2Y73
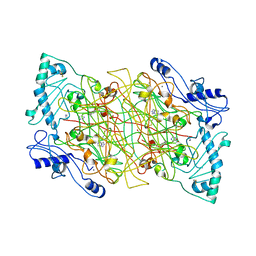 
 | | THE NATIVE STRUCTURES OF SOLUBLE HUMAN PRIMARY AMINE OXIDASE AOC3 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, CALCIUM ION, ... | | Authors: | Elovaara, H, Kidron, H, Parkash, V, Nymalm, Y, Bligt, E, Ollikka, P, Smith, D.J, Pihlavisto, M, Salmi, M, Jalkanen, S, Salminen, T.A. | | Deposit date: | 2011-01-28 | | Release date: | 2011-06-15 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Identification of Two Imidazole Binding Sites and Key Residues for Substrate Specificity in Human Primary Amine Oxidase Aoc3.
Biochemistry, 50, 2011
|
|
7VJT
 
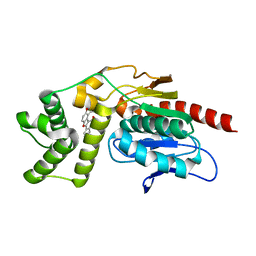 | | Crystal Structure of Mtb Pks13-TE in complex with inhibitor coumestan derivative 8 | | Descriptor: | 3,8-bis(oxidanyl)-7-(piperidin-1-ylmethyl)-[1]benzofuro[3,2-c]chromen-6-one, Polyketide synthase Pks13 (Termination polyketide synthase) | | Authors: | Zhang, W, Wang, S.S, Yu, L.F. | | Deposit date: | 2021-09-28 | | Release date: | 2022-09-28 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.94 Å) | | Cite: | Structure-Based Optimization of Coumestan Derivatives as Polyketide Synthase 13-Thioesterase(Pks13-TE) Inhibitors with Improved hERG Profiles for Mycobacterium tuberculosis Treatment.
J.Med.Chem., 65, 2022
|
|
2LV6
 
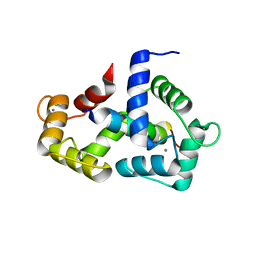 | | The complex between Ca-Calmodulin and skeletal muscle myosin light chain kinase from combination of NMR and aqueous and contrast-matched SAXS data | | Descriptor: | CALCIUM ION, Calmodulin, Myosin light chain kinase 2, ... | | Authors: | Grishaev, A.V, Anthis, N.J, Clore, G.M. | | Deposit date: | 2012-06-29 | | Release date: | 2013-02-20 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR, SOLUTION SCATTERING | | Cite: | Contrast-matched small-angle X-ray scattering from a heavy-atom-labeled protein in structure determination: application to a lead-substituted calmodulin-peptide complex.
J.Am.Chem.Soc., 134, 2012
|
|
7X2U
 
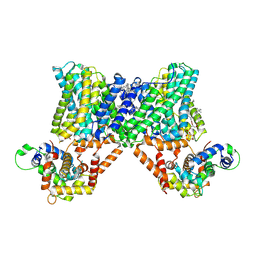 | | Structure of a human NHE3-CHP1 complex in the autoinhibited state | | Descriptor: | (1S)-2-{[{[(2R)-2,3-DIHYDROXYPROPYL]OXY}(HYDROXY)PHOSPHORYL]OXY}-1-[(PALMITOYLOXY)METHYL]ETHYL STEARATE, Calcineurin B homologous protein 1, Sodium/hydrogen exchanger 3, ... | | Authors: | Dong, Y, Li, H, Gao, Y, Zhang, X.C, Zhao, Y. | | Deposit date: | 2022-02-26 | | Release date: | 2022-04-20 | | Last modified: | 2024-06-26 | | Method: | ELECTRON MICROSCOPY (3.2 Å) | | Cite: | Structure of a human NHE3-CHP1 complex in the autoinhibited state
Sci Adv, 2022
|
|
2K0E
 
