+ Open data
Open data
- Basic information
Basic information
| Entry | 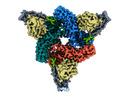 | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of beta-arrestin2 in complex with M2Rpp | |||||||||||||||
 Map data Map data | ||||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords |  GPCR / GPCR /  Arrestin / Arrestin /  SIGNALING PROTEIN / SIGNALING PROTEIN /  SIGNALING PROTEIN-IMMUNE SYSTEM complex SIGNALING PROTEIN-IMMUNE SYSTEM complex | |||||||||||||||
| Function / homology |  Function and homology information Function and homology information angiotensin receptor binding / angiotensin receptor binding /  Muscarinic acetylcholine receptors / symmetric synapse / phospholipase C-activating G protein-coupled acetylcholine receptor signaling pathway / G protein-coupled acetylcholine receptor activity / regulation of smooth muscle contraction / desensitization of G protein-coupled receptor signaling pathway / cholinergic synapse / Muscarinic acetylcholine receptors / symmetric synapse / phospholipase C-activating G protein-coupled acetylcholine receptor signaling pathway / G protein-coupled acetylcholine receptor activity / regulation of smooth muscle contraction / desensitization of G protein-coupled receptor signaling pathway / cholinergic synapse /  inositol hexakisphosphate binding / adenylate cyclase-inhibiting G protein-coupled acetylcholine receptor signaling pathway ... inositol hexakisphosphate binding / adenylate cyclase-inhibiting G protein-coupled acetylcholine receptor signaling pathway ... angiotensin receptor binding / angiotensin receptor binding /  Muscarinic acetylcholine receptors / symmetric synapse / phospholipase C-activating G protein-coupled acetylcholine receptor signaling pathway / G protein-coupled acetylcholine receptor activity / regulation of smooth muscle contraction / desensitization of G protein-coupled receptor signaling pathway / cholinergic synapse / Muscarinic acetylcholine receptors / symmetric synapse / phospholipase C-activating G protein-coupled acetylcholine receptor signaling pathway / G protein-coupled acetylcholine receptor activity / regulation of smooth muscle contraction / desensitization of G protein-coupled receptor signaling pathway / cholinergic synapse /  inositol hexakisphosphate binding / adenylate cyclase-inhibiting G protein-coupled acetylcholine receptor signaling pathway / G protein-coupled serotonin receptor activity / arrestin family protein binding / G protein-coupled receptor internalization / inositol hexakisphosphate binding / adenylate cyclase-inhibiting G protein-coupled acetylcholine receptor signaling pathway / G protein-coupled serotonin receptor activity / arrestin family protein binding / G protein-coupled receptor internalization /  regulation of heart contraction / G protein-coupled receptor signaling pathway, coupled to cyclic nucleotide second messenger / phosphatidylinositol-3,4,5-trisphosphate binding / positive regulation of receptor internalization / endocytic vesicle / asymmetric synapse / axon terminus / regulation of heart contraction / G protein-coupled receptor signaling pathway, coupled to cyclic nucleotide second messenger / phosphatidylinositol-3,4,5-trisphosphate binding / positive regulation of receptor internalization / endocytic vesicle / asymmetric synapse / axon terminus /  clathrin-coated pit / presynaptic modulation of chemical synaptic transmission / clathrin-coated pit / presynaptic modulation of chemical synaptic transmission /  phosphatidylinositol binding / clathrin-coated endocytic vesicle membrane / response to virus / adenylate cyclase-modulating G protein-coupled receptor signaling pathway / G protein-coupled acetylcholine receptor signaling pathway / phosphatidylinositol binding / clathrin-coated endocytic vesicle membrane / response to virus / adenylate cyclase-modulating G protein-coupled receptor signaling pathway / G protein-coupled acetylcholine receptor signaling pathway /  receptor internalization / receptor internalization /  protein transport / Cargo recognition for clathrin-mediated endocytosis / protein transport / Cargo recognition for clathrin-mediated endocytosis /  presynaptic membrane / presynaptic membrane /  Clathrin-mediated endocytosis / Clathrin-mediated endocytosis /  nervous system development / G alpha (i) signalling events / chemical synaptic transmission / nervous system