13PF
 
 | | PanDDA analysis group deposition -- Crystal structure of PLpro-C111S in complex with Fr12776 | | Descriptor: | Papain-like protease nsp3, ZINC ION, [4-(morpholin-4-yl)phenyl]methanol | | Authors: | Wang, W, Huang, L, Xu, Q, Zhu, Z, Li, M, Zhou, H, Han, T, Zheng, S, Wang, Q, Yu, F. | | Deposit date: | 2025-09-17 | | Release date: | 2026-02-18 | | Method: | X-RAY DIFFRACTION (2.41 Å) | | Cite: | Crystallographic fragment screening discovers novel micromolar active inhibitors and druggable hotspots of SARS-CoV-2 PL pro.
Int.J.Biol.Macromol., 347, 2026
|
|
13PG
 
 | | PanDDA analysis group deposition -- Crystal structure of PLpro-C111S in complex with Fr16736 | | Descriptor: | Papain-like protease nsp3, ZINC ION, cyclobutyl(morpholin-4-yl)methanone | | Authors: | Wang, W, Huang, L, Xu, Q, Zhu, Z, Li, M, Zhou, H, Han, T, Zheng, S, Wang, Q, Yu, F. | | Deposit date: | 2025-09-17 | | Release date: | 2026-02-18 | | Method: | X-RAY DIFFRACTION (2.29 Å) | | Cite: | Crystallographic fragment screening discovers novel micromolar active inhibitors and druggable hotspots of SARS-CoV-2 PL pro.
Int.J.Biol.Macromol., 347, 2026
|
|
13PH
 
 | | PanDDA analysis group deposition -- Crystal structure of PLpro-C111S in complex with Fr12864 | | Descriptor: | 5-chloranylthiophene-2-sulfonamide, Papain-like protease nsp3, ZINC ION | | Authors: | Wang, W, Huang, L, Xu, Q, Zhu, Z, Li, M, Zhou, H, Han, T, Zheng, S, Wang, Q, Yu, F. | | Deposit date: | 2025-09-17 | | Release date: | 2026-02-18 | | Method: | X-RAY DIFFRACTION (2.84 Å) | | Cite: | Crystallographic fragment screening discovers novel micromolar active inhibitors and druggable hotspots of SARS-CoV-2 PL pro.
Int.J.Biol.Macromol., 347, 2026
|
|
13PI
 
 | | PanDDA analysis group deposition -- Crystal structure of PLpro-C111S in complex with Fr12314 | | Descriptor: | 3,4-dihydro-2~{H}-chromene-6-carboxamide, MALONATE ION, Papain-like protease nsp3, ... | | Authors: | Wang, W, Huang, L, Xu, Q, Zhu, Z, Li, M, Zhou, H, Han, T, Zheng, S, Wang, Q, Yu, F. | | Deposit date: | 2025-09-17 | | Release date: | 2026-02-18 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Crystallographic fragment screening discovers novel micromolar active inhibitors and druggable hotspots of SARS-CoV-2 PL pro.
Int.J.Biol.Macromol., 347, 2026
|
|
13PJ
 
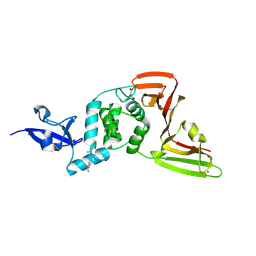 | | PanDDA analysis group deposition -- Crystal structure of PLpro-C111S in complex with Fr13351 | | Descriptor: | 4-[(trifluoromethyl)sulfanyl]benzamide, MALONATE ION, Papain-like protease nsp3, ... | | Authors: | Wang, W, Huang, L, Xu, Q, Zhu, Z, Li, M, Zhou, H, Han, T, Zheng, S, Wang, Q, Yu, F. | | Deposit date: | 2025-09-17 | | Release date: | 2026-02-18 | | Method: | X-RAY DIFFRACTION (2.27 Å) | | Cite: | Crystallographic fragment screening discovers novel micromolar active inhibitors and druggable hotspots of SARS-CoV-2 PL pro.
Int.J.Biol.Macromol., 347, 2026
|
|
13PK
 
