8ITG
 
 | |
8IIP
 
 | |
8IIN
 
 | |
8IIR
 
 | | MsmUdgX H109S/Q53A double mutant | | Descriptor: | BETA-MERCAPTOETHANOL, IRON/SULFUR CLUSTER, Type-4 uracil-DNA glycosylase | | Authors: | Aroli, S. | | Deposit date: | 2023-02-24 | | Release date: | 2023-06-21 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (2.28 Å) | | Cite: | Mutational and structural analyses of UdgX: insights into the active site pocket architecture and its evolution.
Nucleic Acids Res., 51, 2023
|
|
8ILX
 
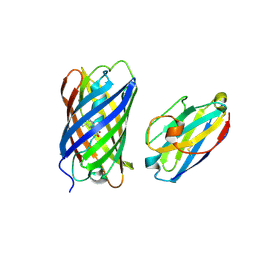 | | mCherry-LaM3 complex | | Descriptor: | LAM3, MCherry fluorescent protein | | Authors: | Liang, H, Liu, R, Ding, Y. | | Deposit date: | 2023-03-05 | | Release date: | 2023-06-21 | | Method: | X-RAY DIFFRACTION (3.29 Å) | | Cite: | Structural Insights into the Binding of Red Fluorescent Protein mCherry-Specific Nanobodies.
Int J Mol Sci, 24, 2023
|
|
8IM1
 | | mCherry-LaM1 complex | | Descriptor: | LaM1, MCherry fluorescent protein, SULFATE ION | | Authors: | Liang, H, Liu, R, Ding, Y. | | Deposit date: | 2023-03-05 | | Release date: | 2023-06-21 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Structural Insights into the Binding of Red Fluorescent Protein mCherry-Specific Nanobodies.
Int J Mol Sci, 24, 2023
|
|
5VYL
 
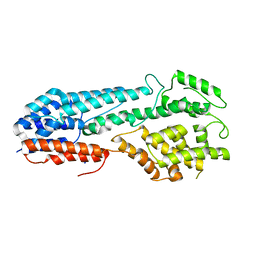 | |
8IGV
 | | Hexameric Ring Complex of Engineered V1-ATPase bound to 5 ADPs: A3(De)3_(ADP-Pi)1cat(ADP)2cat,2non-cat | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, MAGNESIUM ION, PHOSPHATE ION, ... | | Authors: | Kosugi, T, Tanabe, M, Koga, N. | | Deposit date: | 2023-02-21 | | Release date: | 2023-07-12 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (3.15 Å) | | Cite: | Design of allosteric sites into rotary motor V 1 -ATPase by restoring lost function of pseudo-active sites.
Nat.Chem., 15, 2023
|
|
8III
 
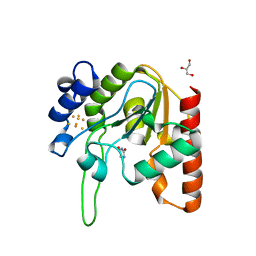 | |
5W4P
 
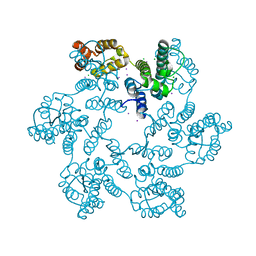 | |
8IIG
 
 | |
5W0C
 
 | | Cytochrome P450 (CYP) 2C9 TCA007 Inhibitor Complex | | Descriptor: | Cytochrome P450 2C9, GLYCEROL, PROTOPORPHYRIN IX CONTAINING FE, ... | | Authors: | Johnson, E.F, Hsu, M.-H. | | Deposit date: | 2017-05-30 | | Release date: | 2017-09-13 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.001 Å) | | Cite: | Determinants of the Inhibition of DprE1 and CYP2C9 by Antitubercular Thiophenes.
Angew. Chem. Int. Ed. Engl., 56, 2017
|
|
5W0N
 
 | | Structure of human TUT7 catalytic module (CM) in complex with UMPNPP and U2 RNA | | Descriptor: | 5'-O-[(S)-hydroxy{[(S)-hydroxy(phosphonooxy)phosphoryl]amino}phosphoryl]uridine, IODIDE ION, MAGNESIUM ION, ... | | Authors: | Faehnle, C.R, Walleshauser, J, Joshua-Tor, L. | | Deposit date: | 2017-05-31 | | Release date: | 2017-06-28 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.497 Å) | | Cite: | Multi-domain utilization by TUT4 and TUT7 in control of let-7 biogenesis.
Nat. Struct. Mol. Biol., 24, 2017
|
|
5W1C
 