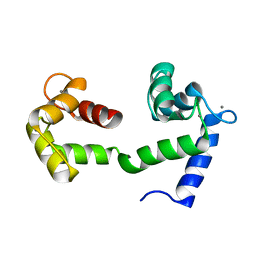 | | A Coupled Equilibrium Shift Mechanism in Calmodulin-Mediated Signal Transduction | | Descriptor: | CALCIUM ION, Calmodulin | | Authors: | Gsponer, J, Christodoulou, J, Cavalli, A, Bui, J.M, Richter, B, Dobson, C.M, Vendruscolo, M. | | Deposit date: | 2008-02-02 | | Release date: | 2008-06-10 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | A coupled equilibrium shift mechanism in calmodulin-mediated signal transduction
Structure, 16, 2008
|
|
7YC4
 
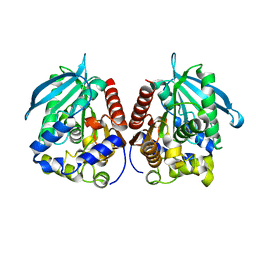 | | Acetylesterase (LgEstI) F207A | | Descriptor: | Alpha/beta hydrolase | | Authors: | Do, H, Lee, J.H. | | Deposit date: | 2022-06-30 | | Release date: | 2023-06-07 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal structure and biochemical analysis of acetylesterase (LgEstI) from Lactococcus garvieae.
Plos One, 18, 2023
|
|
2K61
 
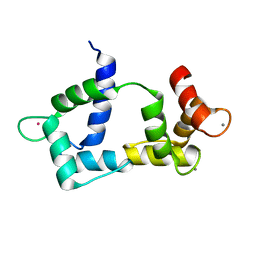 | | Solution structure of CaM complexed to DAPk peptide | | Descriptor: | CALCIUM ION, Calmodulin, TERBIUM(III) ION | | Authors: | Bertini, I, Luchinat, C, Parigi, G, Yuan, J. | | Deposit date: | 2008-07-02 | | Release date: | 2009-05-05 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | Accurate solution structures of proteins from X-ray data and a minimal set of NMR data: calmodulin-peptide complexes as examples.
J.Am.Chem.Soc., 131, 2009
|
|
2JZI
 
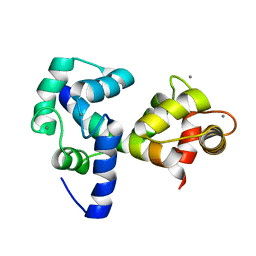 | |
2K0F
 
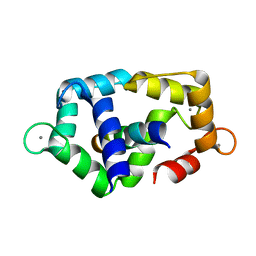 | | Calmodulin complexed with calmodulin-binding peptide from smooth muscle myosin light chain kinase | | Descriptor: | 19-mer peptide from Myosin light chain kinase, CALCIUM ION, calmodulin | | Authors: | Gsponer, J, Christodoulou, J, Cavalli, A, Bui, J.M, Richter, B, Dobson, C.M, Vendruscolo, M. | | Deposit date: | 2008-02-02 | | Release date: | 2008-06-10 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | A coupled equilibrium shift mechanism in calmodulin-mediated signal transduction
Structure, 16, 2008
|
|
2KUG
 
 | | Halothane binds to druggable sites in calcium-calmodulin: Solution Structure of halothane-CaM N-terminal domain | | Descriptor: | 2-BROMO-2-CHLORO-1,1,1-TRIFLUOROETHANE, CALCIUM ION, Calmodulin-1 | | Authors: | Juranic, N, Macura, S, Simeonov, M.V, Jones, K.A, Penheiter, A.R, Hock, T.J, Streiff, J.H. | | Deposit date: | 2010-02-17 | | Release date: | 2010-03-09 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Halothane binds to druggable sites in the [Ca2+]4-calmodulin (CaM) complex, but does not inhibit [Ca2+]4-CaM activation of kinase.
J. Serb. Chem. Soc., 78, 2013
|
|
2N6A
 
 | |
2K0J
 
 | | Solution structure of CaM complexed to DRP1p | | Descriptor: | CALCIUM ION, LANTHANUM (III) ION, calmodulin | | Authors: | Bertini, I, Luchinat, C, Parigi, G, Yuan, J, Structural Proteomics in Europe (SPINE) | | Deposit date: | 2008-02-04 | | Release date: | 2009-03-10 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Accurate solution structures of proteins from X-ray data and a minimal set of NMR data: calmodulin-peptide complexes as examples.
J.Am.Chem.Soc., 131, 2009
|
|
2N27
 
