3MVU
 
 | |
4GN5
 
 | | OBody AM3L15 bound to hen egg-white lysozyme | | Descriptor: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, ACETATE ION, GLYCEROL, ... | | Authors: | Steemson, J.D, Liddament, M.T. | | Deposit date: | 2012-08-16 | | Release date: | 2013-08-21 | | Last modified: | 2014-02-12 | | Method: | X-RAY DIFFRACTION (1.86 Å) | | Cite: | Tracking Molecular Recognition at the Atomic Level with a New Protein Scaffold Based on the OB-Fold.
Plos One, 9, 2014
|
|
4FGJ
 
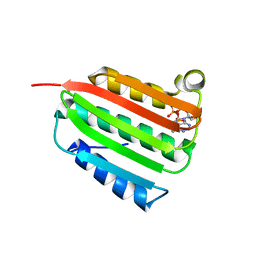
 | | Oxidized quinone reductase 2 in complex with primaquine | | Descriptor: | (4S)-N~4~-(6-methoxyquinolin-8-yl)pentane-1,4-diamine, FLAVIN-ADENINE DINUCLEOTIDE, GLYCEROL, ... | | Authors: | Leung, K.K, Shilton, B.H. | | Deposit date: | 2012-06-04 | | Release date: | 2013-03-13 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.346 Å) | | Cite: | Crystal structures of quinone reductase 2 bound to antimalarial drugs reveal conformational change upon reduction
J.Biol.Chem., 2013
|
|
3MBJ
 
 | |
3MCP
 
 | |
3MCW
 
 | |
4FR9
 
 | |
4FRG
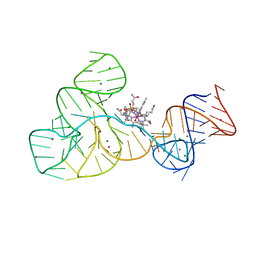 
 | | Crystal structure of the cobalamin riboswitch aptamer domain | | Descriptor: | Hydroxocobalamin, IRIDIUM (III) ION, MAGNESIUM ION, ... | | Authors: | Reyes, F.E, Johnson, J.E, Polaski, J.T, Batey, R.T. | | Deposit date: | 2012-06-26 | | Release date: | 2012-10-17 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.95 Å) | | Cite: | B12 cofactors directly stabilize an mRNA regulatory switch.
Nature, 492, 2012
|
|
4GT8
 | |
3N5L
 
 | |
4H5Q
 
 | |
4HD6
 
 | | Crystal Structure of Tyrosinase from Bacillus megaterium V218F mutant soaked in CuSO4 | | Descriptor: | COPPER (II) ION, Tyrosinase | | Authors: | Goldfeder, M, Kanteev, M, Adir, N, Fishman, A. | | Deposit date: | 2012-10-02 | | Release date: | 2013-01-23 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Influencing the monophenolase/diphenolase activity ratio in tyrosinase.
Biochim.Biophys.Acta, 1834, 2013
|
|
4H6H
 
 | |
4GVP
 
 | |
3NL9
 
 | |
4GOA
 
 | |
4IPB
 
 | |
3MST
 
 | |
4IPM
 
 | | Crystal structure of a GH7 family cellobiohydrolase from Limnoria quadripunctata in complex with thiocellobiose | | Descriptor: | ACETATE ION, CALCIUM ION, GH7 family protein, ... | | Authors: | McGeehan, J.E, Martin, R.N.A, Streeter, S.D, Cragg, S.M, Guille, M.J, Schnorr, K.M, Kern, M, Bruce, N.C, McQueen-Mason, S.J. | | Deposit date: | 2013-01-10 | | Release date: | 2013-06-12 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.14 Å) | | Cite: | Structural characterization of a unique marine animal family 7 cellobiohydrolase suggests a mechanism of cellulase salt tolerance.
Proc.Natl.Acad.Sci.USA, 110, 2013
|
|
4IKL
 
 | |
3M5K
 
 | |
4IN3
 
 | |
3M1U
 
 | |
3M83
 | |
4IPZ
 
 | | SmBz bound to Cyclophilin A | | Descriptor: | CHLORIDE ION, Peptidyl-prolyl cis-trans isomerase A, cyclosporine SmBz-CsA | | Authors: | Price, A.J, Jacques, D.A, James, L.C. | | Deposit date: | 2013-01-10 | | Release date: | 2013-11-06 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.67 Å) | | Cite: | HIV-1 evades innate immune recognition through specific cofactor recruitment.
Nature, 503, 2013
|
|