development / G alpha (i) signalling events / chemical synaptic transmission /  postsynaptic membrane / positive regulation of ERK1 and ERK2 cascade / molecular adaptor activity / G protein-coupled receptor signaling pathway / postsynaptic membrane / positive regulation of ERK1 and ERK2 cascade / molecular adaptor activity / G protein-coupled receptor signaling pathway /  dendrite / neuronal cell body / glutamatergic synapse / dendrite / neuronal cell body / glutamatergic synapse /  synapse / synapse /  signal transduction / signal transduction /  membrane / membrane /  nucleus / nucleus /  plasma membrane / plasma membrane /  cytoplasm cytoplasmSimilarity search - Function | |||||||||||||||
| Biological species |   Bos taurus (cattle) / Bos taurus (cattle) /   Mus musculus (house mouse) / Mus musculus (house mouse) /   Homo sapiens (human) Homo sapiens (human) | |||||||||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 2.9 Å cryo EM / Resolution: 2.9 Å | |||||||||||||||
 Authors Authors | Maharana J / Sano FK / Shihoya W / Banerjee R / Nureki O / Shukla AK | |||||||||||||||
| Funding support |  India, 4 items India, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Science / Year: 2024 Journal: Science / Year: 2024Title: Molecular insights into atypical modes of β-arrestin interaction with seven transmembrane receptors. Authors: Jagannath Maharana / Fumiya K Sano / Parishmita Sarma / Manish K Yadav / Longhan Duan / Tomasz M Stepniewski / Madhu Chaturvedi / Ashutosh Ranjan / Vinay Singh / Sayantan Saha / Gargi ...Authors: Jagannath Maharana / Fumiya K Sano / Parishmita Sarma / Manish K Yadav / Longhan Duan / Tomasz M Stepniewski / Madhu Chaturvedi / Ashutosh Ranjan / Vinay Singh / Sayantan Saha / Gargi Mahajan / Mohamed Chami / Wataru Shihoya / Jana Selent / Ka Young Chung / Ramanuj Banerjee / Osamu Nureki / Arun K Shukla /      Abstract: β-arrestins (βarrs) are multifunctional proteins involved in signaling and regulation of seven transmembrane receptors (7TMRs), and their interaction is driven primarily by agonist-induced receptor ...β-arrestins (βarrs) are multifunctional proteins involved in signaling and regulation of seven transmembrane receptors (7TMRs), and their interaction is driven primarily by agonist-induced receptor activation and phosphorylation. Here, we present seven cryo-electron microscopy structures of βarrs either in the basal state, activated by the muscarinic receptor subtype 2 (M2R) through its third intracellular loop, or activated by the βarr-biased decoy D6 receptor (D6R). Combined with biochemical, cellular, and biophysical experiments, these structural snapshots allow the visualization of atypical engagement of βarrs with 7TMRs and also reveal a structural transition in the carboxyl terminus of βarr2 from a β strand to an α helix upon activation by D6R. Our study provides previously unanticipated molecular insights into the structural and functional diversity encoded in 7TMR-βarr complexes with direct implications for exploring novel therapeutic avenues. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_36078.map.gz emd_36078.map.gz | 86.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-36078-v30.xml emd-36078-v30.xml emd-36078.xml emd-36078.xml | 22.4 KB 22.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_36078_fsc.xml emd_36078_fsc.xml | 9.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_36078.png emd_36078.png | 94 KB | ||
| Filedesc metadata |  emd-36078.cif.gz emd-36078.cif.gz | 7.1 KB | ||
| Others |  emd_36078_half_map_1.map.gz emd_36078_half_map_1.map.gz emd_36078_half_map_2.map.gz emd_36078_half_map_2.map.gz | 84.4 MB 84.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-36078 http://ftp.pdbj.org/pub/emdb/structures/EMD-36078 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36078 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36078 | HTTPS FTP |
-Related structure data
| Related structure data |  8j8rMC  8go9C  8j8vC  8j8zC  8j97C  8j9kC  8ja3C  8jafC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_36078.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_36078.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.1989 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_36078_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_36078_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : beta-arrestin2 in complex with M2Rpp
