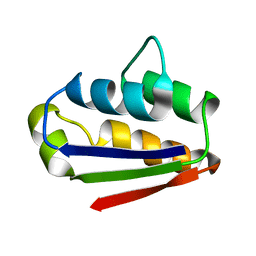
1BTB
 
 | |
2HXX
 
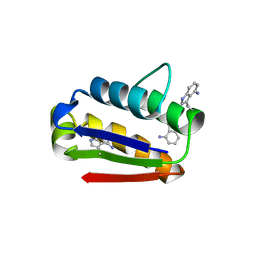 | |
1A19
 
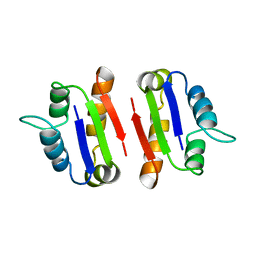 | | BARSTAR (FREE), C82A MUTANT | | Descriptor: | BARSTAR | | Authors: | Ratnaparkhi, G.S, Varadarajan, R. | | Deposit date: | 1997-12-25 | | Release date: | 1998-04-08 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2.76 Å) | | Cite: | Discrepancies between the NMR and X-ray structures of uncomplexed barstar: analysis suggests that packing densities of protein structures determined by NMR are unreliable.
Biochemistry, 37, 1998
|
|
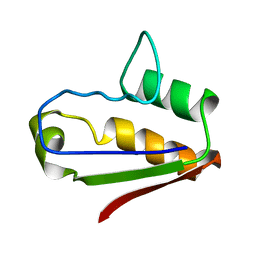
1AB7
 
 | |
1BTA
 
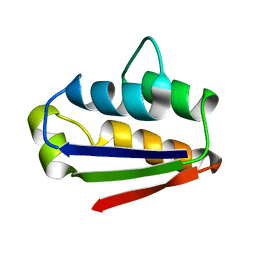 | |
3DA7
 
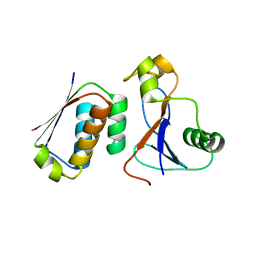 | | A conformationally strained, circular permutant of barnase | | Descriptor: | Barnase circular permutant, Barstar | | Authors: | Mitrousis, G, Butler, J, Loh, S.N, Cingolani, G. | | Deposit date: | 2008-05-28 | | Release date: | 2009-04-14 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Structural and thermodynamic analysis of a conformationally strained circular permutant of barnase.
Biochemistry, 48, 2009
|
|
2ZA4
 
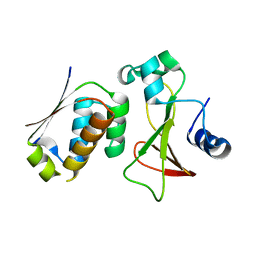 | |
7MRX
 
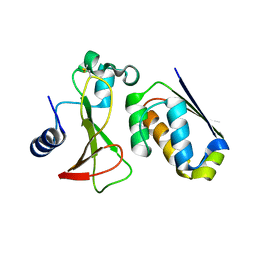 | |
6PQK
 
 | |
1X1Y
 
 | |
1X1U
 
 | |
1X1X
 
 | |
1X1W
 
 | |
1B27
 | |
1AY7
 
 | | RIBONUCLEASE SA COMPLEX WITH BARSTAR | | Descriptor: | BARSTAR, GUANYL-SPECIFIC RIBONUCLEASE SA | | Authors: | Sevcik, J, Urbanikova, L, Dauter, Z, Wilson, K.S. | | Deposit date: | 1997-11-14 | | Release date: | 1999-03-02 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Recognition of RNase Sa by the inhibitor barstar: structure of the complex at 1.7 A resolution.
Acta Crystallogr.,Sect.D, 54, 1998
|
|
1B2S
 | |
1BGS
 
 | | RECOGNITION BETWEEN A BACTERIAL RIBONUCLEASE, BARNASE, AND ITS NATURAL INHIBITOR, BARSTAR | | Descriptor: | BARNASE, BARSTAR | | Authors: | Guillet, V, Lapthorn, A, Mauguen, Y. | | Deposit date: | 1993-11-02 | | Release date: | 1994-04-30 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Recognition between a bacterial ribonuclease, barnase, and its natural inhibitor, barstar.
Structure, 1, 1993
|
|
1B3S
 | |
1B2U
 | |
1BRS
 
 | |