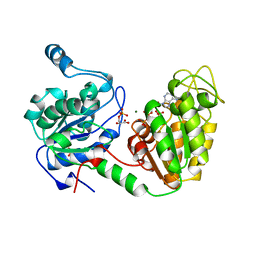 | | TERNARY COMPLEX OF PHOSPHOGLYCERATE KINASE FROM TRYPANOSOMA BRUCEI | | Descriptor: | 3-PHOSPHOGLYCERATE KINASE, 3-PHOSPHOGLYCERIC ACID, ADENOSINE-5'-DIPHOSPHATE, ... | | Authors: | Bernstein, B.E, Michels, P.A.M, Hol, W.G.J. | | Deposit date: | 1996-11-23 | | Release date: | 1997-12-24 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Synergistic effects of substrate-induced conformational changes in phosphoglycerate kinase activation.
Nature, 385, 1997
|
|
13PL
 
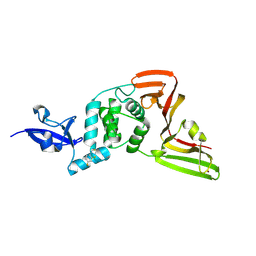 | | PanDDA analysis group deposition -- Crystal structure of PLpro-C111S in complex with Fr12992 | | Descriptor: | N-(2,4-difluorophenyl)hydrazinecarbothioamide, Papain-like protease nsp3, ZINC ION | | Authors: | Wang, W, Huang, L, Xu, Q, Zhu, Z, Li, M, Zhou, H, Han, T, Zheng, S, Wang, Q, Yu, F. | | Deposit date: | 2025-09-17 | | Release date: | 2026-02-18 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Crystallographic fragment screening discovers novel micromolar active inhibitors and druggable hotspots of SARS-CoV-2 PL pro.
Int.J.Biol.Macromol., 347, 2026
|
|
13PM
 
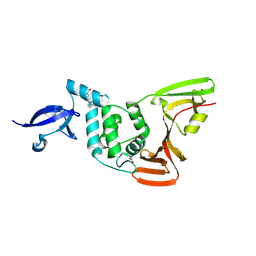 | | PanDDA analysis group deposition -- Crystal structure of PLpro-C111S in complex with Fr12188 | | Descriptor: | MALONATE ION, Papain-like protease nsp3, ZINC ION, ... | | Authors: | Wang, W, Huang, L, Xu, Q, Zhu, Z, Li, M, Zhou, H, Han, T, Zheng, S, Wang, Q, Yu, F. | | Deposit date: | 2025-09-17 | | Release date: | 2026-02-18 | | Method: | X-RAY DIFFRACTION (2.44 Å) | | Cite: | Crystallographic fragment screening discovers novel micromolar active inhibitors and druggable hotspots of SARS-CoV-2 PL pro.
Int.J.Biol.Macromol., 347, 2026
|
|
13PN
 
 | | PanDDA analysis group deposition -- Crystal structure of PLpro-C111S in complex with Fr16619 | | Descriptor: | 1,3-diazepane-2-thione, MALONATE ION, Papain-like protease nsp3, ... | | Authors: | Wang, W, Huang, L, Xu, Q, Zhu, Z, Li, M, Zhou, H, Han, T, Zheng, S, Wang, Q, Yu, F. | | Deposit date: | 2025-09-17 | | Release date: | 2026-02-18 | | Method: | X-RAY DIFFRACTION (2.11 Å) | | Cite: | Crystallographic fragment screening discovers novel micromolar active inhibitors and druggable hotspots of SARS-CoV-2 PL pro.
Int.J.Biol.Macromol., 347, 2026
|
|
13PO
 
