1XN4
 
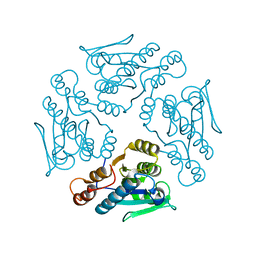 | |
3OL3
 
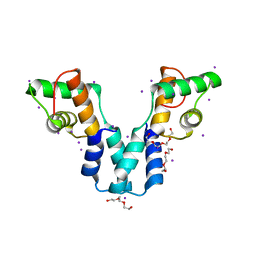 | |
3OL4
 
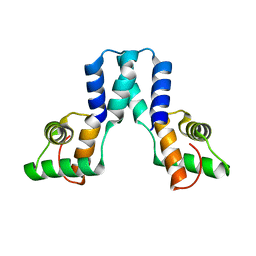 | |
5XKY
 
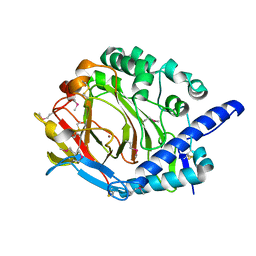 | | Crystal structure of DddY Se derivative | | Descriptor: | Uncharacterized protein, ZINC ION | | Authors: | Li, C.Y, Zhang, Y.Z. | | Deposit date: | 2017-05-10 | | Release date: | 2017-11-01 | | Last modified: | 2018-01-24 | | Method: | X-RAY DIFFRACTION (2.303 Å) | | Cite: | Mechanistic Insights into Dimethylsulfoniopropionate Lyase DddY, a New Member of the Cupin Superfamily.
J. Mol. Biol., 429, 2017
|
|
1YZV
 
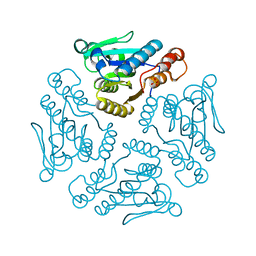 | |
3JZU
 
 | | Crystal structure of Dipeptide Epimerase from Enterococcus faecalis V583 complexed with Mg and dipeptide L-Leu-L-Tyr | | Descriptor: | Dipeptide Epimerase, LEUCINE, MAGNESIUM ION, ... | | Authors: | Fedorov, A.A, Fedorov, E.V, Imker, H.J, Sakai, A, Gerlt, J.A, Almo, S.C. | | Deposit date: | 2009-09-24 | | Release date: | 2010-08-18 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Homology models guide discovery of diverse enzyme specificities among dipeptide epimerases in the enolase superfamily.
Proc.Natl.Acad.Sci.USA, 109, 2012
|
|
4DYY
 
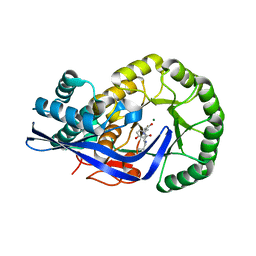
 | | Crystal Structure of the Cu-adduct of Human H-Ferritin variant MIC1 | | Descriptor: | 1,2-ETHANEDIOL, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, CALCIUM ION, ... | | Authors: | Tezcan, F.A, Huard, D.J.E. | | Deposit date: | 2012-02-29 | | Release date: | 2013-01-23 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Re-engineering protein interfaces yields copper-inducible ferritin cage assembly.
Nat.Chem.Biol., 9, 2013
|
|
4V2T
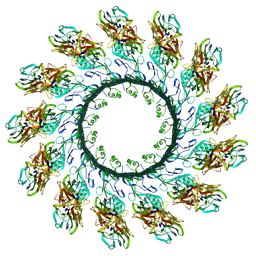 
 | | Membrane embedded pleurotolysin pore with 13 fold symmetry | | Descriptor: | PLEUROTOLYSIN A, PLEUROTOLYSIN B | | Authors: | Lukoyanova, N, Kondos, S.C, Farabella, I, Law, R.H.P, Reboul, C.F, Caradoc-Davies, T.T, Spicer, B.A, Kleifeld, O, Perugini, M, Ekkel, S, Hatfaludi, T, Oliver, K, Hotze, E.M, Tweten, R.K, Whisstock, J.C, Topf, M, Dunstone, M.A, Saibil, H.R. | | Deposit date: | 2014-10-15 | | Release date: | 2015-02-18 | | Last modified: | 2024-05-08 | | Method: | ELECTRON MICROSCOPY (11 Å) | | Cite: | Conformational Changes During Pore Formation by the Perforin-Related Protein Pleurotolysin.
Plos Biol., 13, 2015
|
|
6MQA
 