 | | Competitive inhibition of TRPV1 calmodulin interaction by vanilloids | | Descriptor: | (6E)-N-(4-hydroxy-3-methoxybenzyl)-8-methylnon-6-enamide, CALCIUM ION, Calmodulin | | Authors: | Hetenyi, A, Nemeth, L, Weber, E, Szakonyi, G, Winter, Z, Josvay, K, Bartus, E, Olah, Z, Martinek, T.A. | | Deposit date: | 2015-04-29 | | Release date: | 2016-07-06 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Competitive inhibition of TRPV1-calmodulin interaction by vanilloids.
Febs Lett., 590, 2016
|
|
2KNE
 
 | | Calmodulin wraps around its binding domain in the plasma membrane CA2+ pump anchored by a novel 18-1 motif | | Descriptor: | ATPase, Ca++ transporting, plasma membrane 4, ... | | Authors: | Juranic, N, Atanasova, E, Filoteo, A.G, Macura, S, Prendergast, F.G, Penniston, J.T, Strehler, E.E. | | Deposit date: | 2009-08-21 | | Release date: | 2009-11-24 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | Calmodulin wraps around its binding domain in the plasma membrane Ca2+ pump anchored by a novel 18-1 motif.
J.Biol.Chem., 285, 2010
|
|
2Y74
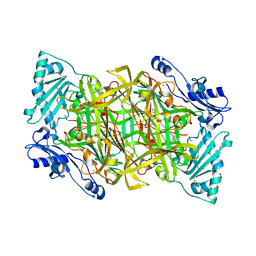 
 | | THE CRYSTAL STRUCTURE OF HUMAN SOLUBLE PRIMARY AMINE OXIDASE AOC3 IN THE OFF-COPPER CONFORMATION | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, CALCIUM ION, ... | | Authors: | Elovaara, H, Kidron, H, Parkash, V, Nymalm, Y, Bligt, E, Ollikka, P, Smith, D.J, Pihlavisto, M, Salmi, M, Jalkanen, S, Salminen, T.A. | | Deposit date: | 2011-01-28 | | Release date: | 2011-06-15 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.95 Å) | | Cite: | Identification of Two Imidazole Binding Sites and Key Residues for Substrate Specificity in Human Primary Amine Oxidase Aoc3.
Biochemistry, 50, 2011
|
|
2ZF8
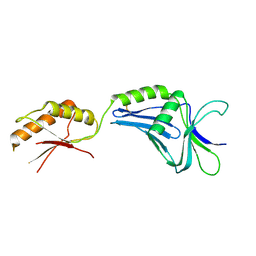 
 | | Crystal structure of MotY | | Descriptor: | Component of sodium-driven polar flagellar motor | | Authors: | Imada, K, Kojima, S, Namba, K, Homma, M. | | Deposit date: | 2007-12-25 | | Release date: | 2008-07-08 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.85 Å) | | Cite: | Insights into the stator assembly of the Vibrio flagellar motor from the crystal structure of MotY
Proc.Natl.Acad.Sci.Usa, 105, 2008
|
|
2MG5
 
 | |
2KBK
 
 | | Solution Structure of BmK-M10 | | Descriptor: | Neurotoxin BmK-M10 | | Authors: | Zhu, J, Wu, H. | | Deposit date: | 2008-11-28 | | Release date: | 2009-12-22 | | Last modified: | 2019-12-11 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of BmK-M10
To be Published
|
|
2WMZ
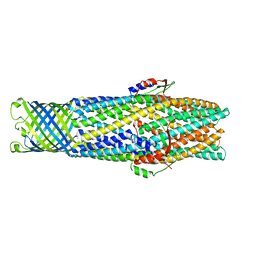 
 | | Structure of a mutated TolC | | Descriptor: | CHLORIDE ION, OUTER MEMBRANE PROTEIN TOLC, SODIUM ION, ... | | Authors: | Hinchliffe, P, Pei, X.Y, Symmons, M.F, Koronakis, E, Hughes, C, Koronakis, V. | | Deposit date: | 2009-07-03 | | Release date: | 2011-01-26 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Structures of Sequential Open States in a Symmetrical Opening Transition of the Tolc Exit Duct.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
1XSW
 
 | | The solid-state NMR structure of Kaliotoxin | | Descriptor: | Kaliotoxin 1 | | Authors: | Lange, A, Becker, S, Seidel, K, Giller, K, Pongs, O, Baldus, M. | | Deposit date: | 2004-10-20 | | Release date: | 2005-04-05 | | Last modified: | 2022-03-02 | | Method: | SOLID-STATE NMR | | Cite: | A Concept for Rapid Protein-Structure Determination by Solid-State NMR Spectroscopy
Angew.Chem.Int.Ed.Engl., 44, 2005
|
|
1YAI
 
