4U6O
 
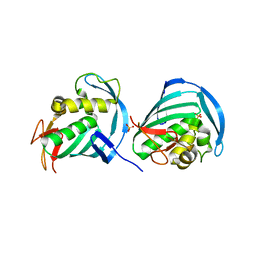 | |
4DOQ
 
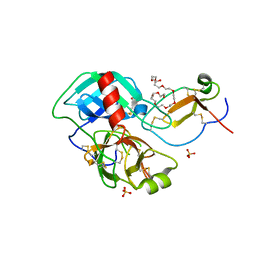 | | Crystal structure of the complex of Porcine Pancreatic Trypsin with 1/2SLPI | | Descriptor: | 3,6,9,12,15,18,21,24,27-NONAOXANONACOSANE-1,29-DIOL, Antileukoproteinase, CALCIUM ION, ... | | Authors: | Fukushima, K, Takimoto-Kamimura, M. | | Deposit date: | 2012-02-10 | | Release date: | 2013-08-14 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure basis 1/2SLPI and porcine pancreas trypsin interaction
J.SYNCHROTRON RADIAT., 20, 2013
|
|
1IYR
 
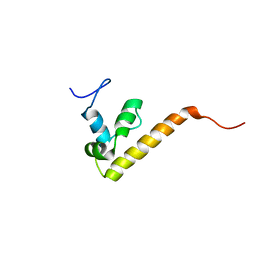 | | NMR Structure Ensemble Of Dff-C Domain | | Descriptor: | DNA FRAGMENTATION FACTOR ALPHA SUBUNIT | | Authors: | Fukushima, K, Kikuchi, J, Koshiba, S, Kigawa, T, Kuroda, Y, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2002-09-05 | | Release date: | 2002-09-25 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of the Dff-C Domain of Dff45/Icad. A Structural Basis for the Regulation of Apoptotic DNA Fragmentation
J.Mol.Biol., 321, 2002
|
|
1KOY
 
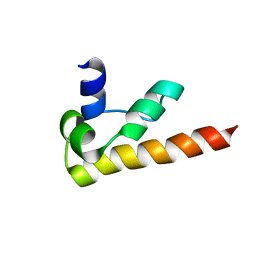 | | NMR structure of DFF-C domain | | Descriptor: | DNA fragmentation factor alpha subunit | | Authors: | Fukushima, K, Kikuchi, J, Koshiba, S, Kigawa, T, Kuroda, Y, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2001-12-25 | | Release date: | 2002-09-04 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the DFF-C domain of DFF45/ICAD. A structural basis for the regulation of apoptotic DNA fragmentation.
J.Mol.Biol., 321, 2002
|
|
5XEJ
 
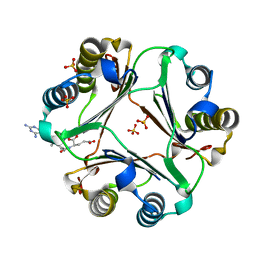 | |
2Z7F
 
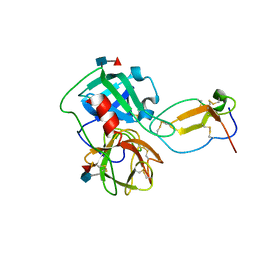 | |
2CZP
 
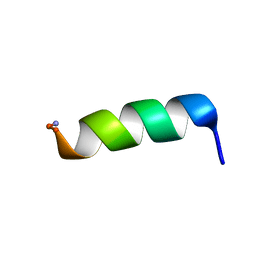 | | Structural analysis of membrane-bound mastoparan-X by solid-state NMR | | Descriptor: | Mastoparan X | | Authors: | Todokoro, Y, Fujiwara, T, Yumen, I, Fukushima, K, Kang, S.-W, Park, J.-S, Kohno, T, Wakamatsu, K, Akutsu, H. | | Deposit date: | 2005-07-14 | | Release date: | 2006-07-04 | | Last modified: | 2022-03-09 | | Method: | SOLID-STATE NMR | | Cite: | Structure of tightly membrane-bound mastoparan-x, a g-protein-activating Peptide, determined by solid-state NMR.
Biophys.J., 91, 2006
|
|
3WBL
 
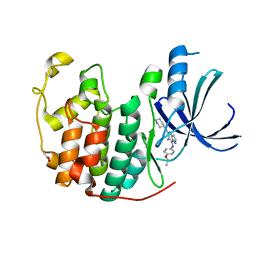 | | Crystal structure of CDK2 in complex with pyrazolopyrimidine inhibitor | | Descriptor: | ACETATE ION, Cyclin-dependent kinase 2, N~7~-(4-ethoxyphenyl)-6-methyl-N~5~-[(3S)-piperidin-3-yl]pyrazolo[1,5-a]pyrimidine-5,7-diamine | | Authors: | Fujino, A, Fukushima, K, Kubota, T, Kosugi, T, Takimoto-Kamimura, M. | | Deposit date: | 2013-05-20 | | Release date: | 2013-10-30 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structure of human cyclin-dependent kinase-2 complex with MK2 inhibitor TEI-I01800: insight into the selectivity.
J.SYNCHROTRON RADIAT., 20, 2013
|
|
3WI6
 
