4GQ3
 
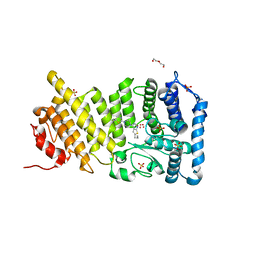 | | Human menin with bound inhibitor MI-2 | | Descriptor: | 4-[4-(5,5-dimethyl-4,5-dihydro-1,3-thiazol-2-yl)piperazin-1-yl]-6-propylthieno[2,3-d]pyrimidine, DI(HYDROXYETHYL)ETHER, Menin, ... | | Authors: | Shi, A, Murai, M.J, He, S, Lund, G.L, Hartley, T, Purohit, T, Reddy, G, Chruszcz, M, Grembecka, J, Cierpicki, T. | | Deposit date: | 2012-08-22 | | Release date: | 2012-09-19 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.56 Å) | | Cite: | Structural insights into inhibition of the bivalent menin-MLL interaction by small molecules in leukemia.
Blood, 120, 2012
|
|
4GQ4
 
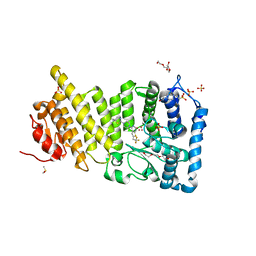 | | Human menin with bound inhibitor MI-2-2 | | Descriptor: | 2-{2-[2-(2-{2-[2-(2-ETHOXY-ETHOXY)-ETHOXY]-ETHOXY}-ETHOXY)-ETHOXY]-ETHOXY}-ETHANOL, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, 4-[4-(5,5-dimethyl-4,5-dihydro-1,3-thiazol-2-yl)piperazin-1-yl]-6-(2,2,2-trifluoroethyl)thieno[2,3-d]pyrimidine, ... | | Authors: | Shi, A, Murai, M.J, He, S, Lund, G.L, Hartley, T, Purohit, T, Reddy, G, Chruszcz, M, Grembecka, J, Cierpicki, T. | | Deposit date: | 2012-08-22 | | Release date: | 2012-09-19 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.27 Å) | | Cite: | Structural insights into inhibition of the bivalent menin-MLL interaction by small molecules in leukemia.
Blood, 120, 2012
|
|
4GQ6
 
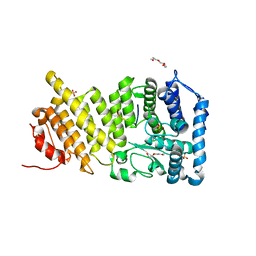 | | Human menin in complex with MLL peptide | | Descriptor: | Histone-lysine N-methyltransferase MLL, Menin, PENTAETHYLENE GLYCOL, ... | | Authors: | Shi, A, Murai, M.J, He, S, Lund, G.L, Hartley, T, Purohit, T, Reddy, G, Chruszcz, M, Grembecka, J, Cierpicki, T. | | Deposit date: | 2012-08-22 | | Release date: | 2012-09-19 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Structural insights into inhibition of the bivalent menin-MLL interaction by small molecules in leukemia.
Blood, 120, 2012
|
|
4GPQ
 
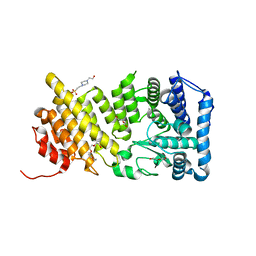 | | Structural insights into inhibition of the bivalent menin-MLL interaction by small molecules in leukemia | | Descriptor: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, DI(HYDROXYETHYL)ETHER, Menin, ... | | Authors: | Shi, A, Murai, M.J, He, S, Lund, G.L, Hartley, T, Purohit, T, Reddy, G, Chruszcz, M, Grembecka, J, Cierpicki, T. | | Deposit date: | 2012-08-21 | | Release date: | 2012-09-19 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.46 Å) | | Cite: | Structural insights into inhibition of the bivalent menin-MLL interaction by small molecules in leukemia.
Blood, 120, 2012
|
|
3RE2
 