 | | PanDDA analysis group deposition -- Crystal structure of PLpro-C111S in complex with 5T-0834 | | Descriptor: | 1-[5-(4-methylpiperazin-1-yl)thiophen-2-yl]ethan-1-one, MALONATE ION, Papain-like protease nsp3, ... | | Authors: | Wang, W, Huang, L, Xu, Q, Zhu, Z, Li, M, Zhou, H, Han, T, Zheng, S, Wang, Q, Yu, F. | | Deposit date: | 2025-09-17 | | Release date: | 2026-02-18 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystallographic fragment screening discovers novel micromolar active inhibitors and druggable hotspots of SARS-CoV-2 PL pro.
Int.J.Biol.Macromol., 347, 2026
|
|
13PP
 
 | | PanDDA analysis group deposition -- Crystal structure of PLpro-C111S in complex with PS-5144 | | Descriptor: | N-methyl-1-[3-(pyridin-3-yl)phenyl]methanamine, Papain-like protease nsp3, ZINC ION | | Authors: | Wang, W, Huang, L, Xu, Q, Zhu, Z, Li, M, Zhou, H, Han, T, Zheng, S, Wang, Q, Yu, F. | | Deposit date: | 2025-09-17 | | Release date: | 2026-02-18 | | Method: | X-RAY DIFFRACTION (2.98 Å) | | Cite: | Crystallographic fragment screening discovers novel micromolar active inhibitors and druggable hotspots of SARS-CoV-2 PL pro.
Int.J.Biol.Macromol., 347, 2026
|
|
13PQ
 
 | | PanDDA analysis group deposition -- Crystal structure of PLpro-C111S in complex with Fr12920 | | Descriptor: | (4-phenoxyphenyl)methanol, Papain-like protease nsp3, ZINC ION | | Authors: | Wang, W, Huang, L, Xu, Q, Zhu, Z, Li, M, Zhou, H, Han, T, Zheng, S, Wang, Q, Yu, F. | | Deposit date: | 2025-09-17 | | Release date: | 2026-02-18 | | Method: | X-RAY DIFFRACTION (2.23 Å) | | Cite: | Crystallographic fragment screening discovers novel micromolar active inhibitors and druggable hotspots of SARS-CoV-2 PL pro.
Int.J.Biol.Macromol., 347, 2026
|
|
13PR
 
 | | PanDDA analysis group deposition -- Crystal structure of PLpro-C111S in complex with Fr12546 | | Descriptor: | 3-(pyridin-2-yloxy)aniline, Papain-like protease nsp3, ZINC ION | | Authors: | Wang, W, Huang, L, Xu, Q, Zhu, Z, Li, M, Zhou, H, Han, T, Zheng, S, Wang, Q, Yu, F. | | Deposit date: | 2025-09-17 | | Release date: | 2026-02-18 | | Method: | X-RAY DIFFRACTION (2.37 Å) | | Cite: | Crystallographic fragment screening discovers novel micromolar active inhibitors and druggable hotspots of SARS-CoV-2 PL pro.
Int.J.Biol.Macromol., 347, 2026
|
|
13PS
 
 | | PanDDA analysis group deposition -- Crystal structure of PLpro-C111S in complex with Fr14256 | | Descriptor: | 3-amino-4-methylbenzamide, Papain-like protease nsp3, ZINC ION | | Authors: | Wang, W, Huang, L, Xu, Q, Zhu, Z, Li, M, Zhou, H, Han, T, Zheng, S, Wang, Q, Yu, F. | | Deposit date: | 2025-09-17 | | Release date: | 2026-02-18 | | Method: | X-RAY DIFFRACTION (2.41 Å) | | Cite: | Crystallographic fragment screening discovers novel micromolar active inhibitors and druggable hotspots of SARS-CoV-2 PL pro.
Int.J.Biol.Macromol., 347, 2026
|
|
13PT
 
 | | PanDDA analysis group deposition -- Crystal structure of PLpro-C111S in complex with Fr13240 | | Descriptor: | Papain-like protease nsp3, ZINC ION, [3-(phenoxymethyl)phenyl]methanol | | Authors: | Wang, W, Huang, L, Xu, Q, Zhu, Z, Li, M, Zhou, H, Han, T, Zheng, S, Wang, Q, Yu, F. | | Deposit date: | 2025-09-17 | | Release date: | 2026-02-18 | | Method: | X-RAY DIFFRACTION (2.64 Å) | | Cite: | Crystallographic fragment screening discovers novel micromolar active inhibitors and druggable hotspots of SARS-CoV-2 PL pro.
Int.J.Biol.Macromol., 347, 2026
|
|
13PU
 
