-検索条件
-検索結果
検索 (著者・登録者: edwards & c)の結果376件中、1から50件目までを表示しています

EMDB-40853: 
CH505 Disulfide Stapled SOSIP Bound to b12 Fab
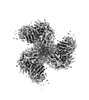
EMDB-40854: 
CH505 Disulfide Stapled SOSIP Bound to CH235.12 Fab

PDB-8sxi: 
CH505 Disulfide Stapled SOSIP Bound to b12 Fab

PDB-8sxj: 
CH505 Disulfide Stapled SOSIP Bound to CH235.12 Fab

EMDB-16453: 
SARS-CoV-2 Omicron Variant Spike Trimer in complex with three 17T2 Fabs

EMDB-16473: 
SARS-CoV-2 spike in complex with the 17T2 neutralizing antibody Fab fragment (local refinement of RBD and Fab)

PDB-8c89: 
SARS-CoV-2 spike in complex with the 17T2 neutralizing antibody Fab fragment (local refinement of RBD and Fab)
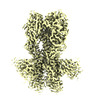
EMDB-28833: 
Cryo-EM structure of X6 COBRA (H1N1) hemagglutinin bound to CR6261 Fab

EMDB-16110: 
Human Urea Transporter UT-A (N-Terminal Domain Model)

EMDB-16111: 
Map of Human Urea Transporter UT-A Collected with 0 and 30 Degree Tilts

EMDB-16112: 
Human Urea Transporter UT-B/UT1 in Complex with Inhibitor UTBinh-14

PDB-8blo: 
Human Urea Transporter UT-A (N-Terminal Domain Model)

PDB-8blp: 
Human Urea Transporter UT-B/UT1 in Complex with Inhibitor UTBinh-14

EMDB-16963: 
Leishmania tarentolae proteasome 20S subunit in complex with 1-Benzyl-N-(3-(cyclopropylcarbamoyl)phenyl)-6-oxo-1,6-dihydropyridazine-3-carboxamide

PDB-8olu: 
Leishmania tarentolae proteasome 20S subunit in complex with 1-Benzyl-N-(3-(cyclopropylcarbamoyl)phenyl)-6-oxo-1,6-dihydropyridazine-3-carboxamide

EMDB-17756: 
Structure of the murine trace amine-associated receptor TAAR7f bound to N,N-dimethylcyclohexylamine (DMCH) in complex with mini-Gs trimeric G protein

PDB-8pm2: 
Structure of the murine trace amine-associated receptor TAAR7f bound to N,N-dimethylcyclohexylamine (DMCH) in complex with mini-Gs trimeric G protein

EMDB-27703: 
Structure of RBD directed antibody DH1047 in complex with SARS-CoV-2 spike: Local refinement of RBD-Fab interace

PDB-8dtk: 
Structure of RBD directed antibody DH1047 in complex with SARS-CoV-2 spike: Local refinement of RBD-Fab interace
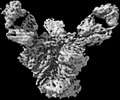
EMDB-27624: 
Cryo-EM structure of HIV-1 Env(CH848 10.17 DS.SOSIP_DT) in complex with DH1030.1 Fab
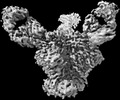
EMDB-27628: 
Cryo-EM structure of HIV-1 Env(BG505.T332N SOSIP) in complex with DH1030.1 Fab

EMDB-27441: 
CryoEM structure of Influenza A virus A/Melbourne/1/1946 (H1N1) hemagglutinin bound to CR6261 Fab

PDB-8sbe: 
Structure of the rat vesicular glutamate transporter 2 determined by single-particle Cryo-EM

EMDB-28162: 
SARS-CoV-2 polyprotein substrate regulates the stepwise Mpro cleavage reaction

EMDB-28200: 
Cryo-EM structure of SARS CoV-2 Mpro WT protease

PDB-8eir: 
SARS-CoV-2 polyprotein substrate regulates the stepwise Mpro cleavage reaction

