7OP5
 
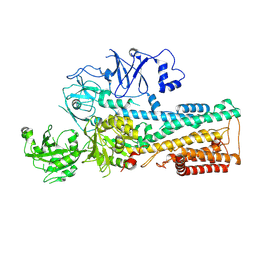 | | Cryo-EM structure of P5B-ATPase E2P | | Descriptor: | BERYLLIUM TRIFLUORIDE ION, MAGNESIUM ION, P5B-ATPase | | Authors: | Li, P, Gourdon, P. | | Deposit date: | 2021-05-29 | | Release date: | 2021-06-30 | | Last modified: | 2024-07-17 | | Method: | ELECTRON MICROSCOPY (3.7 Å) | | Cite: | Structure and transport mechanism of P5B-ATPases.
Nat Commun, 12, 2021
|
|
7OP8
 
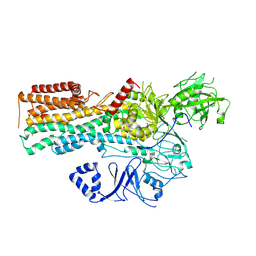 | | Cryo-EM structure of P5B-ATPase E2Pinhibit | | Descriptor: | BERYLLIUM TRIFLUORIDE ION, Cation-transporting ATPase, MAGNESIUM ION | | Authors: | Li, P, Gourdon, P. | | Deposit date: | 2021-05-31 | | Release date: | 2021-06-30 | | Last modified: | 2024-07-17 | | Method: | ELECTRON MICROSCOPY (3.5 Å) | | Cite: | Structure and transport mechanism of P5B-ATPases.
Nat Commun, 12, 2021
|
|
7P2P
 
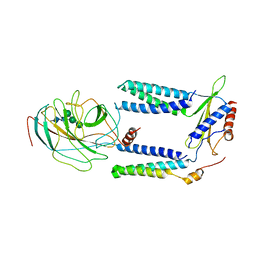 | | Human Signal Peptidase Complex Paralog A (SPC-A) | | Descriptor: | Signal peptidase complex catalytic subunit SEC11A, Signal peptidase complex subunit 1, Signal peptidase complex subunit 2, ... | | Authors: | Liaci, A.M, Foerster, F. | | Deposit date: | 2021-07-06 | | Release date: | 2021-10-06 | | Last modified: | 2022-12-21 | | Method: | ELECTRON MICROSCOPY (4.9 Å) | | Cite: | Structure of the human signal peptidase complex reveals the determinants for signal peptide cleavage.
Mol.Cell, 81, 2021
|
|
1C7I
 
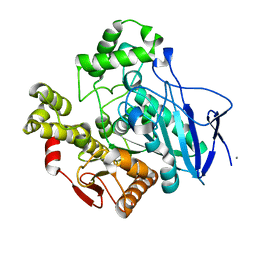 | | THERMOPHYLIC PNB ESTERASE | | Descriptor: | CALCIUM ION, PROTEIN (PARA-NITROBENZYL ESTERASE) | | Authors: | Spiller, B, Gershenson, A, Arnold, F, Stevens, R. | | Deposit date: | 2000-02-21 | | Release date: | 2000-03-29 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | A structural view of evolutionary divergence.
Proc.Natl.Acad.Sci.USA, 96, 1999
|
|
7P2Q
 
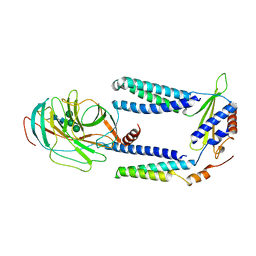 | | Human Signal Peptidase Complex Paralog C (SPC-C) | | Descriptor: | Signal peptidase complex catalytic subunit SEC11C, Signal peptidase complex subunit 1, Signal peptidase complex subunit 2, ... | | Authors: | Liaci, A.M, Foerster, F. | | Deposit date: | 2021-07-06 | | Release date: | 2021-10-06 | | Last modified: | 2022-12-21 | | Method: | ELECTRON MICROSCOPY (4.9 Å) | | Cite: | Structure of the human signal peptidase complex reveals the determinants for signal peptide cleavage.
Mol.Cell, 81, 2021
|
|
5JS1
 