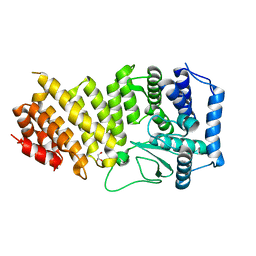 | | Crystal structure of menin reveals the binding site for Mixed Lineage Leukemia (MLL) protein | | Descriptor: | GLYCEROL, Predicted protein | | Authors: | Murai, M.J, Chruszcz, M, Reddy, G, Grembecka, J, Cierpicki, T. | | Deposit date: | 2011-04-02 | | Release date: | 2011-07-13 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Crystal Structure of Menin Reveals Binding Site for Mixed Lineage Leukemia (MLL) Protein.
J.Biol.Chem., 286, 2011
|
|
7ZRN
 
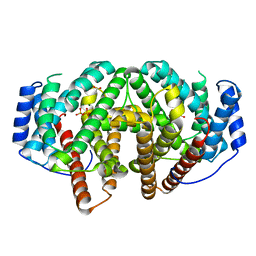 | |
1WS4
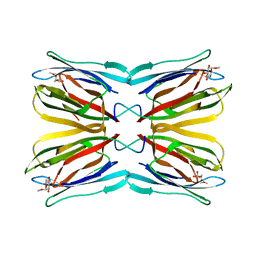 
 | | Crystal structure of Jacalin- Me-alpha-Mannose complex: Promiscuity vs Specificity | | Descriptor: | Agglutinin alpha chain, Agglutinin beta-3 chain, methyl alpha-D-galactopyranoside, ... | | Authors: | Jeyaprakash, A.A, Jayashree, G, Mahanta, S.K, Sekar, K, Surolia, A, Vijayan, M. | | Deposit date: | 2004-10-31 | | Release date: | 2005-03-29 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural basis for the energetics of jacalin-sugar interactions: promiscuity versus specificity
J.Mol.Biol., 347, 2005
|
|
1WS5
 
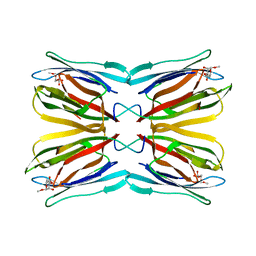 | | Crystal structure of Jacalin-Me-alpha-Mannose complex: Promiscuity vs Specificity | | Descriptor: | Agglutinin alpha chain, Agglutinin beta-3 chain, methyl alpha-D-mannopyranoside | | Authors: | Jeyaprakash, A.A, Jayashree, G, Mahanta, S.K, Sekar, K, Surolia, A, Vijayan, M. | | Deposit date: | 2004-10-31 | | Release date: | 2005-03-29 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural basis for the energetics of jacalin-sugar interactions: promiscuity versus specificity
J.Mol.Biol., 347, 2005
|
|
4PG4
 
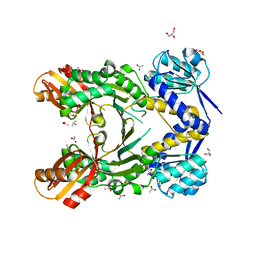 | |
4PG8
 
 | |
4PG5
 
 | |
4PG7
 
 | |
4PG6
 
 | |
7W22
 
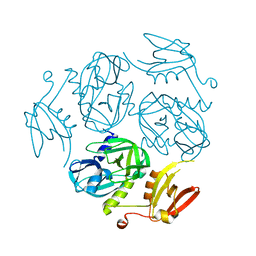 | | Structure of the M. tuberculosis HtrA K436A mutant | | Descriptor: | Probable serine protease HtrA1 | | Authors: | Gupta, A.K, Gopal, B. | | Deposit date: | 2021-11-21 | | Release date: | 2022-11-23 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.01 Å) | | Cite: | Allosteric Determinants in High Temperature Requirement A Enzymes Are Conserved and Regulate the Population of Active Conformations.
Acs Chem.Biol., 18, 2023
|
|
7W25
 
