1MMM
 
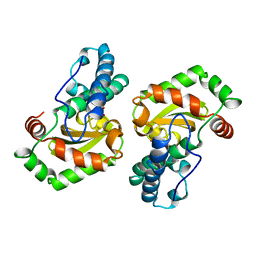 | | DISTINCT METAL ENVIRONMENT IN IRON-SUBSTITUTED MANGANESE SUPEROXIDE DISMUTASE PROVIDES A STRUCTURAL BASIS OF METAL SPECIFICITY | | Descriptor: | FE (III) ION, HYDROXIDE ION, PROTEIN (IRON-SUBSTITUTED MANGANESE SUPEROXIDE DISMUTASE) | | Authors: | Edwards, R.A, Whittaker, M.M, Whittaker, J.W, Jameson, G.B, Baker, E.N. | | Deposit date: | 1998-08-26 | | Release date: | 1998-09-02 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Distinct Metal Environment in Fe-Substituted Manganese Superoxide Dismutase Provides a Structural Basis of Metal Specificity
J.Am.Chem.Soc., 120, 1998
|
|
1M4V
 
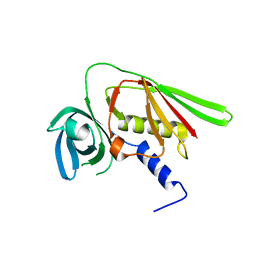 | | Crystal structure of SET3, a superantigen-like protein from Staphylococcus aureus | | Descriptor: | SET3, superantigen-like protein | | Authors: | Arcus, V.L, Langley, R, Proft, T, Fraser, J.D, Baker, E.N. | | Deposit date: | 2002-07-05 | | Release date: | 2002-07-17 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | The three-dimensional structure of a superantigen-like protein, SET3, from a pathogenicity island of the Staphylococcus aureus genome
J.Biol.Chem., 277, 2002
|
|
1ND4
 
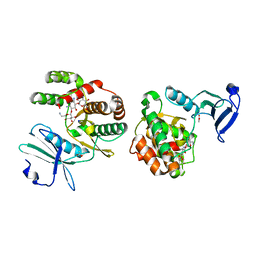 | | Crystal structure of aminoglycoside-3'-phosphotransferase-IIa | | Descriptor: | ACETATE ION, Aminoglycoside 3'-phosphotransferase, KANAMYCIN A, ... | | Authors: | Nurizzo, D, Shewry, S.C, Baker, E.N, Smith, C.A. | | Deposit date: | 2002-12-06 | | Release date: | 2003-09-16 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | The crystal structure of aminoglycoside-3'-phosphotransferase-IIa, an enzyme responsible for antibiotic resistance
J.Mol.Biol., 327, 2003
|
|
1E3J
 
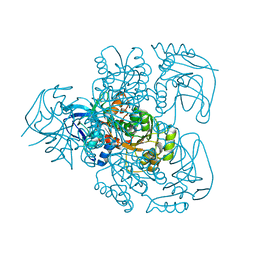 | | Ketose reductase (sorbitol dehydrogenase) from silverleaf whitefly | | Descriptor: | BORIC ACID, NADP(H)-DEPENDENT KETOSE REDUCTASE, PHOSPHATE ION, ... | | Authors: | Banfield, M.J, Salvucci, M.E, Baker, E.N, Smith, C.A. | | Deposit date: | 2000-06-19 | | Release date: | 2001-02-04 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal Structure of Nadp(H)-Dependent Ketose Reductase from Besimia Argentifolii at 2.3 Angstrom Resolution
J.Mol.Biol., 306, 2001
|
|
1EEJ
 
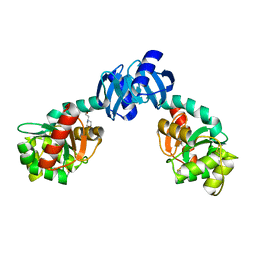 | | CRYSTAL STRUCTURE OF THE PROTEIN DISULFIDE BOND ISOMERASE, DSBC, FROM ESCHERICHIA COLI | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, THIOL:DISULFIDE INTERCHANGE PROTEIN | | Authors: | McCarthy, A.A, Haebel, P.W, Torronen, A, Rybin, V, Baker, E.N, Metcalf, P. | | Deposit date: | 2000-01-31 | | Release date: | 2000-08-03 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structure of the protein disulfide bond isomerase, DsbC, from Escherichia coli.
Nat.Struct.Biol., 7, 2000
|
|
1EH3
 
