION | | Authors: | Hu, L, Li, Z, Wang, P, Lin, Y, Xu, Y. | | Deposit date: | 2011-06-16 | | Release date: | 2011-08-24 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of PHD domain of UHRF1 and insights into recognition of unmodified histone H3 arginine residue 2.
Cell Res., 2011
|
|
5VX4
 
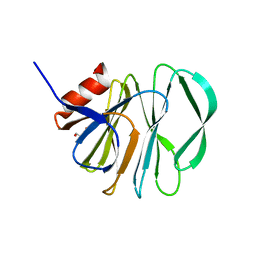 | | VP8* of a G2P[4] Human Rotavirus | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Outer capsid protein VP4 | | Authors: | Hu, L, Venkataram Prasad, B.V. | | Deposit date: | 2017-05-23 | | Release date: | 2018-07-18 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Glycan recognition in globally dominant human rotaviruses.
Nat Commun, 9, 2018
|
|
5VX8
 
 | | VP8* of P[6] Human Rotavirus RV3 | | Descriptor: | Outer capsid protein VP4, SULFATE ION | | Authors: | Hu, L, Venkataram Prasad, B.V. | | Deposit date: | 2017-05-23 | | Release date: | 2018-07-18 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Glycan recognition in globally dominant human rotaviruses.
Nat Commun, 9, 2018
|
|
5VX5
 
 | | VP8* of a G2P[4] Human Rotavirus in complex with LNFP1 | | Descriptor: | Outer capsid protein VP4, alpha-L-fucopyranose-(1-2)-beta-D-galactopyranose-(1-3)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-3)-beta-D-galactopyranose-(1-4)-beta-D-glucopyranose | | Authors: | Hu, L, Venkataram Prasad, B.V. | | Deposit date: | 2017-05-23 | | Release date: | 2018-07-18 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.285 Å) | | Cite: | Glycan recognition in globally dominant human rotaviruses.
Nat Commun, 9, 2018
|
|
5VTH
 
 | | CTX-M-14 P167S:E166A mutant | | Descriptor: | Beta-lactamase | | Authors: | Hu, L, Patel, M, Sankaran, B, Prasad, B.V.V, Palzkill, T. | | Deposit date: | 2017-05-17 | | Release date: | 2017-06-28 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | The Drug-Resistant Variant P167S Expands the Substrate Profile of CTX-M beta-Lactamases for Oxyimino-Cephalosporin Antibiotics by Enlarging the Active Site upon Acylation.
Biochemistry, 56, 2017
|
|
8G0W
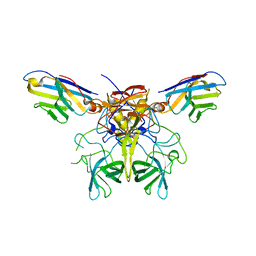 
 | |