 | | PanDDA analysis group deposition -- Crystal structure of PLpro-C111S in complex with Fr14262 | | Descriptor: | MALONATE ION, N-(2-methylphenyl)-N'-[2-(pyridin-2-yl)ethyl]thiourea, Papain-like protease nsp3, ... | | Authors: | Wang, W, Huang, L, Xu, Q, Zhu, Z, Li, M, Zhou, H, Han, T, Zheng, S, Wang, Q, Yu, F. | | Deposit date: | 2025-09-17 | | Release date: | 2026-02-18 | | Method: | X-RAY DIFFRACTION (2.18 Å) | | Cite: | Crystallographic fragment screening discovers novel micromolar active inhibitors and druggable hotspots of SARS-CoV-2 PL pro.
Int.J.Biol.Macromol., 347, 2026
|
|
13PV
 | | PanDDA analysis group deposition -- Crystal structure of PLpro-C111S in complex with Fr14220 | | Descriptor: | N-(2-chlorophenyl)-N'-[(furan-2-yl)methyl]thiourea, Papain-like protease nsp3, ZINC ION | | Authors: | Wang, W, Huang, L, Xu, Q, Zhu, Z, Li, M, Zhou, H, Han, T, Zheng, S, Wang, Q, Yu, F. | | Deposit date: | 2025-09-17 | | Release date: | 2026-02-18 | | Method: | X-RAY DIFFRACTION (2.17 Å) | | Cite: | Crystallographic fragment screening discovers novel micromolar active inhibitors and druggable hotspots of SARS-CoV-2 PL pro.
Int.J.Biol.Macromol., 347, 2026
|
|
13PW
 
 | | PanDDA analysis group deposition -- Crystal structure of PLpro-C111S in complex with Fr13277 | | Descriptor: | N'-(2,4-difluorophenyl)-N,N-dimethylthiourea, Papain-like protease nsp3, ZINC ION | | Authors: | Wang, W, Huang, L, Xu, Q, Zhu, Z, Li, M, Zhou, H, Han, T, Zheng, S, Wang, Q, Yu, F. | | Deposit date: | 2025-09-17 | | Release date: | 2026-02-18 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Crystallographic fragment screening discovers novel micromolar active inhibitors and druggable hotspots of SARS-CoV-2 PL pro.
Int.J.Biol.Macromol., 347, 2026
|
|
13PX
 
 | | PanDDA analysis group deposition -- Crystal structure of PLpro-C111S in complex with Fr13634 | | Descriptor: | 4-aminobenzamide, Papain-like protease nsp3, ZINC ION | | Authors: | Wang, W, Huang, L, Xu, Q, Zhu, Z, Li, M, Zhou, H, Han, T, Zheng, S, Wang, Q, Yu, F. | | Deposit date: | 2025-09-17 | | Release date: | 2026-02-18 | | Method: | X-RAY DIFFRACTION (2.47 Å) | | Cite: | Crystallographic fragment screening discovers novel micromolar active inhibitors and druggable hotspots of SARS-CoV-2 PL pro.
Int.J.Biol.Macromol., 347, 2026
|
|
13PY
 
 | | PanDDA analysis group deposition -- Crystal structure of PLpro-C111S in complex with Fr14240 | | Descriptor: | 2-chloro-4-(trifluoromethyl)benzene-1-sulfonamide, MALONATE ION, Papain-like protease nsp3, ... | | Authors: | Wang, W, Huang, L, Xu, Q, Zhu, Z, Li, M, Zhou, H, Han, T, Zheng, S, Wang, Q, Yu, F. | | Deposit date: | 2025-09-17 | | Release date: | 2026-02-18 | | Method: | X-RAY DIFFRACTION (2.27 Å) | | Cite: | Crystallographic fragment screening discovers novel micromolar active inhibitors and druggable hotspots of SARS-CoV-2 PL pro.
Int.J.Biol.Macromol., 347, 2026
|
|
13PZ
 
