9F7K
 
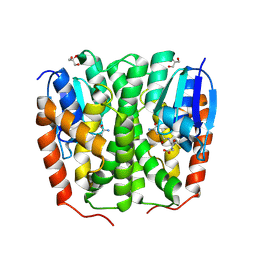 | | Glutathione transferase epsilon 1 from Drosophila melanogaster in complex with glutathione | | Descriptor: | GH14654p, GLUTATHIONE, GLYCEROL, ... | | Authors: | Didierjean, C, Schwartz, M, Neiers, F. | | Deposit date: | 2024-05-03 | | Release date: | 2024-09-04 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural and Thermodynamic Insights into Dimerization Interfaces of Drosophila Glutathione Transferases.
Biomolecules, 14, 2024
|
|
6Z8X
 
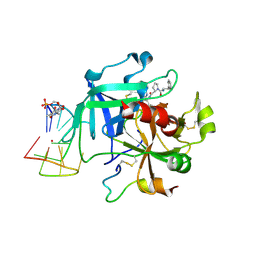 | | X-ray structure of the complex between human alpha thrombin and a thrombin binding aptamer variant (TBA-3Leu), which contains leucyl amide in the side chain of Thy3 at N3. | | Descriptor: | D-phenylalanyl-N-[(2S,3S)-6-{[amino(iminio)methyl]amino}-1-chloro-2-hydroxyhexan-3-yl]-L-prolinamide, POTASSIUM ION, Prothrombin, ... | | Authors: | Troisi, R, Timofeev, E.N, Sica, F. | | Deposit date: | 2020-06-02 | | Release date: | 2021-01-27 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.53 Å) | | Cite: | Expanding the recognition interface of the thrombin-binding aptamer HD1 through modification of residues T3 and T12.
Mol Ther Nucleic Acids, 23, 2021
|
|
2IOH
 
 | | Crystal structure of phosphonoacetaldehyde hydrolase with a K53R mutation | | Descriptor: | MAGNESIUM ION, PHOSPHATE ION, Phosphonoacetaldehyde hydrolase | | Authors: | Allen, K.A, Lahiri, S.D, Zhang, G, Dunaway-Mariano, D, Peisach, E. | | Deposit date: | 2006-10-10 | | Release date: | 2007-08-07 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Diversification of function in the haloacid dehalogenase enzyme superfamily: The role of the cap domain in hydrolytic phosphoruscarbon bond cleavage.
Bioorg.Chem., 34, 2006
|
|
4UX3
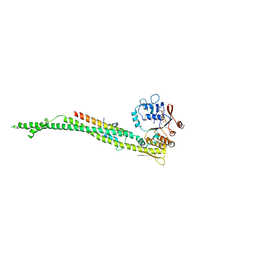 
 | | cohesin Smc3-HD:Scc1-N complex from yeast | | Descriptor: | MAGNESIUM ION, MITOTIC CHROMOSOME DETERMINANT-RELATED PROTEIN, PHOSPHOTHIOPHOSPHORIC ACID-ADENYLATE ESTER, ... | | Authors: | Gligoris, T.G, Nasmyth, K, Lowe, J. | | Deposit date: | 2014-08-18 | | Release date: | 2014-12-03 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | Closing the Cohesin Ring: Structure and Function of its Smc3-Kleisin Interface.
Science, 346, 2014
|
|
3N9B
 
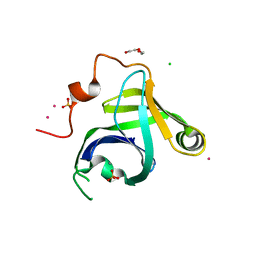 | | Crystal Structure of the P. aeruginosa LigD phosphoesterase domain | | Descriptor: | CHLORIDE ION, DI(HYDROXYETHYL)ETHER, MANGANESE (II) ION, ... | | Authors: | Shuman, S, Nair, P, Smith, P. | | Deposit date: | 2010-05-28 | | Release date: | 2010-08-11 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.92 Å) | | Cite: | Structure of bacterial LigD 3'-phosphoesterase unveils a DNA repair superfamily
Proc.Natl.Acad.Sci.USA, 107, 2010
|
|
6Z8V
 
