3P59
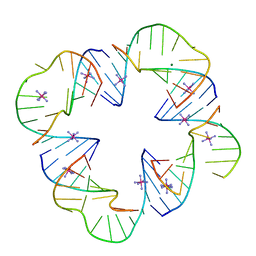 
 | | First Crystal Structure of a RNA Nanosquare | | Descriptor: | COBALT HEXAMMINE(III), MAGNESIUM ION, RNA (5'-R(*CP*CP*GP*GP*AP*GP*GP*AP*AP*CP*UP*AP*CP*(5BU)P*G)-3'), ... | | Authors: | Dibrov, S, Hermann, T, McLean, J, Parsons, J. | | Deposit date: | 2010-10-08 | | Release date: | 2011-04-06 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.1793 Å) | | Cite: | Self-assembling RNA square.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
3TZR
 
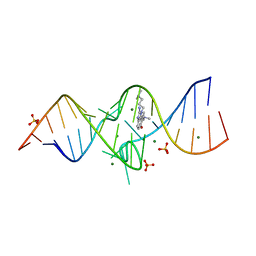 | | Structure of a Riboswitch-like RNA-ligand complex from the Hepatitis C Virus Internal Ribosome Entry Site | | Descriptor: | (8R)-8-[(dimethylamino)methyl]-1-[3-(dimethylamino)propyl]-1,7,8,9-tetrahydrochromeno[5,6-d]imidazol-2-amine, 5'-R(*CP*GP*AP*GP*GP*AP*AP*CP*UP*AP*CP*UP*GP*UP*CP*UP*UP*CP*CP*C)-3', 5'-R(*GP*GP*UP*CP*GP*UP*GP*CP*AP*GP*CP*CP*UP*CP*GP*G)-3', ... | | Authors: | Dibrov, S.M, Ding, K, Brunn, N, Parker, M.A, Bergdahl, B.M, Wyles, D.L, Hermann, T. | | Deposit date: | 2011-09-27 | | Release date: | 2012-03-21 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.212 Å) | | Cite: | Structure of a Riboswitch in the Hepatitis C Virus Internal Ribosome Entry Site
To be Published
|
|
3LOA
 
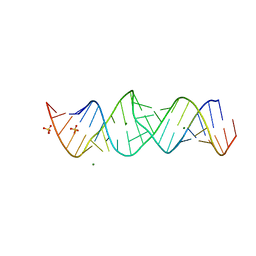 | |
2NOK
 
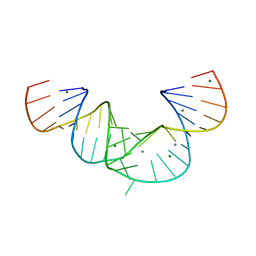 | | Crystal Structure of an RNA domain from Hepatitis C virus. | | Descriptor: | 5'-R(*CP*GP*GP*AP*GP*GP*AP*AP*CP*UP*AP*CP*UP*GP*UP*CP*UP*UP*CP*AP*CP*GP*CP*C)-3', 5'-R(*GP*CP*GP*UP*GP*UP*CP*GP*UP*GP*CP*AP*GP*CP*CP*UP*CP*CP*GP*G)-3', MAGNESIUM ION, ... | | Authors: | Dibrov, S.M, Johnston-Cos, H, Weng, Y.H. | | Deposit date: | 2006-10-25 | | Release date: | 2007-02-13 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Functional architecture of HCV IRES domain II stabilized by divalent metal ions in the crystal and in solution.
Angew.Chem.Int.Ed.Engl., 46, 2007
|
|
3MEI
 
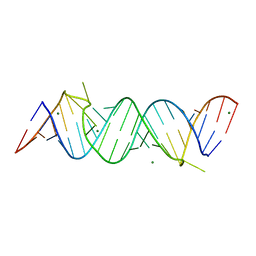 | | Regulatory motif from the thymidylate synthase mRNA | | Descriptor: | MAGNESIUM ION, RNA (5'-R(*CP*CP*GP*CP*CP*GP*CP*GP*CP*CP*AP*(5BU)P*GP*CP*CP*UP*GP*UP*GP*GP*CP*GP*G)-3') | | Authors: | Dibrov, S, McLean, J, Hermann, T. | | Deposit date: | 2010-03-31 | | Release date: | 2011-01-26 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.968 Å) | | Cite: | Structure of an RNA dimer of a regulatory element from human thymidylate synthase mRNA.
Acta Crystallogr.,Sect.D, 67, 2011
|
|
4H1I
 
 | |
4GYH
 
 | |
5CNR
 
 | |
4P97
 
 | |
4PHY
 
 | | Functional conservation despite structural divergence in ligand-responsive RNA switches | | Descriptor: | ACETATE ION, MAGNESIUM ION, RNA (26-MER), ... | | Authors: | Boerneke, M.A, Dibrov, S.M, Hermann, T.H. | | Deposit date: | 2014-05-07 | | Release date: | 2015-02-18 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Functional conservation despite structural divergence in ligand-responsive RNA switches.
Proc.Natl.Acad.Sci.USA, 111, 2014
|
|
