3O17
 
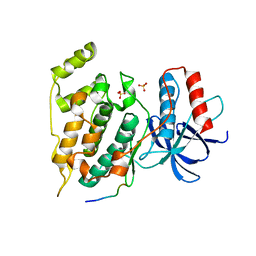 | | Crystal Structure of JNK1-alpha1 isoform | | Descriptor: | C-Jun-amino-terminal kinase-interacting protein 1, JIP1, 10MER PEPTIDE, ... | | Authors: | Abad-Zapatero, C. | | Deposit date: | 2010-07-20 | | Release date: | 2011-01-12 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Crystal Structure of JNK1-alpha1 isoform
TO BE PUBLISHED
|
|
3E7O
 
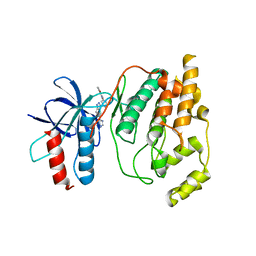 | | Crystal Structure of JNK2 | | Descriptor: | Mitogen-activated protein kinase 9, N-{3-[5-(1H-1,2,4-triazol-3-yl)-1H-indazol-3-yl]phenyl}furan-2-carboxamide | | Authors: | Kuglstatter, A, Villasenor, A.G. | | Deposit date: | 2008-08-18 | | Release date: | 2008-09-09 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.14 Å) | | Cite: | The crystal structure of JNK2 reveals conformational flexibility in the MAP kinase insert and indicates its involvement in the regulation of catalytic activity.
J.Mol.Biol., 383, 2008
|
|
4E73
 
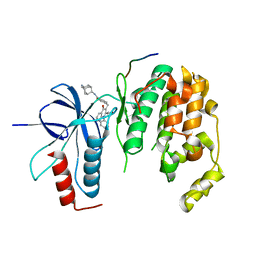 | | Crystal structure of JNK1beta-JIP in complex with an azaquinolone inhbitor | | Descriptor: | C-Jun-amino-terminal kinase-interacting protein 1, Mitogen-activated protein kinase 8, methyl 3-(4-{[(1R,2S,3S,5S,7s)-5-aminotricyclo[3.3.1.1~3,7~]dec-2-yl]carbamoyl}benzyl)-4-oxo-1-phenyl-1,4-dihydro-1,8-naphthyridine-2-carboxylate | | Authors: | Lukacs, C.M, Janson, C.A. | | Deposit date: | 2012-03-16 | | Release date: | 2013-05-29 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.27 Å) | | Cite: | Identification of an Adamantyl Azaquinolone JNK Selective Inhibitor.
ACS Med Chem Lett, 3, 2012
|
|
7XMH
 
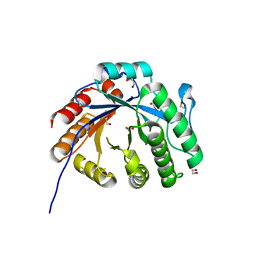 | | Crystal structure of a rice class IIIb chitinase, Oschib2 | | Descriptor: | 1,2-ETHANEDIOL, ACETATE ION, Putative class III chitinase | | Authors: | Jun, T, Tomoya, T, Tomoyuki, N, Takayuki, O. | | Deposit date: | 2022-04-25 | | Release date: | 2023-05-03 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.18 Å) | | Cite: | Characterization of two rice GH18 chitinases belonging to family 8 of plant pathogenesis-related proteins.
Plant Sci., 326, 2023
|
|
1FMH
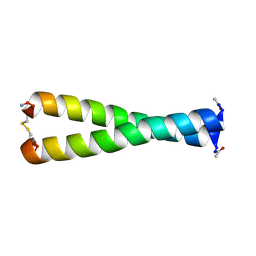 
 | |
4H3B
 
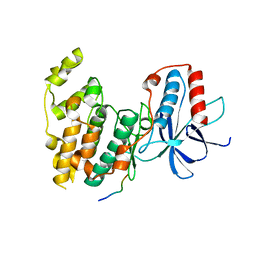 | | Crystal Structure of JNK3 in Complex with SAB Peptide | | Descriptor: | Mitogen-activated protein kinase 10, SH3 domain-binding protein 5 | | Authors: | Nwachukwu, J.C, Laughlin, J.D, Figuera-Losada, M, Cherry, L, Nettles, K.W, LoGrasso, P.V. | | Deposit date: | 2012-09-13 | | Release date: | 2012-11-21 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.08 Å) | | Cite: | Structural Mechanisms of Allostery and Autoinhibition in JNK Family Kinases.
Structure, 20, 2012
|
|
4H36
 