PDB-8eke: 
Cryo-EM structure of SARS CoV-2 Mpro WT protease

EMDB-28608: 
Cryo-EM structure of CH848 10.17DT DS-SOSIP-2P Env

PDB-8eu8: 
Cryo-EM structure of CH848 10.17DT DS-SOSIP-2P Env

EMDB-27443: 
Cryo-EM structure of influenza A virus A/Bayern/7/1995 hemagglutinin bound to CR6261 Fab
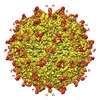
EMDB-25606: 
Zika Virus particle bound with IgM antibody DH1017 Fab fragment

PDB-7t17: 
Zika Virus asymmetric unit bound with IgM antibody DH1017 Fab fragment

EMDB-27440: 
CryoEM structure of Influenza A virus A/Ohio/09/2015 hemagglutinin bound to CR6261 Fab
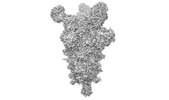
EMDB-26676: 
Three RBD-down state of SARS-CoV-2 D614G spike in complex with the SP1-77 neutralizing antibody Fab fragment

EMDB-26677: 
Three RBD-down state of SARS-CoV-2 D614G spike in complex with the SP1-77 neutralizing antibody Fab fragment (local refinement of the RBD and Fab variable domains)

EMDB-26678: 
An antibody from single human VH-rearranging mouse neutralizes all SARS-CoV-2 variants through BA.5 by inhibiting membrane fusion

EMDB-27043: 
Negative stain electron microscopy single particle reconstruction of monoclonal antibody SP1-77 Fab in complex with SARS-CoV2 2P spike
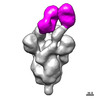
EMDB-27044: 
Negative stain electron microscopy single particle reconstruction of monoclonal antibody VHH7-7-53 Fab in complex with SARS-CoV2 2P spike

EMDB-27046: 
Negative stain electron microscopy single particle reconstruction of monoclonal antibody VHH7-5-82 Fab in complex with SARS-CoV2 2P spike

PDB-7upw: 
Three RBD-down state of SARS-CoV-2 D614G spike in complex with the SP1-77 neutralizing antibody Fab fragment

PDB-7upx: 
Three RBD-down state of SARS-CoV-2 D614G spike in complex with the SP1-77 neutralizing antibody Fab fragment (local refinement of the RBD and Fab variable domains)

PDB-7upy: 
An antibody from single human VH-rearranging mouse neutralizes all SARS-CoV-2 variants through BA.5 by inhibiting membrane fusion

EMDB-26961: 
Triple mutant (K417N-E484K-N501Y) SARS-CoV-2 spike protein in the 3-RBD-Down conformation (S-GSAS-D614G-K417N-E484K-N501Y)

EMDB-11847: 
Organisation of a helical Bunyamwera virus ribonucleoprotein

EMDB-11849: 
Asymmetric reconstruction of a native Bunyamwera virus ribonucleoprotein

PDB-7aoy: 
Helical arrangement of Bunyamwera virus nucleocapsid protein within a native ribonucleoprotein

EMDB-26433: 
SARS-CoV-2 Omicron-BA.2 3-RBD down Spike Protein Trimer without the P986-P987 stabilizing mutations (S-GSAS-Omicron-BA.2)
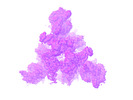
EMDB-26435: 
SARS-CoV-2 Omicron-BA.2 3-RBD down Spike Protein Trimer without the P986-P987 stabilizing mutations (S-GSAS-Omicron-BA.2)

EMDB-26436: 
SARS-CoV-2 Omicron-BA.2 3-RBD down Spike Protein Trimer without the P986-P987 stabilizing mutations (S-GSAS-Omicron-BA.2)

EMDB-26643: 
SARS-CoV-2 Omicron-BA.2 1.5-RBD up Spike Protein Trimer without the P986-P987 stabilizing mutations (S-GSAS-Omicron-BA.2)
ページ:
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア EMN検索について
EMN検索について



 wwPDBはEMDBデータモデルのバージョン3へ移行します
wwPDBはEMDBデータモデルのバージョン3へ移行します
