1QH2
 
 | |
2JZM
 
 | |
2C6E
 
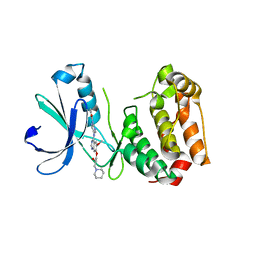 | | Aurora A kinase activated mutant (T287D) in complex with a 5- aminopyrimidinyl quinazoline inhibitor | | Descriptor: | N-{5-[(7-{[(2S)-2-HYDROXY-3-PIPERIDIN-1-YLPROPYL]OXY}-6-METHOXYQUINAZOLIN-4-YL)AMINO]PYRIMIDIN-2-YL}BENZAMIDE, SERINE/THREONINE-PROTEIN KINASE 6 | | Authors: | Pauptit, R.A, Pannifer, A.D, Breed, J, McMiken, H.H.J, Rowsell, S, Anderson, M. | | Deposit date: | 2005-11-09 | | Release date: | 2006-01-11 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Sar and Inhibitor Complex Structure Determination of a Novel Class of Potent and Specific Aurora Kinase Inhibitors.
Bioorg.Med.Chem.Lett., 16, 2006
|
|
2C6D
 
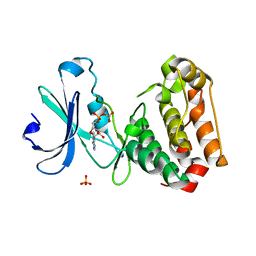 | | Aurora A kinase activated mutant (T287D) in complex with ADPNP | | Descriptor: | GLYCEROL, PHOSPHATE ION, PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER, ... | | Authors: | Pauptit, R.A, Pannifer, A.D, Breed, J, McMiken, H.H.J, Rowsell, S, Anderson, M. | | Deposit date: | 2005-11-09 | | Release date: | 2006-01-11 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Sar and Inhibitor Complex Structure Determination of a Novel Class of Potent and Specific Aurora Kinase Inhibitors.
Bioorg.Med.Chem.Lett., 16, 2006
|
|
1WN4
 
 | | NMR Structure of VoNTR | | Descriptor: | VoNTR protein | | Authors: | Dutton, J.L, Renda, R.F, Waine, C, Clark, R.J, Daly, N.L, Jennings, C.V, Anderson, M.A, Craik, D.J. | | Deposit date: | 2004-07-27 | | Release date: | 2004-09-14 | | Last modified: | 2022-03-02 | | Method: | SOLUTION NMR | | Cite: | Conserved structural and sequence elements implicated in the processing of gene-encoded circular proteins
J.Biol.Chem., 279, 2004
|
|
1WN8
 
 | | NMR Structure of OaNTR | | Descriptor: | Kalata B3/B6 | | Authors: | Dutton, J.L, Renda, R.F, Waine, C, Clark, R.J, Daly, N.L, Jennings, C.V, Anderson, M.A, Craik, D.J. | | Deposit date: | 2004-07-28 | | Release date: | 2004-09-14 | | Last modified: | 2022-03-02 | | Method: | SOLUTION NMR | | Cite: | Conserved structural and sequence elements implicated in the processing of gene-encoded circular proteins
J.Biol.Chem., 279, 2004
|
|
1YTP
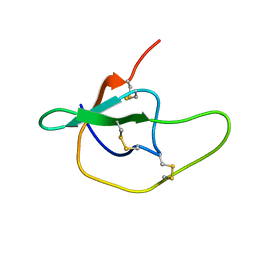 
 | |
4E3R
 
 | | PLP-bound aminotransferase mutant crystal structure from Vibrio fluvialis | | Descriptor: | Pyruvate transaminase, SODIUM ION, SULFATE ION | | Authors: | Midelfort, K.S, Kumar, R, Han, S, Karmilowicz, M.J, McConnell, K, Gehlhaar, D.K, Mistry, A, Chang, J.S, Anderson, M, Vilalobos, A, Minshull, J, Govindarajan, S, Wong, J.W. | | Deposit date: | 2012-03-10 | | Release date: | 2012-10-10 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Redesigning and characterizing the substrate specificity and activity of Vibrio fluvialis aminotransferase for the synthesis of imagabalin.
Protein Eng.Des.Sel., 26, 2013
|
|
4E3Q
 
 | | PMP-bound form of Aminotransferase crystal structure from Vibrio fluvialis | | Descriptor: | 4'-DEOXY-4'-AMINOPYRIDOXAL-5'-PHOSPHATE, BENZAMIDINE, Pyruvate transaminase, ... | | Authors: | Midelfort, K.S, Kumar, R, Han, S, Karmilowicz, M.J, McConnell, K, Gehlhaar, D.K, Mistry, A, Chang, J.S, Anderson, M, Vilalobos, A, Minshull, J, Govindarajan, S, Wong, J.W. | | Deposit date: | 2012-03-10 | | Release date: | 2012-10-10 | | Last modified: | 2013-01-02 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Redesigning and characterizing the substrate specificity and activity of Vibrio fluvialis aminotransferase for the synthesis of imagabalin.
Protein Eng.Des.Sel., 26, 2013
|
|
1PT4
 
