-Search query
-Search result
Showing 1 - 50 of 98 items for (author: mah & lee & ng)
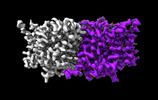
EMDB-29883: 
CLC-ec1 E202Y at pH 4.5 100mM Cl Turn
Method: single particle / : Fortea E, Lee S, Argyos Y, Chadda R, Ciftci D, Huysmans G, Robertson JL, Boudker O, Accardi A
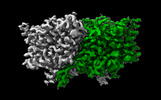
EMDB-29884: 
CLC-ec1 R230C/L249C/C85A at pH 4.5 100mM Cl Swap
Method: single particle / : Fortea E, Lee S, Argyos Y, Chadda R, Ciftci D, Huysmans G, Robertson JL, Boudker O, Accardi A
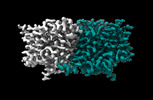
EMDB-29885: 
CLC-ec1 R230C/L249C/C85A at pH 4.5 100mM Cl Turn
Method: single particle / : Fortea E, Lee S, Argyos Y, Chadda R, Ciftci D, Huysmans G, Robertson JL, Boudker O, Accardi A
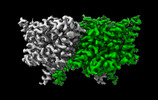
EMDB-29890: 
CLC-ec1 L25C/A450C/C85A at pH 4.5 100mM Cl Intermediate
Method: single particle / : Fortea E, Lee S, Argyos Y, Chadda R, Ciftci D, Huysmans G, Robertson JL, Boudker O, Accardi A
EMDB-29899: 
CLC-ec1 L25C/A450C/C85A at pH 4.5 100mM Cl Twist
Method: single particle / : Fortea E, Lee S, Argyos Y, Chadda R, Ciftci D, Huysmans G, Robertson JL, Boudker O, Accardi A

PDB-8ga0: 
CLC-ec1 E202Y at pH 4.5 100mM Cl Turn
Method: single particle / : Fortea E, Lee S, Argyos Y, Chadda R, Ciftci D, Huysmans G, Robertson JL, Boudker O, Accardi A

PDB-8ga1: 
CLC-ec1 R230C/L249C/C85A at pH 4.5 100mM Cl Swap
Method: single particle / : Fortea E, Lee S, Argyos Y, Chadda R, Ciftci D, Huysmans G, Robertson JL, Boudker O, Accardi A

PDB-8ga3: 
CLC-ec1 R230C/L249C/C85A at pH 4.5 100mM Cl Turn
Method: single particle / : Fortea E, Lee S, Argyos Y, Chadda R, Ciftci D, Huysmans G, Robertson JL, Boudker O, Accardi A

PDB-8ga5: 
CLC-ec1 L25C/A450C/C85A at pH 4.5 100mM Cl Intermediate
Method: single particle / : Fortea E, Lee S, Argyos Y, Chadda R, Ciftci D, Huysmans G, Robertson JL, Boudker O, Accardi A

PDB-8gah: 
CLC-ec1 L25C/A450C/C85A at pH 4.5 100mM Cl Twist
Method: single particle / : Fortea E, Lee S, Argyos Y, Chadda R, Ciftci D, Huysmans G, Robertson JL, Boudker O, Accardi A

EMDB-29980: 
Cryo-EM structure of serine 87 O-GlcNAc-modified alpha-synuclein fibrils
Method: helical / : Balana JA, Nguyen AB, Saelices L, Pratt RM

PDB-8gf7: 
Cryo-EM structure of serine 87 O-GlcNAc-modified alpha-synuclein fibrils
Method: helical / : Balana JA, Nguyen AB, Saelices L, Pratt RM

EMDB-16433: 
Cryo-EM structure of NADH bound SLA dehydrogenase RlGabD from Rhizobium leguminosarum bv. trifolii SRD1565
Method: single particle / : Sharma M, Meek RW, Armstrong Z, Blaza JN, Alhifthi A, Li J, Goddard-Borger ED, Williams SJ, Davies GJ

PDB-8c54: 
Cryo-EM structure of NADH bound SLA dehydrogenase RlGabD from Rhizobium leguminosarum bv. trifolii SRD1565
Method: single particle / : Sharma M, Meek RW, Armstrong Z, Blaza JN, Alhifthi A, Li J, Goddard-Borger ED, Williams SJ, Davies GJ

EMDB-26855: 
Arabidopsis DDM1 bound to nucleosome (H2A.W, H2B, H3.3, H4, with 147 bp DNA)
Method: single particle / : Ipsaro JJ, Adams DW, Joshua-Tor L

PDB-7ux9: 
Arabidopsis DDM1 bound to nucleosome (H2A.W, H2B, H3.3, H4, with 147 bp DNA)
Method: single particle / : Ipsaro JJ, Adams DW, Joshua-Tor L

EMDB-27703: 
Structure of RBD directed antibody DH1047 in complex with SARS-CoV-2 spike: Local refinement of RBD-Fab interace
Method: single particle / : May AJ, Manne K, Acharya P

PDB-8dtk: 
Structure of RBD directed antibody DH1047 in complex with SARS-CoV-2 spike: Local refinement of RBD-Fab interace
Method: single particle / : May AJ, Manne K, Acharya P

EMDB-27943: 
9H2 Fab-poliovirus 1 complex
Method: single particle / : Charnesky AJ
EMDB-27947: 
9H2 Fab-Sabin poliovirus 3 complex
Method: single particle / : Charnesky AJ