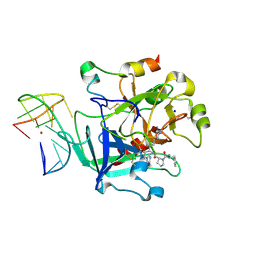 | | X-ray structure of the complex between human alpha thrombin and a thrombin binding aptamer variant (TBA-3L), which contains 1-beta-D-lactopyranosyl residue in the side chain of Thy3 at N3. | | Descriptor: | D-phenylalanyl-N-[(2S,3S)-6-{[amino(iminio)methyl]amino}-1-chloro-2-hydroxyhexan-3-yl]-L-prolinamide, POTASSIUM ION, Prothrombin, ... | | Authors: | Troisi, R, Timofeev, E.N, Sica, F. | | Deposit date: | 2020-06-02 | | Release date: | 2021-01-27 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.58 Å) | | Cite: | Expanding the recognition interface of the thrombin-binding aptamer HD1 through modification of residues T3 and T12.
Mol Ther Nucleic Acids, 23, 2021
|
|
5SDY
 
 | | Crystal Structure of human phosphodiesterase 10 in complex with 6-cyclopropyl-N-[5-(dimethylcarbamoyl)-1-(2-methoxyethyl)pyrazol-4-yl]-3-(pyrimidin-5-ylamino)pyridine-2-carboxamide | | Descriptor: | 6-cyclopropyl-N-[5-(dimethylcarbamoyl)-1-(2-methoxyethyl)-1H-pyrazol-4-yl]-3-[(pyrimidin-5-yl)amino]pyridine-2-carboxamide, MAGNESIUM ION, ZINC ION, ... | | Authors: | Joseph, C, Rodriguez-Sarmiento, R.M, Benz, J, Schlatter, D, Rudolph, M.G. | | Deposit date: | 2022-01-21 | | Release date: | 2022-10-12 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | A high quality, industrial data set for binding affinity prediction: performance comparison in different early drug discovery scenarios.
J.Comput.Aided Mol.Des., 36, 2022
|
|
5SDU
 
 | | Crystal Structure of human phosphodiesterase 10 in complex with 5-methyl-4-[2-(1-methyl-4-phenylimidazol-2-yl)ethyl]-2-phenyl-1,3-oxazole | | Descriptor: | 5-methyl-4-[2-(1-methyl-4-phenyl-1H-imidazol-2-yl)ethyl]-2-phenyl-1,3-oxazole, MAGNESIUM ION, ZINC ION, ... | | Authors: | Joseph, C, Groebke-Zbinden, K, Benz, J, Schlatter, D, Rudolph, M.G. | | Deposit date: | 2022-01-21 | | Release date: | 2022-10-12 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | A high quality, industrial data set for binding affinity prediction: performance comparison in different early drug discovery scenarios.
J.Comput.Aided Mol.Des., 36, 2022
|
|
5SDV
 
 | | Crystal Structure of human phosphodiesterase 10 in complex with 2-methyl-4-N-(1,3-oxazol-4-ylmethyl)-3-N-(2-phenylimidazo[1,2-a]pyrimidin-7-yl)pyrazole-3,4-dicarboxamide | | Descriptor: | 1-methyl-N~4~-[(1,3-oxazol-4-yl)methyl]-N~5~-[(4R)-2-phenylimidazo[1,2-a]pyrimidin-7-yl]-1H-pyrazole-4,5-dicarboxamide, MAGNESIUM ION, ZINC ION, ... | | Authors: | Joseph, C, Peters, J.U, Benz, J, Schlatter, D, Rudolph, M.G. | | Deposit date: | 2022-01-21 | | Release date: | 2022-10-12 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | A high quality, industrial data set for binding affinity prediction: performance comparison in different early drug discovery scenarios.
J.Comput.Aided Mol.Des., 36, 2022
|
|
5SDW
 