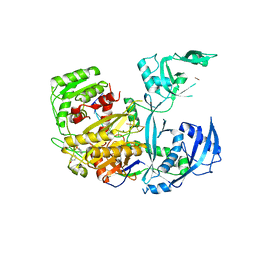 | | Human Argonaute2 Bound to an siRNA | | Descriptor: | MAGNESIUM ION, PHENOL, Protein argonaute-2, ... | | Authors: | Schirle, N.T, MacRae, I.J. | | Deposit date: | 2016-05-07 | | Release date: | 2016-07-20 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.499 Å) | | Cite: | Structural Analysis of Human Argonaute-2 Bound to a Modified siRNA Guide.
J.Am.Chem.Soc., 138, 2016
|
|
1JEW
 
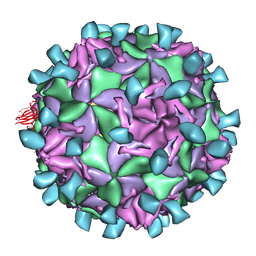 | |
1AZ6
 | |
5KHO
 
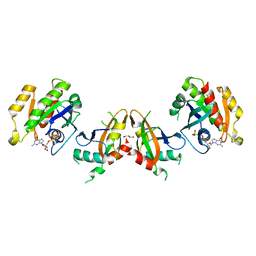 | | Rasip1 RA domain in complex with Rap1B | | Descriptor: | GLYCEROL, MAGNESIUM ION, PHOSPHOAMINOPHOSPHONIC ACID-GUANYLATE ESTER, ... | | Authors: | Gingras, A.R. | | Deposit date: | 2016-06-15 | | Release date: | 2016-10-19 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.78 Å) | | Cite: | Structural Basis of Dimeric Rasip1 RA Domain Recognition of the Ras Subfamily of GTP-Binding Proteins.
Structure, 24, 2016
|
|
7P1J
 
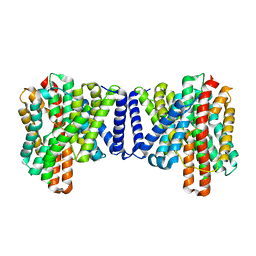 | | Cryo EM structure of bison NHA2 in detergent structure | | Descriptor: | mitochondrial sodium/hydrogen exchanger 9B2 | | Authors: | Matsuoka, R, Fudim, R, Jung, S, Drew, D. | | Deposit date: | 2021-07-01 | | Release date: | 2022-01-26 | | Last modified: | 2024-07-17 | | Method: | ELECTRON MICROSCOPY (3.04 Å) | | Cite: | Structure, mechanism and lipid-mediated remodeling of the mammalian Na + /H + exchanger NHA2.
Nat.Struct.Mol.Biol., 29, 2022
|
|
7P1I
 
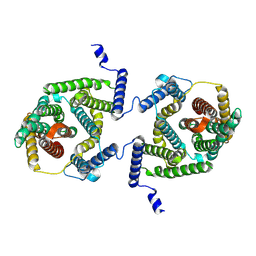 | | Cryo EM structure of bison NHA2 in detergent and N-terminal extension helix | | Descriptor: | mitochondrial sodium/hydrogen exchanger 9B2 | | Authors: | Matsuoka, R, Fudim, R, Jung, S, Drew, D. | | Deposit date: | 2021-07-01 | | Release date: | 2022-01-26 | | Last modified: | 2024-07-17 | | Method: | ELECTRON MICROSCOPY (3.15 Å) | | Cite: | Structure, mechanism and lipid-mediated remodeling of the mammalian Na + /H + exchanger NHA2.
Nat.Struct.Mol.Biol., 29, 2022
|
|
4YVY
 