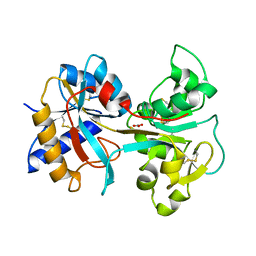 | | R210K N-TERMINAL LOBE HUMAN LACTOFERRIN | | Descriptor: | CARBONATE ION, FE (III) ION, LACTOFERRIN | | Authors: | Peterson, N.A, Anderson, B.F, Jameson, G.B, Tweedie, J.W, Baker, E.N. | | Deposit date: | 2000-02-18 | | Release date: | 2000-03-01 | | Last modified: | 2021-11-03 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structure and iron-binding properties of the R210K mutant of the N-lobe of human lactoferrin: implications for iron release from transferrins.
Biochemistry, 39, 2000
|
|
2GL0
 
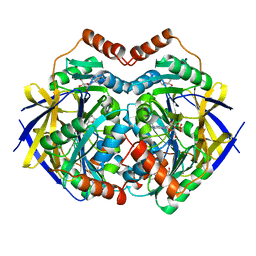 | | Structure of PAE2307 in complex with adenosine | | Descriptor: | ADENOSINE, PHOSPHATE ION, conserved hypothetical protein | | Authors: | Lott, J.S, Paget, B, Johnston, J.M, Baker, E.N. | | Deposit date: | 2006-04-04 | | Release date: | 2006-06-06 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | The Structure of an Ancient Conserved Domain Establishes a Structural Basis for Stable Histidine Phosphorylation and Identifies a New Family of Adenosine-specific Kinases.
J.Biol.Chem., 281, 2006
|
|
2HYX
 
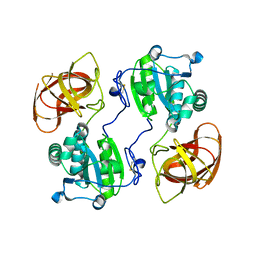 | |
2HUO
 
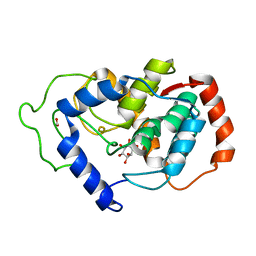 | | Crystal structure of mouse myo-inositol oxygenase in complex with substrate | | Descriptor: | 1,2,3,4,5,6-HEXAHYDROXY-CYCLOHEXANE, FE (III) ION, FORMIC ACID, ... | | Authors: | Brown, P.M, Caradoc-Davies, T.T, Dickson, J.M.J, Cooper, G.J.S, Loomes, K.M, Baker, E.N. | | Deposit date: | 2006-07-27 | | Release date: | 2006-09-26 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structure of a substrate complex of myo-inositol oxygenase, a di-iron oxygenase with a key role in inositol metabolism.
Proc.Natl.Acad.Sci.Usa, 103, 2006
|
|
1F5J
 
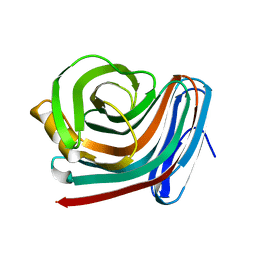 | | CRYSTAL STRUCTURE OF XYNB, A HIGHLY THERMOSTABLE BETA-1,4-XYLANASE FROM DICTYOGLOMUS THERMOPHILUM RT46B.1, AT 1.8 A RESOLUTION | | Descriptor: | BETA-1,4-XYLANASE, SULFATE ION | | Authors: | McCarthy, A.A, Baker, E.N. | | Deposit date: | 2000-07-26 | | Release date: | 2000-11-15 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structure of XynB, a highly thermostable beta-1,4-xylanase from Dictyoglomus thermophilum Rt46B.1, at 1.8 A resolution.
Acta Crystallogr.,Sect.D, 56, 2000
|
|
1H6A
 