 | | PanDDA analysis group deposition -- Crystal structure of PLpro-C111S in complex with Fr12779 | | Descriptor: | MALONATE ION, Papain-like protease nsp3, ZINC ION, ... | | Authors: | Wang, W, Huang, L, Xu, Q, Zhu, Z, Li, M, Zhou, H, Han, T, Zheng, S, Wang, Q, Yu, F. | | Deposit date: | 2025-09-17 | | Release date: | 2026-02-18 | | Method: | X-RAY DIFFRACTION (2.56 Å) | | Cite: | Crystallographic fragment screening discovers novel micromolar active inhibitors and druggable hotspots of SARS-CoV-2 PL pro.
Int.J.Biol.Macromol., 347, 2026
|
|
13QA
 
 | | PanDDA analysis group deposition -- Crystal structure of PLpro-C111S in complex with Fr12206 | | Descriptor: | 4-(2-methyl-1H-imidazol-1-yl)aniline, MALONATE ION, Papain-like protease nsp3, ... | | Authors: | Wang, W, Huang, L, Xu, Q, Zhu, Z, Li, M, Zhou, H, Han, T, Zheng, S, Wang, Q, Yu, F. | | Deposit date: | 2025-09-17 | | Release date: | 2026-02-18 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystallographic fragment screening discovers novel micromolar active inhibitors and druggable hotspots of SARS-CoV-2 PL pro.
Int.J.Biol.Macromol., 347, 2026
|
|
13QB
 
 | | PanDDA analysis group deposition -- Crystal structure of PLpro-C111S in complex with Fr13881 | | Descriptor: | 3-(trifluoromethyl)-1,4-dihydropyrazol-5-one, Papain-like protease nsp3, ZINC ION | | Authors: | Wang, W, Huang, L, Xu, Q, Zhu, Z, Li, M, Zhou, H, Han, T, Zheng, S, Wang, Q, Yu, F. | | Deposit date: | 2025-09-17 | | Release date: | 2026-02-18 | | Method: | X-RAY DIFFRACTION (2.42 Å) | | Cite: | Crystallographic fragment screening discovers novel micromolar active inhibitors and druggable hotspots of SARS-CoV-2 PL pro.
Int.J.Biol.Macromol., 347, 2026
|
|
13QC
 
 | | PanDDA analysis group deposition -- Crystal structure of PLpro-C111S in complex with FS-2015 | | Descriptor: | 4-methyl-N-[(pyridin-3-yl)methyl]pyridin-2-amine, MALONATE ION, Papain-like protease nsp3, ... | | Authors: | Wang, W, Huang, L, Xu, Q, Zhu, Z, Li, M, Zhou, H, Han, T, Zheng, S, Wang, Q, Yu, F. | | Deposit date: | 2025-09-17 | | Release date: | 2026-02-18 | | Method: | X-RAY DIFFRACTION (2.22 Å) | | Cite: | Crystallographic fragment screening discovers novel micromolar active inhibitors and druggable hotspots of SARS-CoV-2 PL pro.
Int.J.Biol.Macromol., 347, 2026
|
|
13QD
 
 | | PanDDA analysis group deposition -- Crystal structure of PLpro-C111S in complex with Fr13673 | | Descriptor: | 4-methylthiophene-2-carboxamide, MALONATE ION, Papain-like protease nsp3, ... | | Authors: | Wang, W, Huang, L, Xu, Q, Zhu, Z, Li, M, Zhou, H, Han, T, Zheng, S, Wang, Q, Yu, F. | | Deposit date: | 2025-09-17 | | Release date: | 2026-02-18 | | Method: | X-RAY DIFFRACTION (1.92 Å) | | Cite: | Crystallographic fragment screening discovers novel micromolar active inhibitors and druggable hotspots of SARS-CoV-2 PL pro.
Int.J.Biol.Macromol., 347, 2026
|
|