 | | Crystal Structure of human phosphodiesterase 10 in complex with 4-N,4-N,2-trimethyl-3-N-(2-phenyl-1H-benzimidazol-5-yl)pyrazole-3,4-dicarboxamide | | Descriptor: | MAGNESIUM ION, N~4~,N~4~,1-trimethyl-N~5~-(2-phenyl-1H-benzimidazol-5-yl)-1H-pyrazole-4,5-dicarboxamide, ZINC ION, ... | | Authors: | Joseph, C, Peters, J.U, Benz, J, Schlatter, D, Rudolph, M.G. | | Deposit date: | 2022-01-21 | | Release date: | 2022-10-12 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | A high quality, industrial data set for binding affinity prediction: performance comparison in different early drug discovery scenarios.
J.Comput.Aided Mol.Des., 36, 2022
|
|
5SE1
 
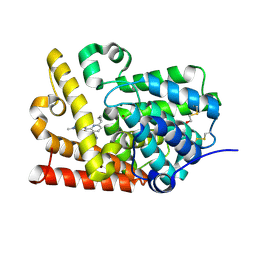 | | Crystal Structure of human phosphodiesterase 10 in complex with 6-cyclopropyl-N-[3-(dimethylcarbamoyl)-1-methylpyrazol-4-yl]-3-(pyrimidin-5-ylamino)pyridine-2-carboxamide | | Descriptor: | 6-cyclopropyl-N-[3-(dimethylcarbamoyl)-1-methyl-1H-pyrazol-4-yl]-3-[(pyrimidin-5-yl)amino]pyridine-2-carboxamide, MAGNESIUM ION, ZINC ION, ... | | Authors: | Joseph, C, Rodriguez-Sarmiento, R.M, Benz, J, Schlatter, D, Rudolph, M.G. | | Deposit date: | 2022-01-21 | | Release date: | 2022-10-12 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | A high quality, industrial data set for binding affinity prediction: performance comparison in different early drug discovery scenarios.
J.Comput.Aided Mol.Des., 36, 2022
|
|
1C08
 
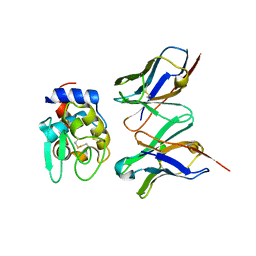 | | CRYSTAL STRUCTURE OF HYHEL-10 FV-HEN LYSOZYME COMPLEX | | Descriptor: | ANTI-HEN EGG WHITE LYSOZYME ANTIBODY (HYHEL-10), LYSOZYME | | Authors: | Shiroishi, M, Kondo, H, Matsushima, M, Tsumoto, K, Kumagai, I. | | Deposit date: | 1999-07-15 | | Release date: | 2000-07-19 | | Last modified: | 2023-05-31 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal structure of anti-Hen egg white lysozyme antibody (HyHEL-10) Fv-antigen complex. Local structural changes in the protein antigen and water-mediated interactions of Fv-antigen and light chain-heavy chain interfaces.
J.Biol.Chem., 274, 1999
|
|
5SE4
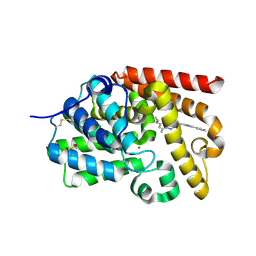 
 | | Crystal Structure of human phosphodiesterase 10 in complex with 5-cyclopropyl-7-methyl-2-[2-(1-methyl-4-phenylimidazol-2-yl)ethyl]-[1,2,4]triazolo[1,5-a]pyrimidine | | Descriptor: | (8S)-5-cyclopropyl-7-methyl-2-[2-(1-methyl-4-phenyl-1H-imidazol-2-yl)ethyl][1,2,4]triazolo[1,5-a]pyrimidine, MAGNESIUM ION, ZINC ION, ... | | Authors: | Joseph, C, Groebke-Zbinden, K, Benz, J, Schlatter, D, Rudolph, M.G. | | Deposit date: | 2022-01-21 | | Release date: | 2022-10-12 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | A high quality, industrial data set for binding affinity prediction: performance comparison in different early drug discovery scenarios.
J.Comput.Aided Mol.Des., 36, 2022
|
|
5SDZ
 