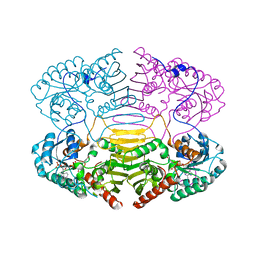 | |
1H6D
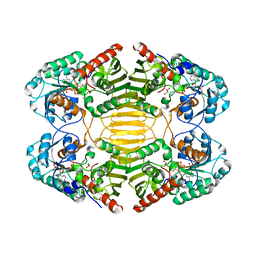 
 | |
1H5Y
 
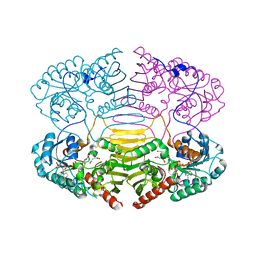
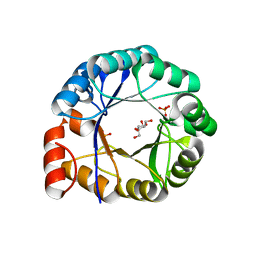 | | HisF protein from Pyrobaculum aerophilum | | Descriptor: | GLYCEROL, HISF, PHOSPHATE ION | | Authors: | Banfield, M.J, Lott, J.S, McCarthy, A.A, Baker, E.N. | | Deposit date: | 2001-05-31 | | Release date: | 2001-06-01 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure of Hisf, a Histidine Biosynthetic Protein from Pyrobaculum Aerophilum
Acta Crystallogr.,Sect.D, 57, 2001
|
|
1H6B
 
 | |
1H6C
 
 | |
4INS
 
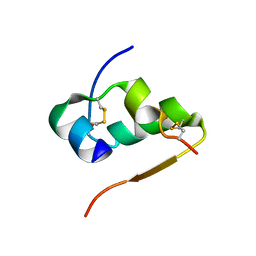 | | THE STRUCTURE OF 2ZN PIG INSULIN CRYSTALS AT 1.5 ANGSTROMS RESOLUTION | | Descriptor: | INSULIN (CHAIN A), INSULIN (CHAIN B), ZINC ION | | Authors: | Dodson, G.G, Dodson, E.J, Hodgkin, D.C, Isaacs, N.W, Vijayan, M. | | Deposit date: | 1989-07-10 | | Release date: | 1990-04-15 | | Last modified: | 2017-11-29 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | The structure of 2Zn pig insulin crystals at 1.5 A resolution.
Philos.Trans.R.Soc.London,Ser.B, 319, 1988
|
|
2R61
 
 | |
1OQH
 
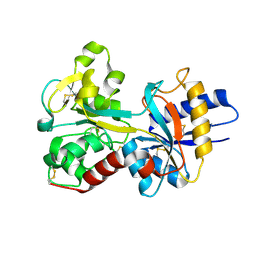 | | Crystal Structure of the R124A mutant of the N-lobe human transferrin | | Descriptor: | CARBONATE ION, FE (III) ION, POTASSIUM ION, ... | | Authors: | Baker, H.M, He, Q.-Y, Brigg, S.K, Mason, A.B, N Baker, E. | | Deposit date: | 2003-03-10 | | Release date: | 2003-03-18 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structural and functional consequences of binding site mutations in transferrin: crystal structures of the Asp63Glu and Arg124Ala mutants of the N-lobe of human transferrin
Biochemistry, 42, 2003
|
|
2AKQ
 
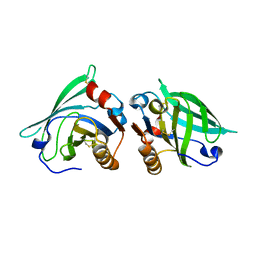 | | The structure of bovine B-lactoglobulin A in crystals grown at very low ionic strength | | Descriptor: | Beta-lactoglobulin variant A | | Authors: | Adams, J.J, Anderson, B.F, Norris, G.E, Creamer, L.K, Jameson, G.B. | | Deposit date: | 2005-08-03 | | Release date: | 2005-08-16 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structure of bovine beta-lactoglobulin (variant A) at very low ionic strength
J.Struct.Biol., 154, 2006
|
|
4X58
 