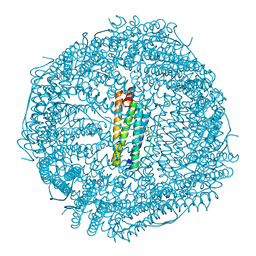
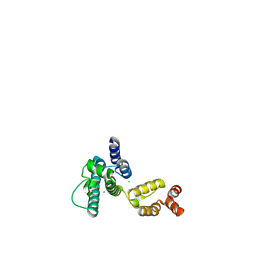 | | Structure of HIV-1 CA P207S | | Descriptor: | CHLORIDE ION, Capsid protein, IODIDE ION | | Authors: | Smaga, S.S, Xiong, Y. | | Deposit date: | 2018-10-09 | | Release date: | 2019-06-19 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (3.199 Å) | | Cite: | MxB Restricts HIV-1 by Targeting the Tri-hexamer Interface of the Viral Capsid.
Structure, 27, 2019
|
|
6MQP
 
 | | Structure of HIV-1 CA T210K | | Descriptor: | CHLORIDE ION, Capsid protein, IODIDE ION | | Authors: | Smaga, S.S, Xiong, Y. | | Deposit date: | 2018-10-10 | | Release date: | 2019-06-19 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (3.296 Å) | | Cite: | MxB Restricts HIV-1 by Targeting the Tri-hexamer Interface of the Viral Capsid.
Structure, 27, 2019
|
|
6FB4
 
 | | human KIBRA C2 domain mutant C771A | | Descriptor: | GLYCEROL, PHOSPHATE ION, Protein KIBRA | | Authors: | Crennell, S.J, Posner, M.G, Bagby, S. | | Deposit date: | 2017-12-18 | | Release date: | 2018-05-16 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.415631 Å) | | Cite: | Distinctive phosphoinositide- and Ca2+-binding properties of normal and cognitive performance-linked variant forms of KIBRA C2 domain.
J. Biol. Chem., 293, 2018
|
|
7SCC
 
 | | T-cylinder of Synechocystis PCC 6803 Phycobilisome, complex with OCP - local refinement | | Descriptor: | Allophycocyanin alpha chain, Allophycocyanin beta chain, Orange carotenoid-binding protein, ... | | Authors: | Sauer, P.V, Sutter, M, Dominguez-Martin, M.A, Kirst, H, Kerfeld, C.A. | | Deposit date: | 2021-09-27 | | Release date: | 2022-08-31 | | Last modified: | 2022-10-05 | | Method: | ELECTRON MICROSCOPY (2.6 Å) | | Cite: | Structures of a phycobilisome in light-harvesting and photoprotected states.
Nature, 609, 2022
|
|
6FJC
 
 | | Human KIBRA C2 domain mutant C771A in complex with phosphatidylinositol 3,4,5-trisphosphate | | Descriptor: | (2R)-3-{[(S)-{[(2S,3R,5S,6S)-2,6-DIHYDROXY-3,4,5-TRIS(PHOSPHONOOXY)CYCLOHEXYL]OXY}(HYDROXY)PHOSPHORYL]OXY}-2-(1-HYDROXY BUTOXY)PROPYL BUTYRATE, GLYCEROL, Protein KIBRA, ... | | Authors: | Crennell, S.J, Posner, M.G, Bagby, S. | | Deposit date: | 2018-01-22 | | Release date: | 2018-05-16 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.598 Å) | | Cite: | Distinctive phosphoinositide- and Ca2+-binding properties of normal and cognitive performance-linked variant forms of KIBRA C2 domain.
J. Biol. Chem., 293, 2018
|
|
7SCB
 
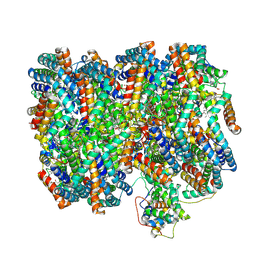 | | B-cylinder of Synechocystis PCC 6803 Phycobilisome, complex with OCP - local refinement | | Descriptor: | Allophycocyanin alpha chain, Allophycocyanin beta chain, Allophycocyanin subunit alpha-B, ... | | Authors: | Sauer, P.V, Sutter, M, Dominguez-Martin, M.A, Kirst, H, Kerfeld, C.A. | | Deposit date: | 2021-09-27 | | Release date: | 2022-08-31 | | Last modified: | 2022-10-05 | | Method: | ELECTRON MICROSCOPY (2.5 Å) | | Cite: | Structures of a phycobilisome in light-harvesting and photoprotected states.
Nature, 609, 2022
|
|
7SCA
 