EMDB-27948: 
9H2 Fab-poliovirus 2 complex
Method: single particle / : Charnesky AJ
EMDB-27949: 
9H2 Fab-Sabin poliovirus 3 complex
Method: single particle / : Charnesky AJ
EMDB-27950: 
9H2 Fab-Sabin poliovirus 2 complex
Method: single particle / : Charnesky AJ
EMDB-27951: 
9H2 Fab-Sabin poliovirus 1 complex
Method: single particle / : Charnesky AJ

PDB-8e8l: 
9H2 Fab-poliovirus 1 complex
Method: single particle / : Charnesky AJ

PDB-8e8r: 
9H2 Fab-Sabin poliovirus 3 complex
Method: single particle / : Charnesky AJ

PDB-8e8s: 
9H2 Fab-poliovirus 2 complex
Method: single particle / : Charnesky AJ

PDB-8e8x: 
9H2 Fab-Sabin poliovirus 3 complex
Method: single particle / : Charnesky AJ

PDB-8e8y: 
9H2 Fab-Sabin poliovirus 2 complex
Method: single particle / : Charnesky AJ

PDB-8e8z: 
9H2 Fab-Sabin poliovirus 1 complex
Method: single particle / : Charnesky AJ

EMDB-28915: 
SIRT6 bound to an H3K9Ac nucleosome
Method: single particle / : Markert J, Whedon S, Wang Z, Cole P, Farnung L

PDB-8f86: 
SIRT6 bound to an H3K9Ac nucleosome
Method: single particle / : Markert J, Whedon S, Wang Z, Cole P, Farnung L

EMDB-34806: 
SARS-CoV-2 Delta Spike in complex with FP-12A
Method: single particle / : Chen X, Wu YM

EMDB-34807: 
SARS-CoV-2 Delta Spike in complex with IS-9A
Method: single particle / : Mohapatra A, Wu YM

EMDB-34808: 
SARS-CoV-2 Omicron BA.1 Spike in complex with IY-2A
Method: single particle / : Chen X, Mohapatra A, Wu YM

PDB-8hhy: 
SARS-CoV-2 Delta Spike in complex with IS-9A
Method: single particle / : Mohapatra A, Wu YM

PDB-8hhz: 
SARS-CoV-2 Omicron BA.1 Spike in complex with IY-2A
Method: single particle / : Chen X, Mohapatra A, Wu YM

EMDB-25202: 
Cryo-EM structure of Torpedo acetylcholine receptor in apo form
Method: single particle / : Rahman MM, Basta T, Teng J, Lee M, Worrell BT, Stowell MHB, Hibbs RE

EMDB-25205: 
Cryo-EM structure of Torpedo acetylcholine receptor in apo form with added cholesterol
Method: single particle / : Rahman MM, Basta T, Teng J, Lee M, Worrell BT, Stowell MHB, Hibbs RE

EMDB-25206: 
Cryo-EM structure of Torpedo acetylcholine receptor in complex with carbachol, desensitized state
Method: single particle / : Rahman MM, Basta T, Teng J, Lee M, Worrell BT, Stowell MHB, Hibbs RE

EMDB-25207: 
Cryo-EM structure of Torpedo acetylcholine receptor in complex with d-tubocurarine
Method: single particle / : Rahman MM, Basta T, Teng J, Lee M, Worrell BT, Stowell MHB, Hibbs RE

EMDB-25208: 
Cryo-EM structure of Torpedo acetylcholine receptor in complex with d-tubocurarine and carbachol
Method: single particle / : Rahman MM, Basta T, Teng J, Lee M, Worrell BT, Stowell MHB, Hibbs RE

PDB-7smm: 
Cryo-EM structure of Torpedo acetylcholine receptor in apo form
Method: single particle / : Rahman MM, Basta T, Teng J, Lee M, Worrell BT, Stowell MHB, Hibbs RE

PDB-7smq: 
Cryo-EM structure of Torpedo acetylcholine receptor in apo form with added cholesterol
Method: single particle / : Rahman MM, Basta T, Teng J, Lee M, Worrell BT, Stowell MHB, Hibbs RE

PDB-7smr: 
Cryo-EM structure of Torpedo acetylcholine receptor in complex with carbachol, desensitized state
Method: single particle / : Rahman MM, Basta T, Teng J, Lee M, Worrell BT, Stowell MHB, Hibbs RE

PDB-7sms: 
Cryo-EM structure of Torpedo acetylcholine receptor in complex with d-tubocurarine
Method: single particle / : Rahman MM, Basta T, Teng J, Lee M, Worrell BT, Stowell MHB, Hibbs RE

PDB-7smt: 
Cryo-EM structure of Torpedo acetylcholine receptor in complex with d-tubocurarine and carbachol
Method: single particle / : Rahman MM, Basta T, Teng J, Lee M, Worrell BT, Stowell MHB, Hibbs RE

EMDB-30381: 
Cryo-EM density of SARS-CoV-2 spike protein
Method: single particle / : Ho M, Chang Y, Wang C, Wu Y, Huang H, Lee K, Chen T, Lo JM, Chen X, Ma C

EMDB-23400: 
SARS-CoV-2 Spike Protein Trimer bound to DH1043 fab
Method: single particle / : Gobeil S, Acharya P
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search




 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