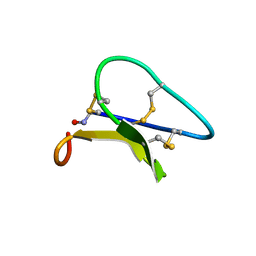 | | Solution structure of the Moebius cyclotide kalata B2 | | Descriptor: | kalata B2 | | Authors: | Jennings, C.V, Anderson, M.A, Daly, N.L, Rosengren, K.J, Craik, D.J. | | Deposit date: | 2003-06-23 | | Release date: | 2004-08-10 | | Last modified: | 2022-03-02 | | Method: | SOLUTION NMR | | Cite: | Isolation, Solution Structure, and Insecticidal Activity of Kalata B2, a Circular Protein with a Twist: Do Mobius Strips Exist in Nature?(,)
Biochemistry, 44, 2005
|
|
1TIH
 
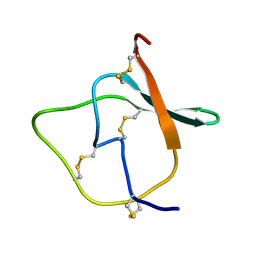 | |
1MR4
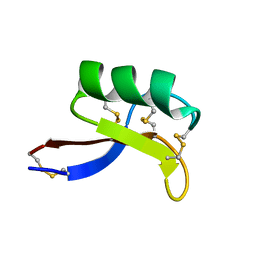 
 | | Solution Structure of NaD1 from Nicotiana alata | | Descriptor: | Nicotiana alata plant defensin 1 (NaD1) | | Authors: | Lay, F.T, Schirra, H.J, Scanlon, M.J, Anderson, M.A, Craik, D.J. | | Deposit date: | 2002-09-18 | | Release date: | 2003-09-18 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | The Three-dimensional Solution Structure of NaD1, a New Floral Defensin from Nicotiana alata and its Application to a Homology Model of the Crop Defense Protein alfAFP
J.MOL.BIOL., 325, 2003
|
|
2JYY
 
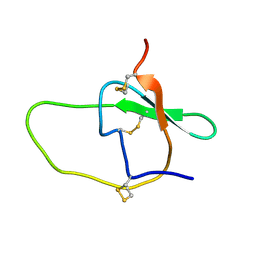 | |
1CE3
 | |
2YIW
 
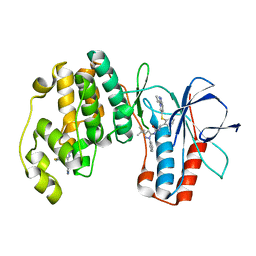 | | triazolopyridine inhibitors of p38 kinase | | Descriptor: | 1-(3-tert-butyl-1-phenyl-1H-pyrazol-5-yl)-3-(2-{[3-(1-methylethyl)[1,2,4]triazolo[4,3-a]pyridin-6-yl]sulfanyl}benzyl)urea, 2-fluoro-4-[4-(4-fluorophenyl)-1H-pyrazol-3-yl]pyridine, MITOGEN-ACTIVATED PROTEIN KINASE 14 | | Authors: | Millan, D.S, Anderson, M, Bunnage, M.E, Burrows, J.L, Butcher, K.J, Dodd, P.G, Evans, T.J, Fairman, D.A, Hughes, S.J, Irving, S.L, Kilty, I.C, Lemaitre, A, Lewthwaite, R.A, Mahnke, A, Mathais, J.P, Philip, J, Phillips, C, Smith, R.T, Stefamiak, M.H, Yeadon, M. | | Deposit date: | 2011-05-17 | | Release date: | 2011-11-30 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Design and Synthesis of Inhaled P38 Inhibitors for the Treatment of Chronic Obstructive Pulmonary Disease.
J.Med.Chem., 54, 2011
|
|
2YIS
 