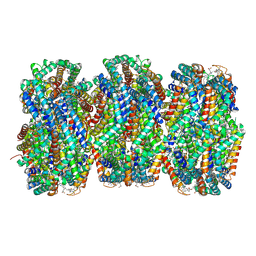 | | Synechocystis PCC 6803 Phycobilisome rod from OCP-PBS complex sample | | Descriptor: | C-phycocyanin alpha subunit, C-phycocyanin beta subunit, PHYCOCYANOBILIN, ... | | Authors: | Sauer, P.V, Sutter, M, Dominguez-Martin, M.A, Kirst, H, Kerfeld, C.A. | | Deposit date: | 2021-09-27 | | Release date: | 2022-08-31 | | Last modified: | 2022-10-05 | | Method: | ELECTRON MICROSCOPY (2.1 Å) | | Cite: | Structures of a phycobilisome in light-harvesting and photoprotected states.
Nature, 609, 2022
|
|
7SC8
 
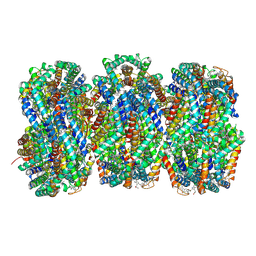 | | Synechocystis PCC 6803 Phycobilisome rod from PBS sample | | Descriptor: | C-phycocyanin alpha subunit, C-phycocyanin beta subunit, PHYCOCYANOBILIN, ... | | Authors: | Sauer, P.V, Sutter, M, Dominguez-Martin, M.A, Kirst, H, Kerfeld, C.A. | | Deposit date: | 2021-09-27 | | Release date: | 2022-08-31 | | Last modified: | 2022-10-05 | | Method: | ELECTRON MICROSCOPY (2.1 Å) | | Cite: | Structures of a phycobilisome in light-harvesting and photoprotected states.
Nature, 609, 2022
|
|
7SC7
 
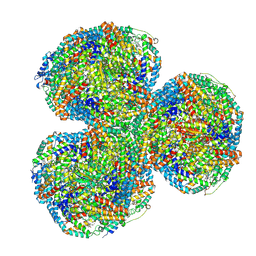 | | Synechocystis PCC 6803 Phycobilisome core from up-down rod conformation | | Descriptor: | Allophycocyanin alpha chain, Allophycocyanin beta chain, Allophycocyanin subunit alpha-B, ... | | Authors: | Sauer, P.V, Sutter, M, Dominguez-Martin, M.A, Kirst, H, Kerfeld, C.A. | | Deposit date: | 2021-09-27 | | Release date: | 2022-08-31 | | Last modified: | 2022-10-05 | | Method: | ELECTRON MICROSCOPY (2.8 Å) | | Cite: | Structures of a phycobilisome in light-harvesting and photoprotected states.
Nature, 609, 2022
|
|
6MQO
 
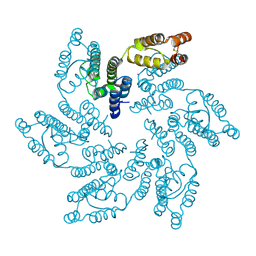 | | Structure of HIV-1 CA G208R | | Descriptor: | Capsid protein, IODIDE ION | | Authors: | Smaga, S.S, Xiong, Y. | | Deposit date: | 2018-10-10 | | Release date: | 2019-06-19 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | MxB Restricts HIV-1 by Targeting the Tri-hexamer Interface of the Viral Capsid.
Structure, 27, 2019
|
|
7SC9
 
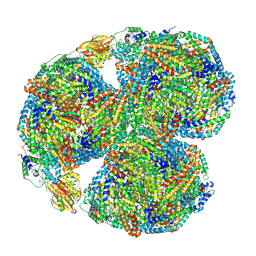 | | Synechocystis PCC 6803 Phycobilisome core, complex with OCP | | Descriptor: | Allophycocyanin alpha chain, Allophycocyanin beta chain, Allophycocyanin subunit alpha-B, ... | | Authors: | Sauer, P.V, Sutter, M, Dominguez-Martin, M.A, Kirst, H, Kerfeld, C.A. | | Deposit date: | 2021-09-27 | | Release date: | 2022-08-31 | | Last modified: | 2022-10-05 | | Method: | ELECTRON MICROSCOPY (2.6 Å) | | Cite: | Structures of a phycobilisome in light-harvesting and photoprotected states.
Nature, 609, 2022
|
|
6FJD
 