 | | Crystal structure of MBP fused activation-induced cytidine deaminase (AID) in complex with cytidine | | Descriptor: | 4-AMINO-1-BETA-D-RIBOFURANOSYL-2(1H)-PYRIMIDINONE, CALCIUM ION, DNA (5'-D(*CP*TP*GP*GP*CP*CP*TP*TP*GP*AP*AP*C)-3'), ... | | Authors: | Qiao, Q, Wang, L, Wu, H. | | Deposit date: | 2017-06-02 | | Release date: | 2017-08-16 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (3.18 Å) | | Cite: | AID Recognizes Structured DNA for Class Switch Recombination.
Mol. Cell, 67, 2017
|
|
5W1I
 
 | |
8IQB
 
 | |
8IQD
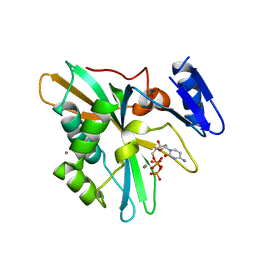 
 | |
5W4H
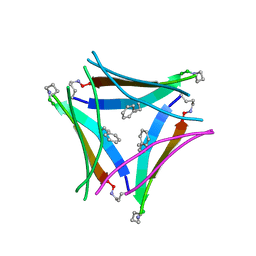 
 | |
5W4I
 
 | |
5W4O
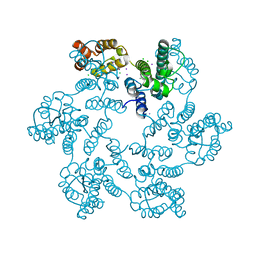 
 | |
5W57
 
 | | Structure of Holo AztC | | Descriptor: | Periplasmic solute binding protein, ZINC ION | | Authors: | Avalos, D, Yukl, E.T. | | Deposit date: | 2017-06-14 | | Release date: | 2017-09-20 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Mechanisms of zinc binding to the solute-binding protein AztC and transfer from the metallochaperone AztD.
J. Biol. Chem., 292, 2017
|
|
8I50
 
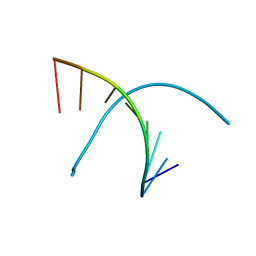 | | Crystal structure of DNA octamer containing GuNA[Me,Me] | | Descriptor: | DNA (5'-D(*GP*(OIQ)P*GP*(BRU)P*AP*CP*AP*C)-3') | | Authors: | Aoyama, H, Obika, S, Yamaguchi, T. | | Deposit date: | 2023-01-21 | | Release date: | 2023-08-09 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (0.95 Å) | | Cite: | Mechanism of the extremely high duplex-forming ability of oligonucleotides modified with N-tert-butylguanidine- or N-tert-butyl-N'- methylguanidine-bridged nucleic acids.
Nucleic Acids Res., 51, 2023
|
|
5W56
 
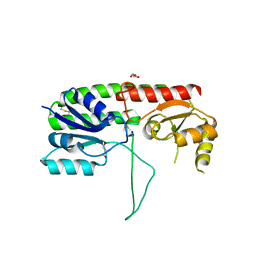 | | Structure of Apo AztC | | Descriptor: | GLYCEROL, Periplasmic solute binding protein, SODIUM ION | | Authors: | Avalos, D, Yukl, E.T. | | Deposit date: | 2017-06-14 | | Release date: | 2017-09-20 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.03 Å) | | Cite: | Mechanisms of zinc binding to the solute-binding protein AztC and transfer from the metallochaperone AztD.
J. Biol. Chem., 292, 2017
|
|
8HY7
 
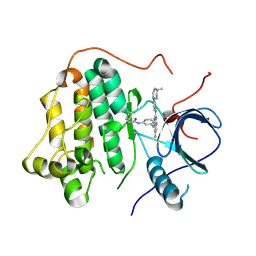 | | EGFR kinase domain mutant "TMLR" with compound 28f | | Descriptor: | Epidermal growth factor receptor, ~{N}-[3-[[6-fluoranyl-2-[[4-(4-methylpiperazin-1-yl)phenyl]amino]quinazolin-4-yl]amino]phenyl]prop-2-enamide | | Authors: | Wang, Y.X, Wu, C.Y. | | Deposit date: | 2023-01-06 | | Release date: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.91 Å) | | Cite: | The crystal structure of EGFR kinase domain(T790M/L858R) in complex with compound 28f
To Be Published
|
|
5W4J
 
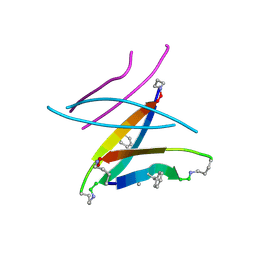 | |