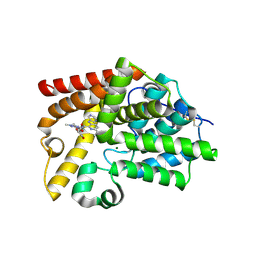 | | Crystal Structure of human phosphodiesterase 10 in complex with 6-cyclopropyl-N-[3-[2,3-dihydroxypropyl(methyl)carbamoyl]-1-methylpyrazol-4-yl]-3-(pyrimidin-5-ylamino)pyridine-2-carboxamide | | Descriptor: | 6-cyclopropyl-N-(3-{[(2S)-2,3-dihydroxypropyl](methyl)carbamoyl}-1-methyl-1H-pyrazol-4-yl)-3-[(pyrimidin-5-yl)amino]pyridine-2-carboxamide, MAGNESIUM ION, ZINC ION, ... | | Authors: | Joseph, C, Groebke-Zbinden, K, Benz, J, Schlatter, D, Rudolph, M.G. | | Deposit date: | 2022-01-21 | | Release date: | 2022-10-12 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | A high quality, industrial data set for binding affinity prediction: performance comparison in different early drug discovery scenarios.
J.Comput.Aided Mol.Des., 36, 2022
|
|
5SEA
 
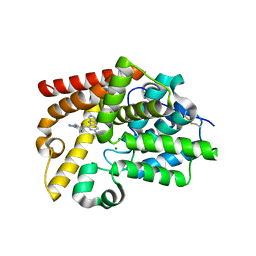 | | Crystal Structure of human phosphodiesterase 10 in complex with 6-cyclopropyl-N-(2-phenylpyrazol-3-yl)-3-(pyrimidin-5-ylamino)pyridine-2-carboxamide | | Descriptor: | 6-cyclopropyl-N-(1-phenyl-1H-pyrazol-5-yl)-3-[(pyrimidin-5-yl)amino]pyridine-2-carboxamide, MAGNESIUM ION, ZINC ION, ... | | Authors: | Joseph, C, Groebke-Zbinden, K, Benz, J, Schlatter, D, Rudolph, M.G. | | Deposit date: | 2022-01-21 | | Release date: | 2022-10-12 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | A high quality, industrial data set for binding affinity prediction: performance comparison in different early drug discovery scenarios.
J.Comput.Aided Mol.Des., 36, 2022
|
|
5SEE
 
 | | Crystal Structure of human phosphodiesterase 10 in complex with 6-cyclopropyl-N-(2-pyridin-2-ylpyrazol-3-yl)-3-(pyrimidin-5-ylamino)pyridine-2-carboxamide | | Descriptor: | 6-cyclopropyl-N-[1-(pyridin-2-yl)-1H-pyrazol-5-yl]-3-[(pyrimidin-5-yl)amino]pyridine-2-carboxamide, CHLORIDE ION, MAGNESIUM ION, ... | | Authors: | Joseph, C, Rodriguez-Sarmiento, R.M, Benz, J, Schlatter, D, Rudolph, M.G. | | Deposit date: | 2022-01-21 | | Release date: | 2022-10-12 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | A high quality, industrial data set for binding affinity prediction: performance comparison in different early drug discovery scenarios.
J.Comput.Aided Mol.Des., 36, 2022
|
|
5SER
 
 | | Crystal Structure of human phosphodiesterase 10 in complex with 4-N-(2-fluoroethyl)-4-N,2-dimethyl-3-N-(2-phenyl-[1,2,4]triazolo[1,5-a]pyridin-7-yl)pyrazole-3,4-dicarboxamide | | Descriptor: | MAGNESIUM ION, N~4~-(2-fluoroethyl)-N~4~,1-dimethyl-N~5~-[(4S)-2-phenyl[1,2,4]triazolo[1,5-a]pyridin-7-yl]-1H-pyrazole-4,5-dicarboxamide, ZINC ION, ... | | Authors: | Joseph, C, Flohr, A, Benz, J, Schlatter, D, Rudolph, M.G. | | Deposit date: | 2022-01-21 | | Release date: | 2022-10-12 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | A high quality, industrial data set for binding affinity prediction: performance comparison in different early drug discovery scenarios.
J.Comput.Aided Mol.Des., 36, 2022
|
|
5SEB
 
