[English] 日本語
 Yorodumi
Yorodumi- EMDB-29980: Cryo-EM structure of serine 87 O-GlcNAc-modified alpha-synuclein ... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of serine 87 O-GlcNAc-modified alpha-synuclein fibrils | ||||||||||||||||||||||||
 Map data Map data | main map. This map was used to build model | ||||||||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||||||||
 Keywords Keywords |  alpha-synuclein / alpha-synuclein /  O-GlcNAc / O-GlcNAc /  amyloid / amyloid /  fibril / posttranslational modification / PROTEIN FIBRIL fibril / posttranslational modification / PROTEIN FIBRIL | ||||||||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of mitochondrial electron transport, NADH to ubiquinone / neutral lipid metabolic process /  regulation of phospholipase activity / negative regulation of monooxygenase activity / regulation of acyl-CoA biosynthetic process / negative regulation of dopamine uptake involved in synaptic transmission / negative regulation of norepinephrine uptake / positive regulation of glutathione peroxidase activity / positive regulation of SNARE complex assembly / positive regulation of hydrogen peroxide catabolic process ...negative regulation of mitochondrial electron transport, NADH to ubiquinone / neutral lipid metabolic process / regulation of phospholipase activity / negative regulation of monooxygenase activity / regulation of acyl-CoA biosynthetic process / negative regulation of dopamine uptake involved in synaptic transmission / negative regulation of norepinephrine uptake / positive regulation of glutathione peroxidase activity / positive regulation of SNARE complex assembly / positive regulation of hydrogen peroxide catabolic process ...negative regulation of mitochondrial electron transport, NADH to ubiquinone / neutral lipid metabolic process /  regulation of phospholipase activity / negative regulation of monooxygenase activity / regulation of acyl-CoA biosynthetic process / negative regulation of dopamine uptake involved in synaptic transmission / negative regulation of norepinephrine uptake / positive regulation of glutathione peroxidase activity / positive regulation of SNARE complex assembly / positive regulation of hydrogen peroxide catabolic process / supramolecular fiber / negative regulation of transporter activity / negative regulation of chaperone-mediated autophagy / mitochondrial membrane organization / regulation of reactive oxygen species biosynthetic process / positive regulation of protein localization to cell periphery / regulation of phospholipase activity / negative regulation of monooxygenase activity / regulation of acyl-CoA biosynthetic process / negative regulation of dopamine uptake involved in synaptic transmission / negative regulation of norepinephrine uptake / positive regulation of glutathione peroxidase activity / positive regulation of SNARE complex assembly / positive regulation of hydrogen peroxide catabolic process / supramolecular fiber / negative regulation of transporter activity / negative regulation of chaperone-mediated autophagy / mitochondrial membrane organization / regulation of reactive oxygen species biosynthetic process / positive regulation of protein localization to cell periphery /  regulation of synaptic vesicle recycling / negative regulation of platelet-derived growth factor receptor signaling pathway / negative regulation of exocytosis / regulation of glutamate secretion / response to iron(II) ion / regulation of norepinephrine uptake / dopamine biosynthetic process / regulation of synaptic vesicle recycling / negative regulation of platelet-derived growth factor receptor signaling pathway / negative regulation of exocytosis / regulation of glutamate secretion / response to iron(II) ion / regulation of norepinephrine uptake / dopamine biosynthetic process /  SNARE complex assembly / positive regulation of neurotransmitter secretion / synaptic vesicle priming / dopamine uptake involved in synaptic transmission / SNARE complex assembly / positive regulation of neurotransmitter secretion / synaptic vesicle priming / dopamine uptake involved in synaptic transmission /  regulation of macrophage activation / regulation of locomotion / positive regulation of inositol phosphate biosynthetic process / mitochondrial ATP synthesis coupled electron transport / negative regulation of microtubule polymerization / synaptic vesicle transport / dynein complex binding / positive regulation of receptor recycling / regulation of dopamine secretion / positive regulation of endocytosis / regulation of macrophage activation / regulation of locomotion / positive regulation of inositol phosphate biosynthetic process / mitochondrial ATP synthesis coupled electron transport / negative regulation of microtubule polymerization / synaptic vesicle transport / dynein complex binding / positive regulation of receptor recycling / regulation of dopamine secretion / positive regulation of endocytosis /  protein kinase inhibitor activity / negative regulation of