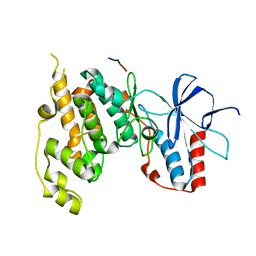 | | Crystal Structure of JNK3 in Complex with ATF2 Peptide | | Descriptor: | Cyclic AMP-dependent transcription factor ATF-2, Mitogen-activated protein kinase 10 | | Authors: | Nwachukwu, J.C, Laughlin, J.D, Figuera-Losada, M, Cherry, L, Nettles, K.W, LoGrasso, P.V. | | Deposit date: | 2012-09-13 | | Release date: | 2012-11-21 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structural Mechanisms of Allostery and Autoinhibition in JNK Family Kinases.
Structure, 20, 2012
|
|
4G1W
 
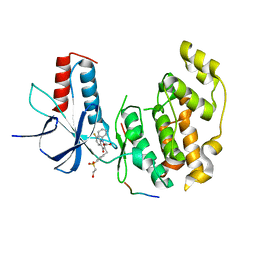 | |
5RAT
 
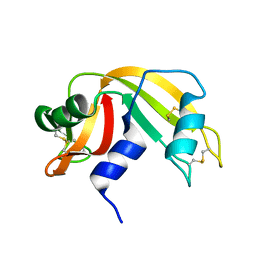 | |
3QD2
 
 | | Crystal structure of mouse PERK kinase domain | | Descriptor: | Eukaryotic translation initiation factor 2-alpha kinase 3 | | Authors: | Wenjun, C, Jingzhi, L, David, R, Bingdong, S. | | Deposit date: | 2011-01-17 | | Release date: | 2011-04-27 | | Last modified: | 2019-01-16 | | Method: | X-RAY DIFFRACTION (2.81 Å) | | Cite: | The structure of the PERK kinase domain suggests the mechanism for its activation.
Acta Crystallogr.,Sect.D, 67, 2011
|
|
4RAT
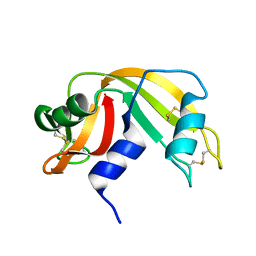 
 | |
7RAT
 
 | |
6N4V
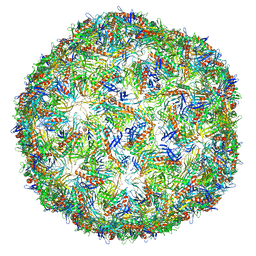 
 | | CryoEM structure of Leviviridae PP7 WT coat protein dimer capsid (PP7PP7-WT) | | Descriptor: | Coat protein | | Authors: | Liangjun, Z, Kopylov, M, Potter, C.S, Carragher, B, Finn, M.G. | | Deposit date: | 2018-11-20 | | Release date: | 2019-04-03 | | Last modified: | 2019-12-18 | | Method: | ELECTRON MICROSCOPY (3 Å) | | Cite: | Engineering the PP7 Virus Capsid as a Peptide Display Platform.
Acs Nano, 13, 2019
|
|
1RAT
 
 | |
2RAT
 | |
3MEN
 
 | |
1FDH
 
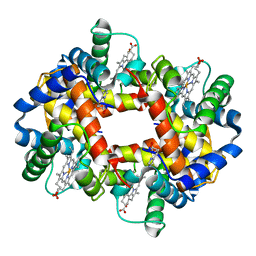 | | STRUCTURE OF HUMAN FOETAL DEOXYHAEMOGLOBIN | | Descriptor: | HEMOGLOBIN F (DEOXY) (ALPHA CHAIN), HEMOGLOBIN F (DEOXY) (GAMMA CHAIN), PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Frierjunior, J.A. | | Deposit date: | 1976-08-18 | | Release date: | 1976-08-19 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structure of human foetal deoxyhaemoglobin.
J.Mol.Biol., 112, 1977
|
|
8RAT
 | |
3LNU
 
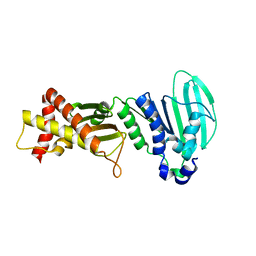 | |
3LPX
 
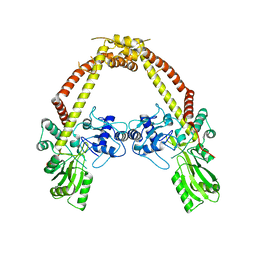 | | Crystal structure of GyrA | | Descriptor: | DNA gyrase, A subunit | | Authors: | Jung, H.Y, Heo, Y.S. | | Deposit date: | 2010-02-07 | | Release date: | 2011-02-16 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Crystal structure of GyrA
To be Published
|
|
3RAT
 | |
4W4X
 