| Entire | Name: beta-arrestin2 in complex with M2Rpp |
|---|---|
| Components |
|
-Supramolecule #1: beta-arrestin2 in complex with M2Rpp
| Supramolecule | Name: beta-arrestin2 in complex with M2Rpp / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|
-Supramolecule #2: beta-arrestin2
| Supramolecule | Name: beta-arrestin2 / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:   Bos taurus (cattle) Bos taurus (cattle) |
-Supramolecule #3: Fab30
| Supramolecule | Name: Fab30 / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2-#3 |
|---|---|
| Source (natural) | Organism:   Mus musculus (house mouse) Mus musculus (house mouse) |
-Supramolecule #4: Muscarinic receptor M2R
| Supramolecule | Name: Muscarinic receptor M2R / type: complex / ID: 4 / Parent: 1 / Macromolecule list: #4 |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) / Synthetically produced: Yes Homo sapiens (human) / Synthetically produced: Yes |
-Macromolecule #1: Beta-arrestin-2
| Macromolecule | Name: Beta-arrestin-2 / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Bos taurus (cattle) Bos taurus (cattle) |
| Molecular weight | Theoretical: 47.217676 KDa |
| Recombinant expression | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Sequence | String: MGEKPGTRVF KKSSPNGKLT VYLGKRDFVD HLDKVDPVDG VVLVDPDYLK DRKVFVTLTV AFRYGREDCD VLGLSFRKDL FIANYQAFP PTPNPPRPPT RLQERLLRKL GQHAHPFFFT IPQNLPSSVT LQPGPEDTGK ALGVDFEIRA FVAKSLEEKS H KRNSVRLV ...String: MGEKPGTRVF KKSSPNGKLT VYLGKRDFVD HLDKVDPVDG VVLVDPDYLK DRKVFVTLTV AFRYGREDCD VLGLSFRKDL FIANYQAFP PTPNPPRPPT RLQERLLRKL GQHAHPFFFT IPQNLPSSVT LQPGPEDTGK ALGVDFEIRA FVAKSLEEKS H KRNSVRLV IRKVQFAPEK PGPQPSAETT RHFLMSDRSL HLEASLDKEL YYHGEPLNVN VHVTNNSTKT VKKIKVSVRQ YA DIVLFST AQYKVPVAQV EQDDQVSPSS TFSKVYTITP FLANNREKRG LALDGKLKHE DTNLASSTIV KEGANKEVLG ILV SYRVKV KLVVSRGGDV SVELPFVLMH PKPHDHIALP RPQSAATHPP TLLPSAVPET DAPVDTNLIE FETNYATDDD IVFE DFARL RLKGLKDEDY DDQFC UniProtKB:  Beta-arrestin-2 Beta-arrestin-2 |
-Macromolecule #2: Fab30 Heavy Chain
| Macromolecule | Name: Fab30 Heavy Chain / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Mus musculus (house mouse) Mus musculus (house mouse) |
| Molecular weight | Theoretical: 25.512354 KDa |
| Recombinant expression | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Sequence | String: EISEVQLVES GGGLVQPGGS LRLSCAASGF NVYSSSIHWV RQAPGKGLEW VASISSYYGY TYYADSVKGR FTISADTSKN TAYLQMNSL RAEDTAVYYC ARSRQFWYSG LDYWGQGTLV TVSSASTKGP SVFPLAPSSK STSGGTAALG CLVKDYFPEP V TVSWNSGA ...String: EISEVQLVES GGGLVQPGGS LRLSCAASGF NVYSSSIHWV RQAPGKGLEW VASISSYYGY TYYADSVKGR FTISADTSKN TAYLQMNSL RAEDTAVYYC ARSRQFWYSG LDYWGQGTLV TVSSASTKGP SVFPLAPSSK STSGGTAALG CLVKDYFPEP V TVSWNSGA LTSGVHTFPA VLQSSGLYSL SSVVTVPSSS LGTQTYICNV NHKPSNTKVD KKVEPKSCDK THHHHHHHH |
-Macromolecule #3: Fab30 Light Chain
| Macromolecule | Name: Fab30 Light Chain / type: protein_or_peptide / ID: 3 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Mus musculus (house mouse) Mus musculus (house mouse) |
| Molecular weight | Theoretical: 23.435064 KDa |
| Recombinant expression | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Sequence | String: SDIQMTQSPS SLSASVGDRV TITCRASQSV SSAVAWYQQK PGKAPKLLIY SASSLYSGVP SRFSGSRSGT DFTLTISSLQ PEDFATYYC QQYKYVPVTF GQGTKVEIKR TVAAPSVFIF PPSDSQLKSG TASVVCLLNN FYPREAKVQW KVDNALQSGN S QESVTEQD ...String: SDIQMTQSPS SLSASVGDRV TITCRASQSV SSAVAWYQQK PGKAPKLLIY SASSLYSGVP SRFSGSRSGT DFTLTISSLQ PEDFATYYC QQYKYVPVTF GQGTKVEIKR TVAAPSVFIF PPSDSQLKSG TASVVCLLNN FYPREAKVQW KVDNALQSGN S QESVTEQD SKDSTYSLSS TLTLSKADYE KHKVYACEVT HQGLSSPVTK SFNRGEC |
-Macromolecule #4: Muscarinic acetylcholine receptor M2
| Macromolecule | Name: Muscarinic acetylcholine receptor M2 / type: protein_or_peptide / ID: 4 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 2.362918 KDa |
| Sequence | String: EI(TPO)QDEN(TPO)V(SEP) (TPO)SLGH(SEP)KD UniProtKB:  Muscarinic acetylcholine receptor M2 Muscarinic acetylcholine receptor M2 |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 Component:
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 1.6 µm / Nominal defocus min: 0.8 µm Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 1.6 µm / Nominal defocus min: 0.8 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Detector mode: COUNTING / Number real images: 2596 / Average electron dose: 56.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller













 Z
Z Y
Y X
X



















