5OAQ
 
 | |
5NO6
 
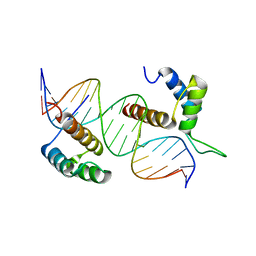 | | TEAD4-HOXB13 complex bound to DNA | | Descriptor: | DNA, Homeobox protein Hox-B13, Transcriptional enhancer factor TEF-3 | | Authors: | Morgunova, E, Jolma, A, Yin, Y, Popov, A, Taipale, J. | | Deposit date: | 2017-04-11 | | Release date: | 2018-05-16 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.88 Å) | | Cite: | TEAD4-HOXB13 complex bound to DNA
To Be Published
|
|
3JUA
 
 | |
5GZB
 
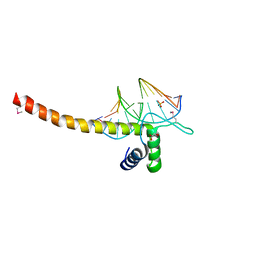 | | Crystal Structure of Transcription Factor TEAD4 in Complex with M-CAT DNA | | Descriptor: | DNA (5'-D(*GP*AP*GP*AP*GP*GP*AP*AP*TP*GP*CP*AP*A)-3'), DNA (5'-D(*TP*TP*GP*CP*AP*TP*TP*CP*CP*TP*CP*TP*C)-3'), GLYCEROL, ... | | Authors: | He, F, Shi, Z.B, Zhou, Z.C. | | Deposit date: | 2016-09-28 | | Release date: | 2017-04-19 | | Last modified: | 2017-08-09 | | Method: | X-RAY DIFFRACTION (2.704 Å) | | Cite: | DNA-binding mechanism of the Hippo pathway transcription factor TEAD4
Oncogene, 36, 2017
|
|
6L9F
 
 | |
5Z2Q
 
 | | Vgll1-TEAD4 core complex | | Descriptor: | PHOSPHATE ION, Transcription cofactor vestigial-like protein 1, Transcriptional enhancer factor TEF-3 | | Authors: | Pobbati, A.V, Song, H. | | Deposit date: | 2018-01-03 | | Release date: | 2018-01-31 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.74 Å) | | Cite: | Structural and functional similarity between the Vgll1-TEAD and the YAP-TEAD complexes.
Structure, 20, 2012
|
|
8A8R
 
 | | Crystal structure of TEAD4 in complex with YAP peptide | | Descriptor: | Isoform 7 of Transcriptional coactivator YAP1, MYRISTIC ACID, Transcriptional enhancer factor TEF-3 | | Authors: | Scheufler, C, Kallen, J. | | Deposit date: | 2022-06-23 | | Release date: | 2022-12-28 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.696 Å) | | Cite: | N-terminal beta-strand in YAP is critical for stronger binding to scalloped relative to TEAD transcription factor.
Protein Sci., 32, 2023
|
|
6SEN
 
 | | TEAD4 bound to a FAM181A peptide | | Descriptor: | Protein FAM181A, SULFATE ION, Transcriptional enhancer factor TEF-3 | | Authors: | Scheufler, C, Villard, F. | | Deposit date: | 2019-07-30 | | Release date: | 2019-11-20 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Identification of FAM181A and FAM181B as new interactors with the TEAD transcription factors.
Protein Sci., 29, 2020
|
|
6SEO
 
 | | TEAD4 bound to a FAM181B peptide | | Descriptor: | Protein FAM181B, Transcriptional enhancer factor TEF-3 | | Authors: | Scheufler, C, Villard, F. | | Deposit date: | 2019-07-30 | | Release date: | 2019-11-20 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Identification of FAM181A and FAM181B as new interactors with the TEAD transcription factors.
Protein Sci., 29, 2020
|
|
6Q36
 