 | | Crystal Structure of human phosphodiesterase 10 in complex with 2-methyl-3-[2-(2-methyl-5-pyrrolidin-1-yl-1,2,4-triazol-3-yl)ethyl]quinoxaline | | Descriptor: | 2-methyl-3-{2-[1-methyl-3-(pyrrolidin-1-yl)-1H-1,2,4-triazol-5-yl]ethyl}quinoxaline, MAGNESIUM ION, ZINC ION, ... | | Authors: | Joseph, C, Flohr, A, Benz, J, Schlatter, D, Rudolph, M.G. | | Deposit date: | 2022-01-21 | | Release date: | 2022-10-12 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.28 Å) | | Cite: | A high quality, industrial data set for binding affinity prediction: performance comparison in different early drug discovery scenarios.
J.Comput.Aided Mol.Des., 36, 2022
|
|
5SFA
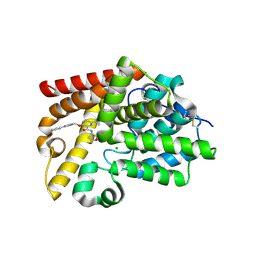 
 | | Crystal Structure of human phosphodiesterase 10 in complex with 6-cyclopropyl-3-[(1-methyl-4-phenylimidazol-2-yl)methoxy]-N-(oxolan-3-yl)pyrazine-2-carboxamide | | Descriptor: | 6-cyclopropyl-3-{[(4S)-1-methyl-4-phenyl-4,5-dihydro-1H-imidazol-2-yl]methoxy}-N-[(3R)-oxolan-3-yl]pyrazine-2-carboxamide, MAGNESIUM ION, ZINC ION, ... | | Authors: | Joseph, C, Koerner, M, Benz, J, Schlatter, D, Rudolph, M.G. | | Deposit date: | 2022-01-21 | | Release date: | 2022-10-12 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.32 Å) | | Cite: | A high quality, industrial data set for binding affinity prediction: performance comparison in different early drug discovery scenarios.
J.Comput.Aided Mol.Des., 36, 2022
|
|
5SED
 
 | | Crystal Structure of human phosphodiesterase 10 in complex with 4-[6-chloro-2-[(1-methyl-4-phenylimidazol-2-yl)methoxy]quinazolin-4-yl]morpholine | | Descriptor: | 6-chloro-2-{[(4S)-1-methyl-4-phenyl-4,5-dihydro-1H-imidazol-2-yl]methoxy}-4-(morpholin-4-yl)quinazoline, MAGNESIUM ION, ZINC ION, ... | | Authors: | Joseph, C, Groebke-Zbinden, K, Benz, J, Schlatter, D, Rudolph, M.G. | | Deposit date: | 2022-01-21 | | Release date: | 2022-10-12 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.32 Å) | | Cite: | A high quality, industrial data set for binding affinity prediction: performance comparison in different early drug discovery scenarios.
J.Comput.Aided Mol.Des., 36, 2022
|
|
5SEC
 
 | | Crystal Structure of human phosphodiesterase 10 in complex with 2-[2-(1,4-diphenylimidazol-2-yl)ethyl]-3-methylquinoxaline | | Descriptor: | 2-[2-(1,4-diphenyl-1H-imidazol-2-yl)ethyl]-3-methylquinoxaline, MAGNESIUM ION, ZINC ION, ... | | Authors: | Joseph, C, Groebke-Zbinden, K, Benz, J, Schlatter, D, Rudolph, M.G. | | Deposit date: | 2022-01-21 | | Release date: | 2022-10-12 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.03 Å) | | Cite: | A high quality, industrial data set for binding affinity prediction: performance comparison in different early drug discovery scenarios.
J.Comput.Aided Mol.Des., 36, 2022
|
|
5SFR
 
