-検索条件
-検索結果
検索 (著者・登録者: thomas & ja)の結果1,170件中、1から50件目までを表示しています

EMDB-42970: 
Model and map from local refinement of a CAB-A17 - Omicron Ba.1 spike complex

PDB-8v4f: 
Model and map from local refinement of a CAB-A17 - Omicron Ba.1 spike complex

EMDB-17449: 
S. cerevisiae nexus-sCMGE after DNA replication initiation

EMDB-17458: 
S. cerevisiae ssDNA-sCMGE after DNA replication initiation

EMDB-17459: 
S. cerevisiae consensus-sCMGE on ssDNA after DNA replication initiation

EMDB-17460: 
S. cerevisiae sCMGE with N-ter Mcm10 density

PDB-8p5e: 
S. cerevisiae nexus-sCMGE after DNA replication initiation

PDB-8p62: 
S. cerevisiae ssDNA-sCMGE after DNA replication initiation

PDB-8p63: 
S. cerevisiae consensus-sCMGE on ssDNA after DNA replication initiation

EMDB-19066: 
TREK2 in OGNG/CHS detergent micelle with biparatopic inhibitory nanobody Nb6158

EMDB-19854: 
PHF type tau filament from R406W mutant

EMDB-19855: 
PHF type tau filament from in vitro V337M mutant

PDB-9eog: 
PHF type tau filament from R406W mutant

PDB-9eoh: 
PHF type tau filament from in vitro V337M mutant

EMDB-19798: 
human PLD3 homodimer structure

PDB-8s86: 
human PLD3 homodimer structure

EMDB-43008: 
Fab fragment of human mAb #58 in complex with computationally optimized broadly reactive H1 influenza hemagglutinin X6

PDB-8v7o: 
Fab fragment of human mAb #58 in complex with computationally optimized broadly reactive H1 influenza hemagglutinin X6

EMDB-19250: 
Pseudoatomic model of a second-order Sierpinski triangle formed by the citrate synthase from Synechococcus elongatus

EMDB-19251: 
Structure of a first order Sierpinski triangle formed by the H369R mutant of the citrate synthase from Synechococcus elongatus

PDB-8rjk: 
Pseudoatomic model of a second-order Sierpinski triangle formed by the citrate synthase from Synechococcus elongatus

PDB-8rjl: 
Structure of a first order Sierpinski triangle formed by the H369R mutant of the citrate synthase from Synechococcus elongatus

EMDB-16903: 
60S ribosomal subunit bound to the E3-UFM1 complex (native, UFM1 pulldown)

EMDB-16880: 
60S ribosomal subunit bound to the E3-UFM1 complex - state 3 (native)

EMDB-16902: 
60S ribosomal subunit bound to the E3-UFM1 complex - state 2 (native)
EMDB-16905: 
60S ribosomal subunit bound to the E3-UFM1 complex - state 3 (in-vitro reconstitution)

EMDB-16908: 
60S ribosomal subunit bound to the E3-UFM1 complex - state 1 (native)

EMDB-15525: 
Cryo-EM structure of the RecA postsynaptic filament from S. pneumoniae

PDB-8amf: 
Cryo-EM structure of the RecA postsynaptic filament from S. pneumoniae
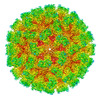

EMDB-43222: 
Cryo-EM structure of Tulane virus 9-6-17 variant capsid protein VP1 5-12-18

EMDB-43292: 
Cryo-EM structure of Tulane virus 9-6-17 variant capsid protein VP1 9-14-18, DTT-treated

EMDB-43293: 
Cryo-EM structure of Tulane virus 9-6-17 variant capsid protein VP1 9-14-18 without DTT treatment

PDB-8vgr: 
Cryo-EM structure of Tulane virus 9-6-17 variant capsid protein VP1 5-12-18

PDB-8vjr: 
Cryo-EM structure of Tulane virus 9-6-17 variant capsid protein VP1 9-14-18, DTT-treated

PDB-8vjs: 
Cryo-EM structure of Tulane virus 9-6-17 variant capsid protein VP1 9-14-18 without DTT treatment

EMDB-43542: 
ELIC5 with cysteamine in 2:1:1 POPC:POPE:POPG nanodisc in open conformation

PDB-8vuw: 
ELIC5 with cysteamine in 2:1:1 POPC:POPE:POPG nanodisc in open conformation

EMDB-41374: 
Antibody N3-1 bound to RBDs in the up and down conformations
EMDB-41382: 
Antibody N3-1 bound to RBD in the up conformation

EMDB-41399: 
Antibody N3-1 bound to SARS-CoV-2 spike

PDB-8tm1: 
Antibody N3-1 bound to RBDs in the up and down conformations

PDB-8tma: 
Antibody N3-1 bound to RBD in the up conformation
EMDB-16424: 
F-actin decorated by SipA497-669
EMDB-16425: 
F-actin decorated by SipA426-685

PDB-8c4c: 
F-actin decorated by SipA497-669

PDB-8c4e: 
F-actin decorated by SipA426-685

EMDB-40856: 
Single particle reconstruction of the human LINE-1 ORF2p without substrate (apo)
EMDB-40858: 
Structure of LINE-1 ORF2p with template:primer hybrid
EMDB-40859: 
Structure of LINE-1 ORF2p with an oligo(A) template

PDB-8sxt: 
Structure of LINE-1 ORF2p with template:primer hybrid
ページ:
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア EMN検索について
EMN検索について



 wwPDBはEMDBデータモデルのバージョン3へ移行します
wwPDBはEMDBデータモデルのバージョン3へ移行します
