[English] 日本語
 Yorodumi
Yorodumi- EMDB-36091: Muscarinic receptor (M2R) in complex with beta-arrestin1 (Low res... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
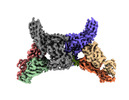
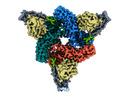
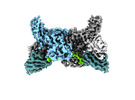
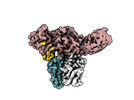
| Title | Muscarinic receptor (M2R) in complex with beta-arrestin1 (Low resolution full map, Cross-linked) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords |  GPCR / GPCR /  Arrestin / Arrestin /  SIGNALING PROTEIN / SIGNALING PROTEIN /  SIGNALING PROTEIN-IMMUNE SYSTEM complex SIGNALING PROTEIN-IMMUNE SYSTEM complex | |||||||||
| Biological species |   Bos taurus (cattle) / Bos taurus (cattle) /   Mus musculus (house mouse) / Mus musculus (house mouse) /   Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 6.53 Å cryo EM / Resolution: 6.53 Å | |||||||||
 Authors Authors | Maharana J / Sano FK / Shihoya W / Banerjee R / Nureki O / Shukla AK | |||||||||
| Funding support |  India, 1 items India, 1 items
| |||||||||
 Citation Citation |  Journal: Science / Year: 2024 Journal: Science / Year: 2024Title: Molecular insights into atypical modes of β-arrestin interaction with seven transmembrane receptors. Authors: Jagannath Maharana / Fumiya K Sano / Parishmita Sarma / Manish K Yadav / Longhan Duan / Tomasz M Stepniewski / Madhu Chaturvedi / Ashutosh Ranjan / Vinay Singh / Sayantan Saha / Gargi ...Authors: Jagannath Maharana / Fumiya K Sano / Parishmita Sarma / Manish K Yadav / Longhan Duan / Tomasz M Stepniewski / Madhu Chaturvedi / Ashutosh Ranjan / Vinay Singh / Sayantan Saha / Gargi Mahajan / Mohamed Chami / Wataru Shihoya / Jana Selent / Ka Young Chung / Ramanuj Banerjee / Osamu Nureki / Arun K Shukla /      Abstract: β-arrestins (βarrs) are multifunctional proteins involved in signaling and regulation of seven transmembrane receptors (7TMRs), and their interaction is driven primarily by agonist-induced receptor ...β-arrestins (βarrs) are multifunctional proteins involved in signaling and regulation of seven transmembrane receptors (7TMRs), and their interaction is driven primarily by agonist-induced receptor activation and phosphorylation. Here, we present seven cryo-electron microscopy structures of βarrs either in the basal state, activated by the muscarinic receptor subtype 2 (M2R) through its third intracellular loop, or activated by the βarr-biased decoy D6 receptor (D6R). Combined with biochemical, cellular, and biophysical experiments, these structural snapshots allow the visualization of atypical engagement of βarrs with 7TMRs and also reveal a structural transition in the carboxyl terminus of βarr2 from a β strand to an α helix upon activation by D6R. Our study provides previously unanticipated molecular insights into the structural and functional diversity encoded in 7TMR-βarr complexes with direct implications for exploring novel therapeutic avenues. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_36091.map.gz emd_36091.map.gz | 59.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-36091-v30.xml emd-36091-v30.xml emd-36091.xml emd-36091.xml | 16.8 KB 16.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_36091_fsc.xml emd_36091_fsc.xml | 8.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_36091.png emd_36091.png | 56.5 KB | ||
| Filedesc metadata |  emd-36091.cif.gz emd-36091.cif.gz | 4.5 KB | ||
| Others |  emd_36091_half_map_1.map.gz emd_36091_half_map_1.map.gz emd_36091_half_map_2.map.gz emd_36091_half_map_2.map.gz | 59.3 MB 59.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-36091 http://ftp.pdbj.org/pub/emdb/structures/EMD-36091 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36091 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36091 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_36091.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_36091.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.3487 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_36091_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_36091_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Muscarinic receptor (M2R) in complex with beta-arrestin1
| Entire | Name: Muscarinic receptor (M2R) in complex with beta-arrestin1 |
|---|---|
| Components |
|
-Supramolecule #1: Muscarinic receptor (M2R) in complex with beta-arrestin1
| Supramolecule | Name: Muscarinic receptor (M2R) in complex with beta-arrestin1 type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 |
|---|
-Supramolecule #2: beta-arrestin1
| Supramolecule | Name: beta-arrestin1 / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:   Bos taurus (cattle) Bos taurus (cattle) |
-Supramolecule #3: Fab30
| Supramolecule | Name: Fab30 / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2-#3 |
|---|---|
| Source (natural) | Organism:   Mus musculus (house mouse) Mus musculus (house mouse) |
-Supramolecule #4: Muscarinic receptor M2R
| Supramolecule | Name: Muscarinic receptor M2R / type: complex / ID: 4 / Parent: 1 / Macromolecule list: #4 |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 1.6 µm / Nominal defocus min: 0.8 µm Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 1.6 µm / Nominal defocus min: 0.8 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Detector mode: COUNTING / Number real images: 17869 / Average electron dose: 50.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT |
|---|
 Movie
Movie Controller
Controller












 Z
Z Y
Y X
X



