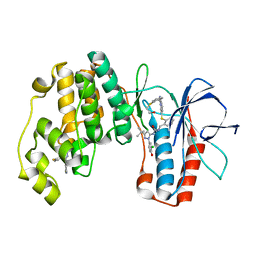 | | triazolopyridine inhibitors of p38 kinase. | | Descriptor: | 1-[3-tert-butyl-1-(3-chloro-4-hydroxyphenyl)-1H-pyrazol-5-yl]-3-{2-[(3-{2-[(2-hydroxyethyl)sulfanyl]phenyl}[1,2,4]triazolo[4,3-a]pyridin-6-yl)sulfanyl]benzyl}urea, 2-fluoro-4-[4-(4-fluorophenyl)-1H-pyrazol-3-yl]pyridine, MITOGEN-ACTIVATED PROTEIN KINASE 14 | | Authors: | Millan, D.S, Anderson, M, Bazin, R, Bunnage, M.E, Burrows, J.L, Butcher, K.J, Dodd, P.G, Evans, T.J, Fairman, D.A, Hughes, S.J, Irving, S.L, Kilty, I.C, Lemaitre, A, Lewthwaite, R.A, Mahnke, A, Mathais, J.P, Philip, J, Phillips, C, Smith, R.T, Stefaniack, M.H, Yeadon, M. | | Deposit date: | 2011-05-16 | | Release date: | 2011-11-30 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Design and Synthesis of Inhaled P38 Inhibitors for the Treatment of Chronic Obstructive Pulmonary Disease.
J.Med.Chem., 54, 2011
|
|
1FYB
 
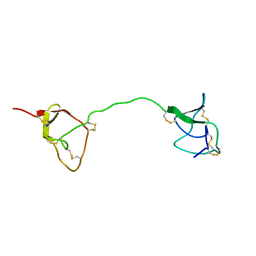 | | SOLUTION STRUCTURE OF C1-T1, A TWO-DOMAIN PROTEINASE INHIBITOR DERIVED FROM THE CIRCULAR PRECURSOR PROTEIN NA-PROPI FROM NICOTIANA ALATA | | Descriptor: | PROTEINASE INHIBITOR | | Authors: | Craik, D.J, Schirra, H.J, Scanlon, M.J, Anderson, M.A. | | Deposit date: | 2000-09-28 | | Release date: | 2001-02-21 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | The solution structure of C1-T1, a two-domain proteinase inhibitor derived from a circular precursor protein from Nicotiana alata.
J.Mol.Biol., 306, 2001
|
|
2VV9
 
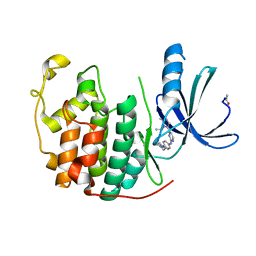 | | CDK2 in complex with an imidazole piperazine | | Descriptor: | 2-{4-[4-({4-[2-methyl-1-(1-methylethyl)-1H-imidazol-5-yl]pyrimidin-2-yl}amino)phenyl]piperazin-1-yl}-2-oxoethanol, CELL DIVISION PROTEIN KINASE 2 | | Authors: | Acton, D.G, Anderson, M, Andrews, D.M, Barker, A.J, Brassington, C.A, Finlay, M.R, Fisher, E, Gerhardt, S, Graham, M.A, Green, C.P, Heaton, D.W, Loddick, S.A, Morgentin, R, Read, J, Roberts, A, Stanway, J, Tucker, J.A, Weir, H.M. | | Deposit date: | 2008-06-04 | | Release date: | 2008-08-05 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Imidazole Piperazines: Sar and Development of a Potent Class of Cyclin-Dependent Kinase Inhibitors with a Novel Binding Mode.
Bioorg.Med.Chem.Lett., 18, 2008
|
|
2YIX
 
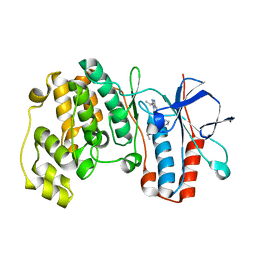 | | Triazolopyridine Inhibitors of p38 | | Descriptor: | 1-ethyl-3-(2-{[3-(1-methylethyl)[1,2,4]triazolo[4,3-a]pyridin-6-yl]sulfanyl}benzyl)urea, MITOGEN-ACTIVATED PROTEIN KINASE 14 | | Authors: | Millan, D.S, Anderson, M, Bunnage, M.E, Burrows, J.L, Butcher, K.J, Dodd, P.G, Evans, T.J, Fairman, D.A, Han, s, Hughes, S.J, Irving, S.L, Kilty, I.C, Lemaitre, A, Lewthawaite, R.A, Mahke, A, Marr, E, Mathias, J.P, Philip, J, Phillips, C, Smith, R.T, Stefaniak, M.H, Yeadon, M. | | Deposit date: | 2011-05-17 | | Release date: | 2011-11-30 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Design and Synthesis of Inhaled P38 Inhibitors for the Treatment of Chronic Obstructive Pulmonary Disease.
J.Med.Chem., 54, 2011
|
|
1N4N
 | | Structure of the Plant Defensin PhD1 from Petunia Hybrida | | Descriptor: | floral defensin-like protein 1 | | Authors: | Janssen, B.J.C, Schirra, H.J, Lay, F.T, Anderson, M.A, Craik, D.J. | | Deposit date: | 2002-11-01 | | Release date: | 2003-11-01 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | Structure of Petunia hybrida defensin 1, a novel plant defensin with five disulfide bonds
Biochemistry, 42, 2003
|
|
1PIC
 