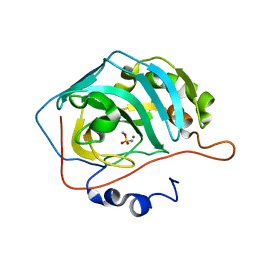 | |
1CGX
 
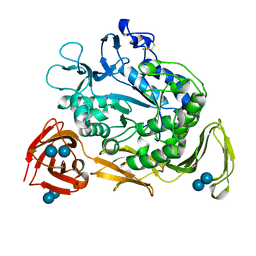 | |
5JME
 | |
2BE7
 
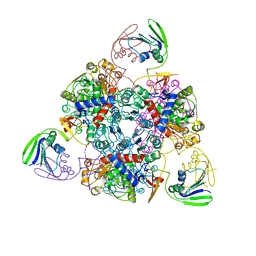 | | Crystal structure of the unliganded (T-state) aspartate transcarbamoylase of the psychrophilic bacterium Moritella profunda | | Descriptor: | Aspartate Carbamoyltransferase Catalytic Chain, Aspartate Carbamoyltransferase Regulatory Chain, SULFATE ION, ... | | Authors: | De Vos, D, Savvides, S.N, Van Beeumen, J. | | Deposit date: | 2005-10-23 | | Release date: | 2006-10-31 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.85 Å) | | Cite: | Structural investigation of cold activity and regulation of aspartate carbamoyltransferase from the extreme psychrophilic bacterium Moritella profunda.
J.Mol.Biol., 365, 2007
|
|
2OFF
 
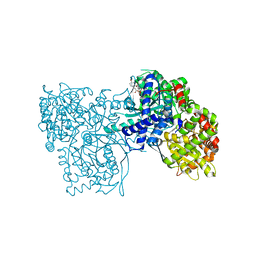 | | The crystal structure of Glycogen Phosphorylase b in complex with a potent allosteric inhibitor | | Descriptor: | 2-DEOXY-3,4-BIS-O-[3-(4-HYDROXYPHENYL)PROPANOYL]-L-THREO-PENTARIC ACID, Glycogen phosphorylase, muscle form | | Authors: | Tiraidis, C, Alexacou, K.-M, Zographos, S.E, Leonidas, D.D, Gimisis, T, Oikonomakos, N.G. | | Deposit date: | 2007-01-03 | | Release date: | 2007-08-07 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | FR258900, a potential anti-hyperglycemic drug, binds at the allosteric site of glycogen phosphorylase
Protein Sci., 16, 2007
|
|
1CGW
 
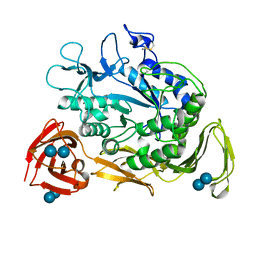 | |
1D4I
 | | HIV-1 protease in complex with the inhibitor BEA425 | | Descriptor: | 2,5-DIBENZYLOXY-3-HYDROXY-HEXANEDIOIC ACID BIS-[(2-HYDROXY-INDAN-1-YL)-AMIDE], HIV-1 PROTEASE | | Authors: | Unge, T. | | Deposit date: | 1999-10-04 | | Release date: | 2002-06-26 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.81 Å) | | Cite: | Optimization of P1-P3 groups in symmetric and asymmetric HIV-1 protease inhibitors.
Eur.J.Biochem., 270, 2003
|
|
4ZEV
 
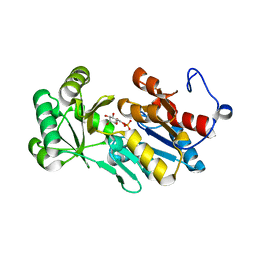 | | Crystal structure of PfHAD1 in complex with mannose-6-phosphate | | Descriptor: | 6-O-phosphono-alpha-D-mannopyranose, MAGNESIUM ION, PHOSPHATE ION, ... | | Authors: | Park, J, Tolia, N.H. | | Deposit date: | 2015-04-20 | | Release date: | 2015-09-09 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Cap-domain closure enables diverse substrate recognition by the C2-type haloacid dehalogenase-like sugar phosphatase Plasmodium falciparum HAD1.
Acta Crystallogr. D Biol. Crystallogr., 71, 2015
|
|
5EJ2
 