 | |
6Q2X
 
 | |
6GE5
 
 | |
6GEC
 
 | |
6GEI
 
 | |
6GEG
 
 | |
6GEK
 
 | |
6GEE
 
 | |
6GE4
 
 | |
4LN0
 
 | | Crystal structure of the VGLL4-TEAD4 complex | | Descriptor: | DI(HYDROXYETHYL)ETHER, GLYCEROL, Transcription cofactor vestigial-like protein 4, ... | | Authors: | Wang, H, Shi, Z, Zhou, Z. | | Deposit date: | 2013-07-11 | | Release date: | 2014-02-26 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.896 Å) | | Cite: | A Peptide Mimicking VGLL4 Function Acts as a YAP Antagonist Therapy against Gastric Cancer.
Cancer Cell, 25, 2014
|
|
5GN0
 
 | | Structure of TAZ-TEAD complex | | Descriptor: | CITRIC ACID, PALMITIC ACID, Transcriptional enhancer factor TEF-3, ... | | Authors: | Kaan, H.Y.K, Song, H. | | Deposit date: | 2016-07-18 | | Release date: | 2017-05-31 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Crystal structure of TAZ-TEAD complex reveals a distinct interaction mode from that of YAP-TEAD complex
Sci Rep, 7, 2017
|
|
8C17
 
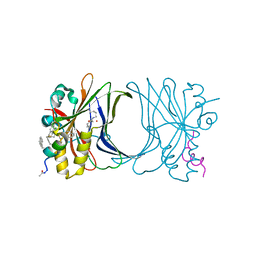 | | Crystal structure of TEAD4 in complex with peptide 1 | | Descriptor: | GLYCEROL, MYRISTIC ACID, Stapled peptide, ... | | Authors: | Scheufler, C, Kallen, J. | | Deposit date: | 2022-12-20 | | Release date: | 2023-03-15 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Biochemical and Structural Characterization of a Peptidic Inhibitor of the YAP:TEAD Interaction That Binds to the alpha-Helix Pocket on TEAD.
Acs Chem.Biol., 18, 2023
|
|
8CAA
 
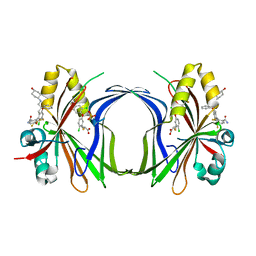 | | Crystal structure of TEAD4 in complex with YTP-13 | | Descriptor: | (2~{R})-2-[2-chloranyl-5-[2-chloranyl-4-(trifluoromethyl)phenoxy]phenyl]sulfanylpropanoic acid, 4-[bis(fluoranyl)methoxy]-2-[(2~{S})-5-chloranyl-6-fluoranyl-2-[[(4-oxidanylcyclohexyl)amino]methyl]-2-phenyl-3~{H}-1-benzofuran-4-yl]-3-fluoranyl-benzamide, PHOSPHATE ION, ... | | Authors: | Scheufler, C, Kallen, J. | | Deposit date: | 2023-01-24 | | Release date: | 2023-04-12 | | Last modified: | 2024-06-19 | | Method: | X-RAY DIFFRACTION (1.999 Å) | | Cite: | Optimization of a Class of Dihydrobenzofurane Analogs toward Orally Efficacious YAP-TEAD Protein-Protein Interaction Inhibitors.
Chemmedchem, 18, 2023
|
|
6GE3
 
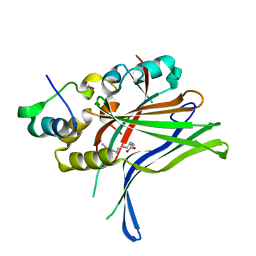 | |
6HIK
 
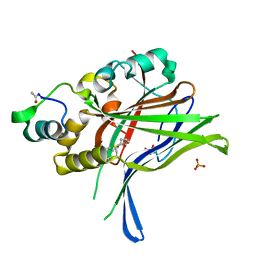 | |
6GE6
 
 | |