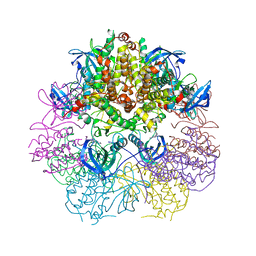 | | Crystal structure of MAPKAP Kinase-2 (MK2) in complex with non-selective inhibitor | | Descriptor: | MAP kinase-activated protein kinase 2, N-[(3S)-piperidin-3-yl]-7,8-dihydro-6H-pyrazolo[1,5-a]pyrrolo[3,2-e]pyrimidin-5-amine | | Authors: | Fujino, A, Fukushima, K, Kubota, T, Matsumoto, Y, Takimoto-Kamimura, M. | | Deposit date: | 2013-09-06 | | Release date: | 2013-12-18 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.99 Å) | | Cite: | Structure of the beta-form of human MK2 in complex with the non-selective kinase inhibitor TEI-L03090
Acta Crystallogr.,Sect.F, 69, 2013
|
|
3A2C
 
 | | Crystal structure of a pyrazolopyrimidine inhibitor complex bound to MAPKAP Kinase-2 (MK2) | | Descriptor: | MAP kinase-activated protein kinase 2, N~7~-(4-ethoxyphenyl)-6-methyl-N~5~-[(3S)-piperidin-3-yl]pyrazolo[1,5-a]pyrimidine-5,7-diamine, SULFATE ION | | Authors: | Fujino, A, Takimoto-Kamimura, M. | | Deposit date: | 2009-05-12 | | Release date: | 2010-05-12 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Structural analysis of an MK2-inhibitor complex: insight into the regulation of the secondary structure of the Gly-rich loop by TEI-I01800
Acta Crystallogr.,Sect.D, 66, 2010
|
|
2DUO
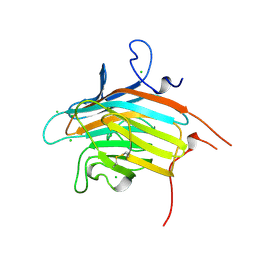 
 | | Crystal structure of VIP36 exoplasmic/lumenal domain, Ca2+-bound form | | Descriptor: | CALCIUM ION, CHLORIDE ION, Vesicular integral-membrane protein VIP36 | | Authors: | Satoh, T, Cowieson, N.P, Kato, R, Wakatsuki, S. | | Deposit date: | 2006-07-25 | | Release date: | 2007-07-24 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural basis for recognition of high mannose type glycoproteins by mammalian transport lectin VIP36
J.Biol.Chem., 282, 2007
|
|
2DUR
 
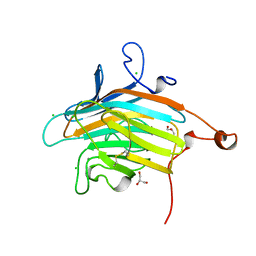 | | Crystal structure of VIP36 exoplasmic/lumenal domain, Ca2+/Man2-bound form | | Descriptor: | CALCIUM ION, CHLORIDE ION, GLYCEROL, ... | | Authors: | Satoh, T, Cowieson, N.P, Kato, R, Wakatsuki, S. | | Deposit date: | 2006-07-25 | | Release date: | 2007-07-24 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Structural basis for recognition of high mannose type glycoproteins by mammalian transport lectin VIP36
J.Biol.Chem., 282, 2007
|
|
2DUP
 
 | | Crystal structure of VIP36 exoplasmic/lumenal domain, metal-free form | | Descriptor: | CALCIUM ION, CHLORIDE ION, GLYCEROL, ... | | Authors: | Satoh, T, Cowieson, N.P, Kato, R, Wakatsuki, S. | | Deposit date: | 2006-07-25 | | Release date: | 2007-07-24 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structural basis for recognition of high mannose type glycoproteins by mammalian transport lectin VIP36
J.Biol.Chem., 282, 2007
|
|
2DUQ
 
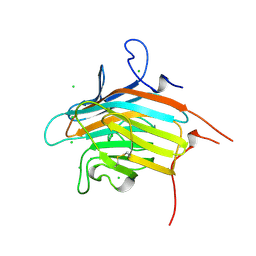 | | Crystal structure of VIP36 exoplasmic/lumenal domain, Ca2+/Man-bound form | | Descriptor: | CALCIUM ION, CHLORIDE ION, Vesicular integral-membrane protein VIP36, ... | | Authors: | Satoh, T, Cowieson, N.P, Kato, R, Wakatsuki, S. | | Deposit date: | 2006-07-25 | | Release date: | 2007-07-24 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural basis for recognition of high mannose type glycoproteins by mammalian transport lectin VIP36
J.Biol.Chem., 282, 2007
|
|
2E6V
 
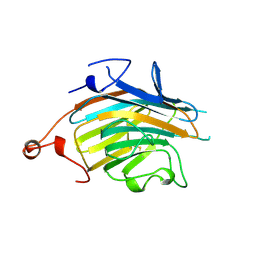 | | Crystal structure of VIP36 exoplasmic/lumenal domain, Ca2+/Man3GlcNAc-bound form | | Descriptor: | CALCIUM ION, Vesicular integral-membrane protein VIP36, alpha-D-mannopyranose, ... | | Authors: | Satoh, T, Kato, R, Wakatsuki, S. | | Deposit date: | 2007-01-04 | | Release date: | 2007-07-24 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural basis for recognition of high mannose type glycoproteins by mammalian transport lectin VIP36
J.Biol.Chem., 282, 2007
|
|
