8EQW
 
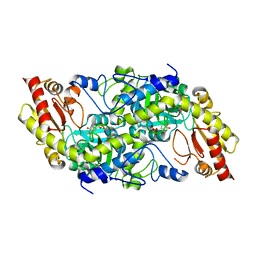 | | Crystal structure of Fub7 | | Descriptor: | Sulfhydrylase FUB7 | | Authors: | Hai, Y. | | Deposit date: | 2022-10-10 | | Release date: | 2023-10-11 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.76 Å) | | Cite: | Molecular and Structural Basis for C gamma-C Bond Formation by PLP-Dependent Enzyme Fub7.
Angew.Chem.Int.Ed.Engl., 63, 2024
|
|
8ERJ
 
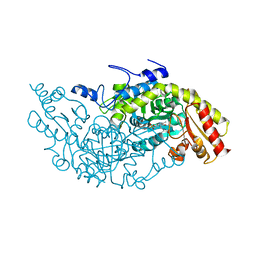 | | Crystal structure of Fub7 in complex with E-2-aminocrotonate | | Descriptor: | (2E)-2-{[(1E)-{3-hydroxy-2-methyl-5-[(phosphonooxy)methyl]pyridin-4-yl}methylidene]amino}but-2-enoic acid, (2S)-2-[({3-hydroxy-2-methyl-5-[(phosphonooxy)methyl]pyridin-4-yl}methyl)amino]but-3-enoic acid, Sulfhydrylase FUB7 | | Authors: | Hai, Y. | | Deposit date: | 2022-10-12 | | Release date: | 2023-10-25 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (2.16 Å) | | Cite: | Molecular and Structural Basis for C gamma-C Bond Formation by PLP-Dependent Enzyme Fub7.
Angew.Chem.Int.Ed.Engl., 63, 2024
|
|
8ERB
 
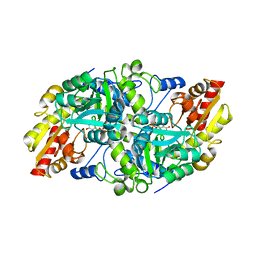 | | Crystal structure of Fub7 in complex with vinylglycine ketimine | | Descriptor: | (2E)-2-[({3-hydroxy-2-methyl-5-[(phosphonooxy)methyl]pyridin-4-yl}methyl)imino]but-3-enoic acid, Sulfhydrylase FUB7 | | Authors: | Hai, Y. | | Deposit date: | 2022-10-11 | | Release date: | 2023-10-18 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.98 Å) | | Cite: | Molecular and Structural Basis for C gamma-C Bond Formation by PLP-Dependent Enzyme Fub7.
Angew.Chem.Int.Ed.Engl., 63, 2024
|
|
5TD7
 
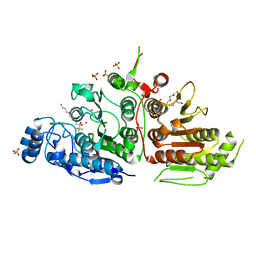 | | Crystal structure of histone deacetylase 10 | | Descriptor: | 7-[(3-aminopropyl)amino]-1,1,1-trifluoroheptane-2,2-diol, PENTAETHYLENE GLYCOL, PHOSPHATE ION, ... | | Authors: | Hai, Y, Shinsky, S.A, Porter, N.J, Christianson, D.W. | | Deposit date: | 2016-09-17 | | Release date: | 2017-05-24 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.85 Å) | | Cite: | Histone deacetylase 10 structure and molecular function as a polyamine deacetylase.
Nat Commun, 8, 2017
|
|
4RHL
 