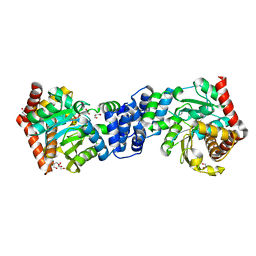 | | Anthranilate phosphoribosyl transferase variant N138A from Mycobacterium tuberculosis in complex with PRPP and Mg | | Descriptor: | 1-O-pyrophosphono-5-O-phosphono-alpha-D-ribofuranose, Anthranilate phosphoribosyltransferase, D-MALATE, ... | | Authors: | Cookson, T.V.M, Evans, G.L, Parker, E.J, Lott, J.S. | | Deposit date: | 2014-12-04 | | Release date: | 2015-11-25 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Structures of Mycobacterium tuberculosis Anthranilate Phosphoribosyltransferase Variants Reveal the Conformational Changes That Facilitate Delivery of the Substrate to the Active Site.
Biochemistry, 54, 2015
|
|
4X59
 
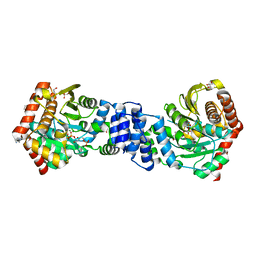 | | Anthranilate phosphoribosyltransferase variant P180A from Mycobacterium tuberculosis in complex with PRPP and Mg | | Descriptor: | 1-O-pyrophosphono-5-O-phosphono-alpha-D-ribofuranose, Anthranilate phosphoribosyltransferase, GLYCEROL, ... | | Authors: | Cookson, T.V.M, Evans, G.L, Parker, E.J, Lott, J.S. | | Deposit date: | 2014-12-04 | | Release date: | 2015-11-25 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structures of Mycobacterium tuberculosis Anthranilate Phosphoribosyltransferase Variants Reveal the Conformational Changes That Facilitate Delivery of the Substrate to the Active Site.
Biochemistry, 54, 2015
|
|
4X5E
 
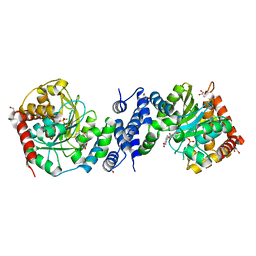 | | Anthranilate phosphoribosyltransferase variant R194A from Mycobacterium tuberculosis with pyrophosphate, Mg2+ and anthranilate bound | | Descriptor: | 2-AMINOBENZOIC ACID, Anthranilate phosphoribosyltransferase, GLYCEROL, ... | | Authors: | Cookson, T.V.M, Evans, G.L, Parker, E.J, Lott, J.S. | | Deposit date: | 2014-12-05 | | Release date: | 2015-11-25 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.77 Å) | | Cite: | Structures of Mycobacterium tuberculosis Anthranilate Phosphoribosyltransferase Variants Reveal the Conformational Changes That Facilitate Delivery of the Substrate to the Active Site.
Biochemistry, 54, 2015
|
|
4X5A
 
 | |
4X5C
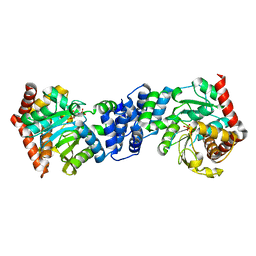 
 | | Anthranilate phosphoribosyltransferase variant R193L from Mycobacterium tuberculosis with pyrophosphate/PRPP and Mg2+ bound | | Descriptor: | 1-O-pyrophosphono-5-O-phosphono-alpha-D-ribofuranose, Anthranilate phosphoribosyltransferase, MAGNESIUM ION, ... | | Authors: | Cookson, T.V.M, Parker, E.J, Lott, J.S. | | Deposit date: | 2014-12-05 | | Release date: | 2015-11-25 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.33 Å) | | Cite: | Structures of Mycobacterium tuberculosis Anthranilate Phosphoribosyltransferase Variants Reveal the Conformational Changes That Facilitate Delivery of the Substrate to the Active Site.
Biochemistry, 54, 2015
|
|
4X5B
 
 | |