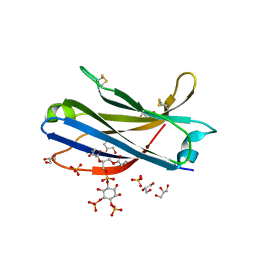 | | Human KIBRA C2 domain mutant C771A in complex with phosphatidylinositol 4,5-bisphosphate | | Descriptor: | (2R)-3-{[(R)-HYDROXY{[(1R,2R,3S,4R,5R,6S)-2,3,6-TRIHYDROXY-4,5-BIS(PHOSPHONOOXY)CYCLOHEXYL]OXY}PHOSPHORYL]OXY}PROPANE-1 ,2-DIYL DIBUTANOATE, GLYCEROL, Protein KIBRA, ... | | Authors: | Crennell, S.J, Posner, M.G, Bagby, S. | | Deposit date: | 2018-01-22 | | Release date: | 2018-05-16 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.898 Å) | | Cite: | Distinctive phosphoinositide- and Ca2+-binding properties of normal and cognitive performance-linked variant forms of KIBRA C2 domain.
J. Biol. Chem., 293, 2018
|
|
6FD0
 
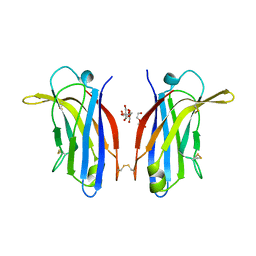 | | Human KIBRA C2 domain mutant M734I S735A | | Descriptor: | DI(HYDROXYETHYL)ETHER, GLYCEROL, Protein KIBRA | | Authors: | Crennell, S.J, Posner, M.G, Bagby, S. | | Deposit date: | 2017-12-21 | | Release date: | 2018-05-16 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.64215851 Å) | | Cite: | Distinctive phosphoinositide- and Ca2+-binding properties of normal and cognitive performance-linked variant forms of KIBRA C2 domain.
J. Biol. Chem., 293, 2018
|
|
3KUM
 
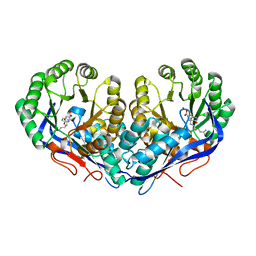 | | Crystal structure of Dipeptide Epimerase from Enterococcus faecalis V583 complexed with Mg and dipeptide L-Arg-L-Tyr | | Descriptor: | ARGININE, Dipeptide Epimerase, MAGNESIUM ION, ... | | Authors: | Fedorov, A.A, Fedorov, E.V, Sakai, A, Gerlt, J.A, Almo, S.C. | | Deposit date: | 2009-11-27 | | Release date: | 2010-10-13 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Homology models guide discovery of diverse enzyme specificities among dipeptide epimerases in the enolase superfamily.
Proc.Natl.Acad.Sci.USA, 109, 2012
|
|
6FXF
 
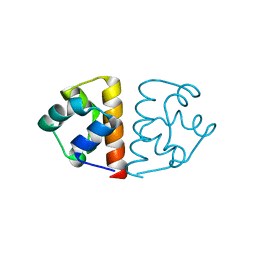 | |
6NDZ
 
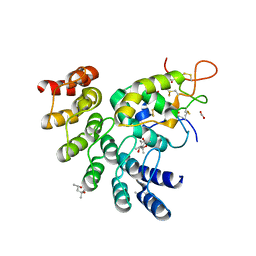 | | Designed repeat protein in complex with Fz8 | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, ACETATE ION, ACETIC ACID, ... | | Authors: | Miao, Y, Jude, K.M, Garcia, K.C. | | Deposit date: | 2018-12-14 | | Release date: | 2019-05-15 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.264 Å) | | Cite: | Receptor subtype discrimination using extensive shape complementary designed interfaces.
Nat.Struct.Mol.Biol., 26, 2019
|
|
6NE2
 
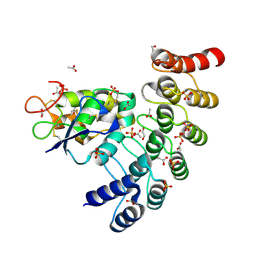 | | Designed repeat protein in complex with Fz7 | | Descriptor: | 1,2-ETHANEDIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose, ACETATE ION, ... | | Authors: | Miao, Y, Jude, K.M, Garcia, K.C. | | Deposit date: | 2018-12-15 | | Release date: | 2019-05-15 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.299 Å) | | Cite: | Receptor subtype discrimination using extensive shape complementary designed interfaces.
Nat.Struct.Mol.Biol., 26, 2019
|
|