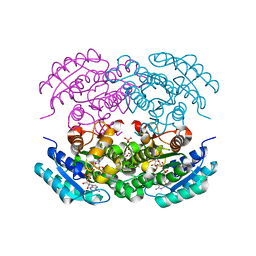 | |
1CGY
 
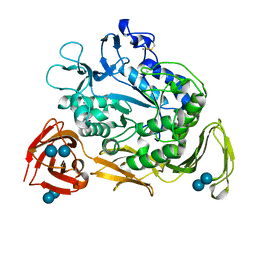 | |
1GRX
 
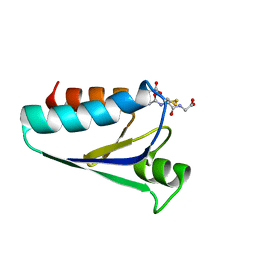 | | STRUCTURE OF E. COLI GLUTAREDOXIN | | Descriptor: | GLUTAREDOXIN, GLUTATHIONE | | Authors: | Bushweller, J.H, Billeter, M, Holmgren, L.A, Wuthrich, K. | | Deposit date: | 1993-10-01 | | Release date: | 1994-01-31 | | Last modified: | 2021-11-03 | | Method: | SOLUTION NMR | | Cite: | NMR structure of oxidized Escherichia coli glutaredoxin: comparison with reduced E. coli glutaredoxin and functionally related proteins.
Protein Sci., 1, 1992
|
|
5J5K
 
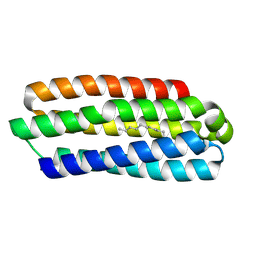 | | CRYSTAL STRUCTURE OF AFMP4P IN COMPLEX WITH PALMITIC ACID | | Descriptor: | PALMITIC ACID, Uncharacterized protein, alpha-D-mannopyranose | | Authors: | Zhang, H, Hao, Q. | | Deposit date: | 2016-04-03 | | Release date: | 2017-04-05 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | A Novel Class Of Virulence Factors In Penicillium Marneffei And Aspergillus Fumigatus Enhances Intracellular Survival In Monocytes By Arachidonic Acid Binding
To Be Published
|
|
1H6N
 
 | | Formation of a tyrosyl radical intermediate in Proteus mirabilis catalase by directed mutagenesis and consequences for nucleotide reactivity | | Descriptor: | ACETATE ION, CATALASE, PROTOPORPHYRIN IX CONTAINING FE, ... | | Authors: | Andreoletti, P, Sainz, G, Jaquinod, M, Gagnon, J, Jouve, H.M. | | Deposit date: | 2001-06-20 | | Release date: | 2003-10-03 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.11 Å) | | Cite: | High Resolution Structure and Biochemical Properties of a Recombinant Proteus Mirabilis Catalase Depleted in Iron.
Proteins: Struct.,Funct., Genet., 50, 2003
|
|
1H7K
 
 | | Formation of a tyrosyl radical intermediate in Proteus mirabilis catalase by directed mutagenesis and consequences for nucleotide reactivity | | Descriptor: | ACETATE ION, CATALASE, PROTOPORPHYRIN IX CONTAINING FE, ... | | Authors: | Andreoletti, P, Gambarelli, S, Gaillard, J, Sainz, G, Stojanoff, V, Jouve, H.M. | | Deposit date: | 2001-07-08 | | Release date: | 2004-02-25 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Formation of a Tyrosyl Radical Intermediate in Proteus Mirabilis Catalase by Directed Mutagenesis and Consequences for Nucleotide Reactivity.
Biochemistry, 40, 2001
|
|