thrombin-activated receptor signaling pathway / response to type II interferon / cuprous ion binding / positive regulation of exocytosis / synaptic vesicle exocytosis / cysteine-type endopeptidase inhibitor activity involved in apoptotic process / response to magnesium ion / protein kinase inhibitor activity / negative regulation of thrombin-activated receptor signaling pathway / response to type II interferon / cuprous ion binding / positive regulation of exocytosis / synaptic vesicle exocytosis / cysteine-type endopeptidase inhibitor activity involved in apoptotic process / response to magnesium ion /  kinesin binding / alpha-tubulin binding / regulation of presynapse assembly / synaptic vesicle endocytosis / negative regulation of serotonin uptake / localization / phospholipid metabolic process / supramolecular fiber organization / axon terminus / kinesin binding / alpha-tubulin binding / regulation of presynapse assembly / synaptic vesicle endocytosis / negative regulation of serotonin uptake / localization / phospholipid metabolic process / supramolecular fiber organization / axon terminus /  inclusion body / cellular response to copper ion / inclusion body / cellular response to copper ion /  Hsp70 protein binding / cellular response to epinephrine stimulus / Hsp70 protein binding / cellular response to epinephrine stimulus /  excitatory postsynaptic potential / response to interleukin-1 / adult locomotory behavior / excitatory postsynaptic potential / response to interleukin-1 / adult locomotory behavior /  SNARE binding / positive regulation of release of sequestered calcium ion into cytosol / fatty acid metabolic process / long-term synaptic potentiation / regulation of transmembrane transporter activity / SNARE binding / positive regulation of release of sequestered calcium ion into cytosol / fatty acid metabolic process / long-term synaptic potentiation / regulation of transmembrane transporter activity /  ferrous iron binding / synapse organization / ferrous iron binding / synapse organization /  phospholipid binding / protein tetramerization / phospholipid binding / protein tetramerization /  phosphoprotein binding / microglial cell activation / regulation of long-term neuronal synaptic plasticity / negative regulation of protein kinase activity / tau protein binding / protein destabilization / PKR-mediated signaling / negative regulation of cysteine-type endopeptidase activity involved in apoptotic process / positive regulation of protein serine/threonine kinase activity / phosphoprotein binding / microglial cell activation / regulation of long-term neuronal synaptic plasticity / negative regulation of protein kinase activity / tau protein binding / protein destabilization / PKR-mediated signaling / negative regulation of cysteine-type endopeptidase activity involved in apoptotic process / positive regulation of protein serine/threonine kinase activity /  receptor internalization / synaptic vesicle membrane / positive regulation of inflammatory response / activation of cysteine-type endopeptidase activity involved in apoptotic process / receptor internalization / synaptic vesicle membrane / positive regulation of inflammatory response / activation of cysteine-type endopeptidase activity involved in apoptotic process /  actin cytoskeleton / positive regulation of peptidyl-serine phosphorylation / cellular response to oxidative stress / actin cytoskeleton / positive regulation of peptidyl-serine phosphorylation / cellular response to oxidative stress /  actin binding / actin binding /  cell cortex / cell cortex /  histone binding / histone binding /  growth cone / chemical synaptic transmission / postsynapse / neuron apoptotic process / negative regulation of neuron apoptotic process / amyloid fibril formation / response to lipopolysaccharide / growth cone / chemical synaptic transmission / postsynapse / neuron apoptotic process / negative regulation of neuron apoptotic process / amyloid fibril formation / response to lipopolysaccharide /  lysosome / molecular adaptor activity / lysosome / molecular adaptor activity /  oxidoreductase activity / transcription cis-regulatory region binding oxidoreductase activity / transcription cis-regulatory region bindingSimilarity search - Function | ||||||||||||||||||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||||||||||||||||||||
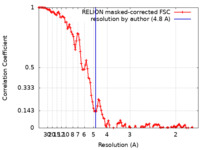
| Method | helical reconstruction /  cryo EM / Resolution: 4.8 Å cryo EM / Resolution: 4.8 Å | ||||||||||||||||||||||||
 Authors Authors | Balana JA / Nguyen AB / Saelices L / Pratt RM | ||||||||||||||||||||||||
| Funding support |  United States, 7 items United States, 7 items
| ||||||||||||||||||||||||