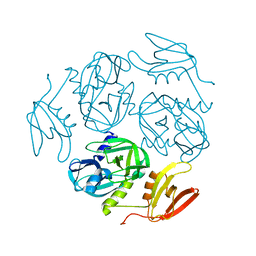 | | Structure of the M. tuberculosis HtrA S413A mutant | | Descriptor: | Probable serine protease HtrA1 | | Authors: | Gupta, A.K, Gopal, B. | | Deposit date: | 2021-11-21 | | Release date: | 2022-11-23 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.65 Å) | | Cite: | Allosteric Determinants in High Temperature Requirement A Enzymes Are Conserved and Regulate the Population of Active Conformations.
Acs Chem.Biol., 18, 2023
|
|
7W23
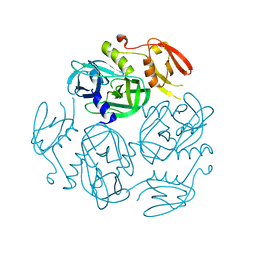 
 | | Structure of the M. tuberculosis HtrA S363A mutant | | Descriptor: | Probable serine protease HtrA1 | | Authors: | Gupta, A.K, Gopal, B. | | Deposit date: | 2021-11-21 | | Release date: | 2022-11-23 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Allosteric Determinants in High Temperature Requirement A Enzymes Are Conserved and Regulate the Population of Active Conformations.
Acs Chem.Biol., 18, 2023
|
|
7W24
 
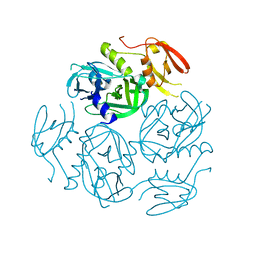 | | Structure of the M. tuberculosis HtrA N383A mutant | | Descriptor: | Probable serine protease HtrA1 | | Authors: | Gupta, A.K, Gopal, B. | | Deposit date: | 2021-11-21 | | Release date: | 2022-11-23 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Allosteric Determinants in High Temperature Requirement A Enzymes Are Conserved and Regulate the Population of Active Conformations.
Acs Chem.Biol., 18, 2023
|
|
7VZ0
 
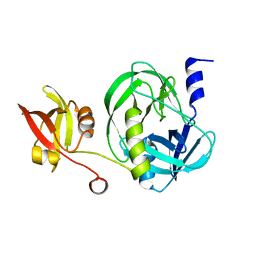 | | Structure of the M. tuberculosis HtrA S407A mutant | | Descriptor: | Probable serine protease HtrA1 | | Authors: | Gupta, A.K, Gopal, B. | | Deposit date: | 2021-11-15 | | Release date: | 2022-11-23 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Allosteric Determinants in High Temperature Requirement A Enzymes Are Conserved and Regulate the Population of Active Conformations.
Acs Chem.Biol., 18, 2023
|
|
7W21
 
 | | Structure of the M. tuberculosis HtrA N269A mutant | | Descriptor: | Probable serine protease HtrA1 | | Authors: | Gupta, A.K, Gopal, B. | | Deposit date: | 2021-11-21 | | Release date: | 2022-11-23 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Allosteric Determinants in High Temperature Requirement A Enzymes Are Conserved and Regulate the Population of Active Conformations.
Acs Chem.Biol., 18, 2023
|
|
7VYZ
 
 | | Structure of the M. tuberculosis HtrA S367A mutant | | Descriptor: | Probable serine protease HtrA1 | | Authors: | Gupta, A.K, Gopal, B. | | Deposit date: | 2021-11-15 | | Release date: | 2022-11-23 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.401 Å) | | Cite: | Allosteric Determinants in High Temperature Requirement A Enzymes Are Conserved and Regulate the Population of Active Conformations.
Acs Chem.Biol., 18, 2023
|
|
7W4R
 
 | |
7W4W
 
 | |
7W4T
 
 | |
7W4U
 
 | |
7W4S
 
 | |