 | | Crystal Structure of human phosphodiesterase 10 in complex with 4-N-ethyl-3-N-[2-[3-(2-fluoroethoxy)phenyl]imidazo[1,2-a]pyrimidin-7-yl]-4-N,2-dimethylpyrazole-3,4-dicarboxamide | | Descriptor: | MAGNESIUM ION, N~4~-ethyl-N~5~-{(4S)-2-[3-(2-fluoroethoxy)phenyl]imidazo[1,2-a]pyrimidin-7-yl}-N~4~,1-dimethyl-1H-pyrazole-4,5-dicarboxamide, ZINC ION, ... | | Authors: | Joseph, C, Gobbi, L, Benz, J, Schlatter, D, Rudolph, M.G. | | Deposit date: | 2022-01-21 | | Release date: | 2022-10-12 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.04 Å) | | Cite: | A high quality, industrial data set for binding affinity prediction: performance comparison in different early drug discovery scenarios.
J.Comput.Aided Mol.Des., 36, 2022
|
|
5SEH
 
 | | Crystal Structure of human phosphodiesterase 10 in complex with 4-(azetidine-1-carbonyl)-2-methyl-N-(2-morpholin-4-yl-[1,2,4]triazolo[1,5-a]pyridin-7-yl)pyrazole-3-carboxamide | | Descriptor: | 4-(azetidine-1-carbonyl)-1-methyl-N-[(4S)-2-(morpholin-4-yl)[1,2,4]triazolo[1,5-a]pyridin-7-yl]-1H-pyrazole-5-carboxamide, MAGNESIUM ION, ZINC ION, ... | | Authors: | Joseph, C, Flohr, A, Benz, J, Schlatter, D, Rudolph, M.G. | | Deposit date: | 2022-01-21 | | Release date: | 2022-10-12 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | A high quality, industrial data set for binding affinity prediction: performance comparison in different early drug discovery scenarios.
J.Comput.Aided Mol.Des., 36, 2022
|
|
5SEJ
 
 | | Crystal Structure of human phosphodiesterase 10 in complex with 6-chloro-2-[2-(2-methyl-5-pyrrolidin-1-yl-1,2,4-triazol-3-yl)ethyl]-[1,2,4]triazolo[1,5-a]pyridine | | Descriptor: | (4R)-6-chloro-2-{2-[1-methyl-3-(pyrrolidin-1-yl)-1H-1,2,4-triazol-5-yl]ethyl}[1,2,4]triazolo[1,5-a]pyridine, MAGNESIUM ION, ZINC ION, ... | | Authors: | Joseph, C, Lerner, C, Benz, J, Schlatter, D, Rudolph, M.G. | | Deposit date: | 2022-01-21 | | Release date: | 2022-10-12 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.44 Å) | | Cite: | A high quality, industrial data set for binding affinity prediction: performance comparison in different early drug discovery scenarios.
J.Comput.Aided Mol.Des., 36, 2022
|
|
5SEQ
 
 | | Crystal Structure of human phosphodiesterase 10 in complex with 5,6,8-trimethyl-2-[2-(2-methyl-5-pyrrolidin-1-yl-1,2,4-triazol-3-yl)ethyl]-[1,2,4]triazolo[1,5-a]pyrazine | | Descriptor: | (4S)-5,6,8-trimethyl-2-{2-[1-methyl-3-(pyrrolidin-1-yl)-1H-1,2,4-triazol-5-yl]ethyl}[1,2,4]triazolo[1,5-a]pyrazine, MAGNESIUM ION, ZINC ION, ... | | Authors: | Joseph, C, Lerner, C, Benz, J, Schlatter, D, Rudolph, M.G. | | Deposit date: | 2022-01-21 | | Release date: | 2022-10-12 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.18 Å) | | Cite: | A high quality, industrial data set for binding affinity prediction: performance comparison in different early drug discovery scenarios.
J.Comput.Aided Mol.Des., 36, 2022
|
|