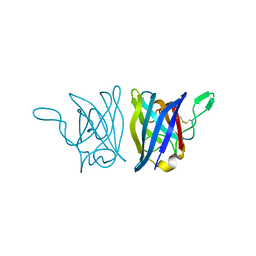 | | X-RAY STRUCTURE OF A BACTERIAL COPPER,ZINC SUPEROXIDE DISMUTASE | | Descriptor: | COPPER (II) ION, COPPER, ZINC SUPEROXIDE DISMUTASE, ... | | Authors: | Bourne, Y, Redford, S.M, Lo, T.P, Tainer, J.A, Getzoff, E.D. | | Deposit date: | 1996-02-03 | | Release date: | 1997-08-20 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Novel dimeric interface and electrostatic recognition in bacterial Cu,Zn superoxide dismutase.
Proc.Natl.Acad.Sci.USA, 93, 1996
|
|
3A9S
 
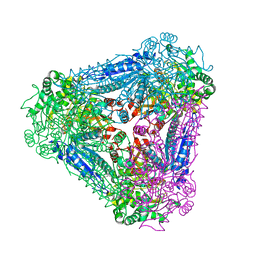 | | X-ray Structure of Bacillus pallidus D-Arabinose Isomerase Complex with Glycerol | | Descriptor: | D-arabinose isomerase, GLYCEROL, MANGANESE (II) ION | | Authors: | Takeda, K, Yoshida, H, Izumori, K, Kamitori, S. | | Deposit date: | 2009-11-05 | | Release date: | 2010-04-07 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | X-ray structures of Bacillus pallidusd-arabinose isomerase and its complex with l-fucitol.
Biochim.Biophys.Acta, 1804, 2010
|
|
3A9R
 
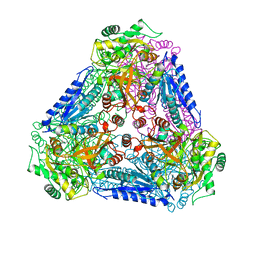 | | X-ray Structures of Bacillus pallidus D-Arabinose IsomeraseComplex with (4R)-2-METHYLPENTANE-2,4-DIOL | | Descriptor: | (4R)-2-METHYLPENTANE-2,4-DIOL, D-arabinose isomerase, MANGANESE (II) ION | | Authors: | Takeda, K, Yoshida, H, Izumori, K, Kamitori, S. | | Deposit date: | 2009-11-05 | | Release date: | 2010-04-07 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.77 Å) | | Cite: | X-ray structures of Bacillus pallidusd-arabinose isomerase and its complex with l-fucitol.
Biochim.Biophys.Acta, 1804, 2010
|
|
3A9T
 
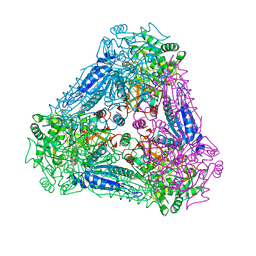 | | X-ray Structure of Bacillus pallidus D-Arabinose Isomerase Complex with L-Fucitol | | Descriptor: | D-arabinose isomerase, FUCITOL, MANGANESE (II) ION | | Authors: | Takeda, K, Yoshida, H, Izumori, K, Kamitori, S. | | Deposit date: | 2009-11-05 | | Release date: | 2010-04-07 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.61 Å) | | Cite: | X-ray structures of Bacillus pallidusd-arabinose isomerase and its complex with l-fucitol.
Biochim.Biophys.Acta, 1804, 2010
|
|
1BF3
 
 | | P-HYDROXYBENZOATE HYDROXYLASE (PHBH) MUTANT WITH CYS 116 REPLACED BY SER (C116S) AND ARG 42 REPLACED BY LYS (R42K), IN COMPLEX WITH FAD AND 4-HYDROXYBENZOIC ACID | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, P-HYDROXYBENZOATE HYDROXYLASE, P-HYDROXYBENZOIC ACID | | Authors: | Eppink, M.H.M, Schreuder, H.A, Van Berkel, W.J.H. | | Deposit date: | 1998-05-26 | | Release date: | 1998-08-12 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Lys42 and Ser42 variants of p-hydroxybenzoate hydroxylase from Pseudomonas fluorescens reveal that Arg42 is essential for NADPH binding.
Eur.J.Biochem., 253, 1998
|
|
