3GWG
 
 | |
3H1F
 
 | |
3H1E
 
 | | Crystal structure of Mg(2+) and BeH(3)(-)-bound CheY of Helicobacter pylori | | Descriptor: | BERYLLIUM TRIFLUORIDE ION, Chemotaxis protein cheY homolog, MAGNESIUM ION, ... | | Authors: | Lam, K.H, Ling, T.K, Au, S.W. | | Deposit date: | 2009-04-12 | | Release date: | 2010-03-09 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal structure of activated CheY1 from Helicobacter pylori.
J.Bacteriol., 192, 2010
|
|
3H1G
 
 | |
2DB7
 
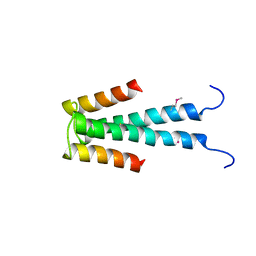 | | Crystal structure of hypothetical protein MS0332 | | Descriptor: | Hairy/enhancer-of-split related with YRPW motif 1 | | Authors: | Wang, H, Takemoto-Hori, C, Murayama, K, Terada, T, Shirouzu, M, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2005-12-15 | | Release date: | 2006-12-19 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structure of hypothetical protein MS0332
To be Published
|
|
3MY0
 
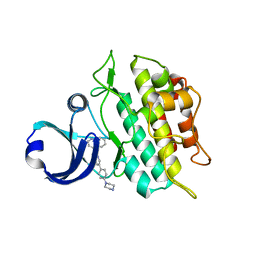 | | Crystal structure of the ACVRL1 (ALK1) kinase domain bound to LDN-193189 | | Descriptor: | 4-[6-(4-piperazin-1-ylphenyl)pyrazolo[1,5-a]pyrimidin-3-yl]quinoline, Serine/threonine-protein kinase receptor R3 | | Authors: | Chaikuad, A, Alfano, I, Cooper, C, Mahajan, P, Daga, N, Sanvitale, C, Fedorov, O, Petrie, K, Savitsky, P, Gileadi, O, Sethi, R, Krojer, T, Muniz, J.R.C, Pike, A.C.W, Vollmar, M, Carpenter, C.P, Ugochukwu, E, Knapp, S, von Delft, F, Weigelt, J, Arrowsmith, C.H, Edwards, A.M, Bountra, C, Bullock, A, Structural Genomics Consortium (SGC) | | Deposit date: | 2010-05-08 | | Release date: | 2010-07-21 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.65 Å) | | Cite: | A small molecule targeting ALK1 prevents Notch cooperativity and inhibits functional angiogenesis.
Angiogenesis, 18, 2015
|
|
3TO5
 
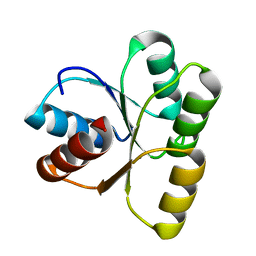 | |
4LX8
 
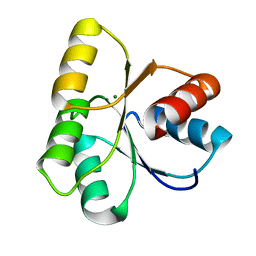 | |
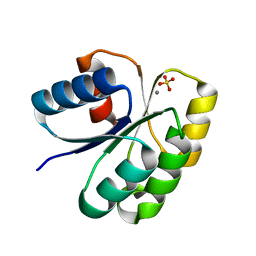
4H60
 
 | |
4HNQ
 
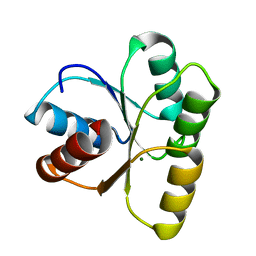 | |
4HNS
 
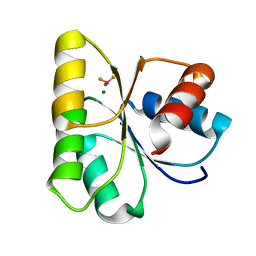 | | Crystal structure of activated CheY3 of Vibrio cholerae | | Descriptor: | BERYLLIUM TRIFLUORIDE ION, Chemotaxis protein CheY, MAGNESIUM ION | | Authors: | Biswas, M, Sen, U, Dasgupta, J. | | Deposit date: | 2012-10-21 | | Release date: | 2013-10-02 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Conformational Barrier of CheY3 and Inability of CheY4 to Bind FliM Control the Flagellar Motor Action in Vibrio cholerae.
Plos One, 8, 2013
|
|
4JP1
 
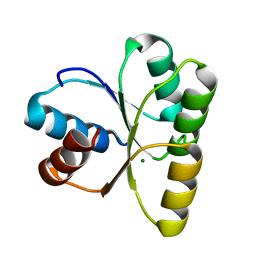 | | Mg2+ bound structure of Vibrio Cholerae CheY3 | | Descriptor: | Chemotaxis protein CheY, MAGNESIUM ION | | Authors: | Biswas, M, Dasgupta, J, Sen, U. | | Deposit date: | 2013-03-19 | | Release date: | 2014-04-02 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.46 Å) | | Cite: | Conformational barrier of CheY3 and inability of CheY4 to bind FliM control the flagellar motor action in Vibrio cholerae
Plos One, 8, 2013
|
|
4HNR
 
 | |
3LXS
 
 | |
2OZ2
 
 | |