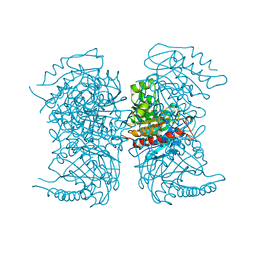 | |
4RHQ
 
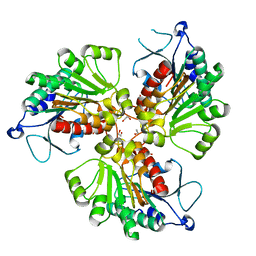 | | Crystal structure of T. brucei arginase-like protein double mutant S149D/S153D | | Descriptor: | 1,2-ETHANEDIOL, Arginase, GLYCEROL | | Authors: | Hai, Y, Barrett, M.P, Christianson, D.W. | | Deposit date: | 2014-10-02 | | Release date: | 2014-12-31 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.18 Å) | | Cite: | Crystal Structure of an Arginase-like Protein from Trypanosoma brucei That Evolved without a Binuclear Manganese Cluster.
Biochemistry, 54, 2015
|
|
4RHJ
 
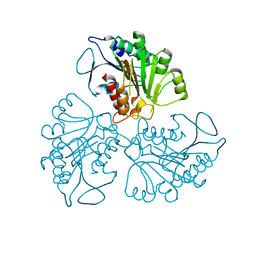 | |
4RHM
 
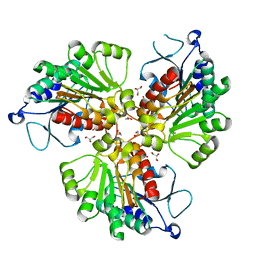 | | Crystal structure of T. brucei arginase-like protein quadruple mutant S149D/R151H/S153D/S226D | | Descriptor: | 1,2-ETHANEDIOL, Arginase, GLYCEROL, ... | | Authors: | Hai, Y, Barrett, M.P, Christianson, D.W. | | Deposit date: | 2014-10-02 | | Release date: | 2014-12-31 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Crystal Structure of an Arginase-like Protein from Trypanosoma brucei That Evolved without a Binuclear Manganese Cluster.
Biochemistry, 54, 2015
|
|
4RHI
 
 | |
4RHK
 
 | |
5HJ9
 
 | | Crystal structure of Leishmania mexicana arginase in complex with inhibitor ABHPE | | Descriptor: | 1,2-ETHANEDIOL, Arginase, GLYCEROL, ... | | Authors: | Hai, Y, Christianson, D.W. | | Deposit date: | 2016-01-13 | | Release date: | 2016-04-13 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.28 Å) | | Cite: | Crystal structures of Leishmania mexicana arginase complexed with alpha , alpha-disubstituted boronic amino-acid inhibitors.
Acta Crystallogr F Struct Biol Commun, 72, 2016
|
|
5HJA
 
 | | Crystal structure of Leishmania mexicana arginase in complex with inhibitor ABHDP | | Descriptor: | (R)-2-amino-6-borono-2-(1-(3,4-dichlorobenzyl)piperidin-4-yl)hexanoic acid, Arginase, GLYCEROL, ... | | Authors: | Hai, Y, Christianson, D.W. | | Deposit date: | 2016-01-13 | | Release date: | 2016-04-13 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Crystal structures of Leishmania mexicana arginase complexed with alpha , alpha-disubstituted boronic amino-acid inhibitors.
Acta Crystallogr F Struct Biol Commun, 72, 2016
|
|
5EF7
 
 | |
5EFN
 
 | |
5EEI
 
 | |
5EEK
 
 | |
5EFG
 
 | |
5EEN
 
 | |
5EFK
 
 | |
5EEF
 
 | |
5EFH
 
 | | Crystal structure of Danio rerio histone deacetylase 6 catalytic domain 2 in complex with trifluoroketone transition state analogue | | Descriptor: | 1,2-ETHANEDIOL, 7-[(3-aminopropyl)amino]-1,1,1-trifluoroheptane-2,2-diol, BROMIDE ION, ... | | Authors: | Hai, Y, Christianson, D.W. | | Deposit date: | 2015-10-23 | | Release date: | 2016-07-27 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.162 Å) | | Cite: | Histone deacetylase 6 structure and molecular basis of catalysis and inhibition.
Nat.Chem.Biol., 12, 2016
|
|
5EDU
 
 | |
5EF8
 
 | |
5EFJ
 
 | |
5EEM
 
 | |