 Citation Citation |  Journal: Nat Chem Biol / Year: 2024 Journal: Nat Chem Biol / Year: 2024Title: O-GlcNAc forces an α-synuclein amyloid strain with notably diminished seeding and pathology. Authors: Aaron T Balana / Anne-Laure Mahul-Mellier / Binh A Nguyen / Mian Horvath / Afraah Javed / Eldon R Hard / Yllza Jasiqi / Preeti Singh / Shumaila Afrin / Rose Pedretti / Virender Singh / ...Authors: Aaron T Balana / Anne-Laure Mahul-Mellier / Binh A Nguyen / Mian Horvath / Afraah Javed / Eldon R Hard / Yllza Jasiqi / Preeti Singh / Shumaila Afrin / Rose Pedretti / Virender Singh / Virginia M-Y Lee / Kelvin C Luk / Lorena Saelices / Hilal A Lashuel / Matthew R Pratt /   Abstract: Amyloid-forming proteins such α-synuclein and tau, which are implicated in Alzheimer's and Parkinson's disease, can form different fibril structures or strains with distinct toxic properties, ...Amyloid-forming proteins such α-synuclein and tau, which are implicated in Alzheimer's and Parkinson's disease, can form different fibril structures or strains with distinct toxic properties, seeding activities and pathology. Understanding the determinants contributing to the formation of different amyloid features could open new avenues for developing disease-specific diagnostics and therapies. Here we report that O-GlcNAc modification of α-synuclein monomers results in the formation of amyloid fibril with distinct core structure, as revealed by cryogenic electron microscopy, and diminished seeding activity in seeding-based neuronal and rodent models of Parkinson's disease. Although the mechanisms underpinning the seeding neutralization activity of the O-GlcNAc-modified fibrils remain unclear, our in vitro mechanistic studies indicate that heat shock proteins interactions with O-GlcNAc fibril inhibit their seeding activity, suggesting that the O-GlcNAc modification may alter the interactome of the α-synuclein fibrils in ways that lead to reduce seeding activity in vivo. Our results show that posttranslational modifications, such as O-GlcNAc modification, of α-synuclein are key determinants of α-synuclein amyloid strains and pathogenicity. | ||||||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_29980.map.gz emd_29980.map.gz | 60.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-29980-v30.xml emd-29980-v30.xml emd-29980.xml emd-29980.xml | 19.9 KB 19.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_29980_fsc.xml emd_29980_fsc.xml | 11.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_29980.png emd_29980.png | 61.3 KB | ||
| Masks |  emd_29980_msk_1.map emd_29980_msk_1.map | 125 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-29980.cif.gz emd-29980.cif.gz | 5.7 KB | ||
| Others |  emd_29980_additional_1.map.gz emd_29980_additional_1.map.gz emd_29980_half_map_1.map.gz emd_29980_half_map_1.map.gz emd_29980_half_map_2.map.gz emd_29980_half_map_2.map.gz | 97.9 MB 98.1 MB 98.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-29980 http://ftp.pdbj.org/pub/emdb/structures/EMD-29980 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29980 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29980 | HTTPS FTP |
-Related structure data
| Related structure data |  8gf7MC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_29980.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_29980.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | main map. This map was used to build model | ||||||||||||||||||||
| Voxel size | X=Y=Z: 0.86 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_29980_msk_1.map emd_29980_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: this map was reconstructed from 3Drefine step in RELION
| File | emd_29980_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | this map was reconstructed from 3Drefine step in RELION | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map 2
| File | emd_29980_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map 1
| File | emd_29980_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : alpha-synuclein fibrils
| Entire | Name: alpha-synuclein fibrils |
|---|---|
| Components |
|
-Supramolecule #1: alpha-synuclein fibrils
| Supramolecule | Name: alpha-synuclein fibrils / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 / Details: The protein is semi-synthesized. |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Alpha-synuclein
| Macromolecule | Name: Alpha-synuclein / type: protein_or_peptide / ID: 1 / Number of copies: 6 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 8.781005 KDa |
| Sequence | String: GLSKAKEGVV AAAEKTKQGV AEAAGKTKEG VLYVGSKTKE GVVHGVATVA EKTKEQVTNV GGAVVTGVTA VAQKTVEGAG SIAAATGFV K UniProtKB:  Alpha-synuclein Alpha-synuclein |
-Macromolecule #2: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 2 / Number of copies: 6 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 / Details: normal PBS |
|---|---|
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 2.2 µm / Nominal defocus min: 0.8 µm Bright-field microscopy / Nominal defocus max: 2.2 µm / Nominal defocus min: 0.8 µm |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number grids imaged: 2 / Number real images: 7270 / Average exposure time: 3.6 sec. / Average electron dose: 40.0 e/Å2 / Details: 3 shots per hole |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller



 Z
Z Y
Y X
X