 | | PHOSPHATIDYLINOSITOL 3-KINASE, P85-ALPHA SUBUNIT: C-TERMINAL SH2 DOMAIN COMPLEXED WITH A TYR751 PHOSPHOPEPTIDE FROM THE PDGF RECEPTOR, NMR, MINIMIZED MEAN STRUCTURE | | Descriptor: | BETA-PLATELET-DERIVED GROWTH FACTOR RECEPTOR, PHOSPHATIDYLINOSITOL 3-KINASE | | Authors: | Breeze, A.L, Kara, B.V, Barratt, D.G, Anderson, M, Smith, J.C, Luke, R.W, Best, J.R, Cartlidge, S.A. | | Deposit date: | 1997-06-23 | | Release date: | 1997-09-17 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | Structure of a specific peptide complex of the carboxy-terminal SH2 domain from the p85 alpha subunit of phosphatidylinositol 3-kinase.
EMBO J., 15, 1996
|
|
1T5X
 
 | | HLA-DR1 in complex with a synthetic peptide (AAYSDQATPLLLSPR) and the superantigen SEC3-3B2 | | Descriptor: | 15-mer peptide fragment of Regulatory protein MIG1, Enterotoxin type C-3, HLA class II histocompatibility antigen, ... | | Authors: | Zavala-Ruiz, Z, Strug, I, Anderson, M.W, Gorski, J, Stern, L.J. | | Deposit date: | 2004-05-05 | | Release date: | 2004-08-17 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | A Polymorphic Pocket at the P10 Position Contributes to Peptide Binding Specificity in Class II MHC Proteins
Chem.Biol., 11, 2004
|
|
1T5W
 | | HLA-DR1 in complex with a synthetic peptide (AAYSDQATPLLLSPR) | | Descriptor: | 15-mer peptide fragment of Regulatory protein MIG1, HLA class II histocompatibility antigen, DR alpha chain, ... | | Authors: | Zavala-Ruiz, Z, Strug, I, Anderson, M.W, Gorski, J, Stern, L.J. | | Deposit date: | 2004-05-05 | | Release date: | 2004-08-17 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | A Polymorphic Pocket at the P10 Position Contributes to Peptide Binding Specificity in Class II MHC Proteins
Chem.Biol., 11, 2004
|
|
4ACM
 
 | | CDK2 IN COMPLEX WITH 3-AMINO-6-(4-{[2-(DIMETHYLAMINO)ETHYL]SULFAMOYL}-PHENYL)-N-PYRIDIN-3-YLPYRAZINE-2-CARBOXAMIDE | | Descriptor: | 3-AMINO-6-(4-{[2-(DIMETHYLAMINO)ETHYL]SULFAMOYL}PHENYL)-N-PYRIDIN-3-YLPYRAZINE-2-CARBOXAMIDE, CYCLIN-DEPENDENT KINASE 2, GLYCEROL | | Authors: | Berg, S, Bhat, R, Anderson, M, Bergh, M, Brassington, C, Hellberg, S, Jerning, E, Hogdin, K, Lo-Alfredsson, Y, Neelissen, J, Nilsson, Y, Ormo, M, Soderman, P, Stanway, J, Tucker, J, von Berg, S, Weigelt, T, Xue, Y. | | Deposit date: | 2011-12-16 | | Release date: | 2012-05-16 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.63 Å) | | Cite: | Discovery of novel potent and highly selective glycogen synthase kinase-3 beta (GSK3 beta ) inhibitors for Alzheimer's disease: design, synthesis, and characterization of pyrazines.
J. Med. Chem., 55, 2012
|
|
1JBL
 | | Solution structure of SFTI-1, A cyclic trypsin inhibitor from sunflower seeds | | Descriptor: | CYCLIC TRYPSIN INHIBITOR | | Authors: | Korsinczky, M.L.J, Schirra, H.J, Rosengren, K.J, West, J, Condie, B.A, Otvos, L, Anderson, M.A, Craik, D.J. | | Deposit date: | 2001-06-05 | | Release date: | 2001-08-22 | | Last modified: | 2015-04-15 | | Method: | SOLUTION NMR | | Cite: | Solution structures by 1H NMR of the novel cyclic trypsin inhibitor SFTI-1 from sunflower seeds and an acyclic permutant.
J.Mol.Biol., 311, 2001
|
|