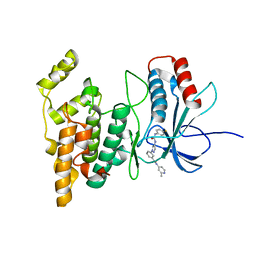 | | JNK2/3 in complex with 3-(4-{[(4-fluorophenyl)carbamoyl]amino}-1H-pyrazol-1-yl)-N-(2-methylpyridin-4-yl)benzamide | | Descriptor: | 3-(4-{[(4-fluorophenyl)carbamoyl]amino}-1H-pyrazol-1-yl)-N-(2-methylpyridin-4-yl)benzamide, c-jun NH2-terminal kinase 3 | | Authors: | Park, H, Iqbal, S, Hernandez, P, Mora, R, Zheng, K, Feng, Y, LoGrasso, P. | | Deposit date: | 2014-08-15 | | Release date: | 2015-02-11 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.65 Å) | | Cite: | Structural Basis and Biological Consequences for JNK2/3 Isoform Selective Aminopyrazoles.
Sci Rep, 5, 2015
|
|
4UX9
 
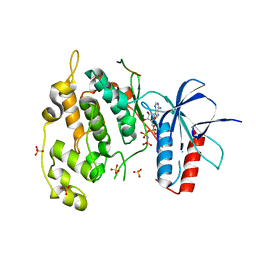 | | Crystal structure of JNK1 bound to a MKK7 docking motif | | Descriptor: | DUAL SPECIFICITY MITOGEN-ACTIVATED PROTEIN KINASE KINASE 7, MITOGEN-ACTIVATED PROTEIN KINASE 8, PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER, ... | | Authors: | Kragelj, J, Palencia, A, Nanao, M.H, Maurin, D, Bouvignies, G, Blackledge, M, Ringkjobing-Jensen, M. | | Deposit date: | 2014-08-20 | | Release date: | 2015-03-25 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.34 Å) | | Cite: | Structure and Dynamics of the Mkk7-Jnk Signaling Complex.
Proc.Natl.Acad.Sci.USA, 112, 2015
|
|
2W83
 
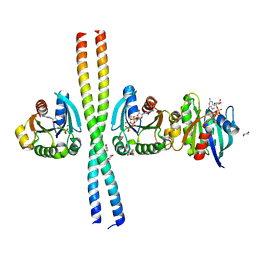 | | Crystal structure of the ARF6 GTPase in complex with a specific effector, JIP4 | | Descriptor: | 1,4-DIETHYLENE DIOXIDE, ADP-RIBOSYLATION FACTOR 6, C-JUN-AMINO-TERMINAL KINASE-INTERACTING PROTEIN 4, ... | | Authors: | Isabet, T, Montagnac, G, Regazzoni, K, Raynal, B, El Khadali, F, Franco, M, England, P, Chavrier, P, Houdusse, A, Menetrey, J. | | Deposit date: | 2009-01-08 | | Release date: | 2009-07-14 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.93 Å) | | Cite: | The Structural Basis of Arf Effector Specificity: The Crystal Structure of Arf6 in a Complex with Jip4.
Embo J., 28, 2009
|
|
4W4Y
 
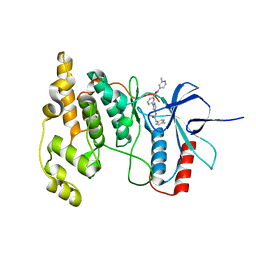 | | JNK2/3 in complex with 3-(4-{[(4-methylphenyl)carbamoyl]amino}-1H-pyrazol-1-yl)-N-(2-methylpyridin-4-yl)benzamide | | Descriptor: | 3-(4-{[(4-methylphenyl)carbamoyl]amino}-1H-pyrazol-1-yl)-N-(2-methylpyridin-4-yl)benzamide, c-jun NH2-terminal kinase 3 | | Authors: | Park, H, Iqbal, S, Hernandez, P, Mora, R, Zheng, K, Feng, Y, LoGrasso, P. | | Deposit date: | 2014-08-15 | | Release date: | 2015-02-11 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structural Basis and Biological Consequences for JNK2/3 Isoform Selective Aminopyrazoles.
Sci Rep, 5, 2015
|
|
