2MRB
 | | THREE-DIMENSIONAL STRUCTURE OF RABBIT LIVER CD-7 METALLOTHIONEIN-2A IN AQUEOUS SOLUTION DETERMINED BY NUCLEAR MAGNETIC RESONANCE | | Descriptor: | CADMIUM ION, CD7 METALLOTHIONEIN-2A | | Authors: | Braun, W, Arseniev, A, Schultze, P, Woergoetter, E, Wagner, G, Vasak, M, Kaegi, J.H.R, Wuthrich, K. | | Deposit date: | 1990-05-14 | | Release date: | 1991-04-15 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Three-dimensional structure of rabbit liver [Cd7]metallothionein-2a in aqueous solution determined by nuclear magnetic resonance.
J.Mol.Biol., 201, 1988
|
|
2LHT
 
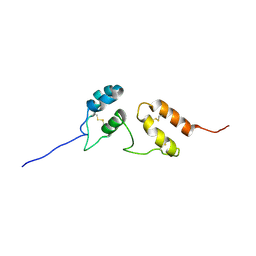 | | Solution structure of Venturia inaequalis cellophane-induced 1 protein (ViCin1) domains 1 and 2 | | Descriptor: | Cellophane-induced protein 1 | | Authors: | Mesarich, C.H, Schmitz, M, Tremouilhac, P, Greenwood, D.R, Mcgillivray, D.J, Templeton, M.D, Dingley, A.J. | | Deposit date: | 2011-08-16 | | Release date: | 2012-07-18 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Structure, dynamics and domain organization of the repeat protein Cin1 from the apple scab fungus.
Biochim.Biophys.Acta, 1824, 2012
|
|
2M3U
 
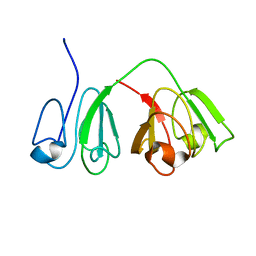 | |
2M3T
 
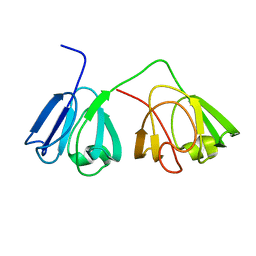 | |
2M0A
 
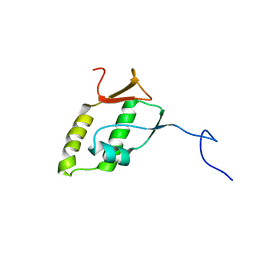 | | Solution structure of MHV nsp3a | | Descriptor: | Non-structural protein 3 | | Authors: | Keane, S.C, Giedroc, D.P. | | Deposit date: | 2012-10-23 | | Release date: | 2013-01-23 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of Mouse Hepatitis Virus (MHV) nsp3a and Determinants of the Interaction with MHV Nucleocapsid (N) Protein.
J.Virol., 87, 2013
|
|
2MGT
 
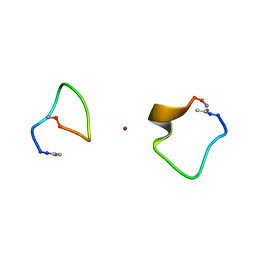 | |
2MXA
 | | Solution structure of the NDH-1 complex subunit CupS from Thermosynechococcus elongatus | | Descriptor: | NDH-1 complex sensory subunit CupS | | Authors: | Korste, A, Wulfhorst, H, Ikegami, T, Nowaczyk, M.M, Stoll, R. | | Deposit date: | 2014-12-17 | | Release date: | 2015-06-10 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the NDH-1 complex subunit CupS from Thermosynechococcus elongatus.
Biochim.Biophys.Acta, 1847, 2015
|
|
2P6J
 
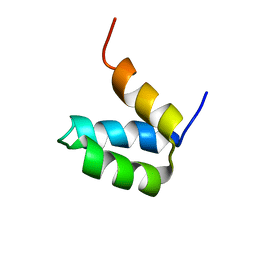 | | Full-sequence computational design and solution structure of a thermostable protein variant | | Descriptor: | designed engrailed homeodomain variant UVF | | Authors: | Shah, P.S, Hom, G.K, Ross, S.A, Lassila, J.K, Crowhurst, K.A, Mayo, S.L. | | Deposit date: | 2007-03-18 | | Release date: | 2007-08-14 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Full-sequence Computational Design and Solution Structure of a Thermostable Protein Variant.
J.Mol.Biol., 372, 2007
|
|
2KAP
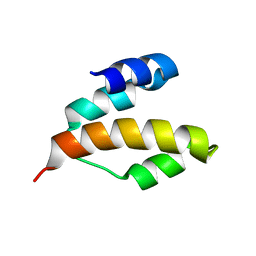 
 | | Solution structure of DLC1-SAM | | Descriptor: | Rho GTPase-activating protein 7 | | Authors: | Yang, S, Yang, D. | | Deposit date: | 2008-11-12 | | Release date: | 2009-10-20 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Characterization of DLC1-SAM equilibrium unfolding at the amino acid residue level
Biochemistry, 48, 2009
|
|
2OA4
 
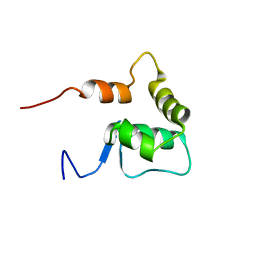 | | Solution NMR Structure: Northeast Structural Genomics Consortium Target SiR5 | | Descriptor: | SiR5 | | Authors: | Wang, L, Rossi, P, Chen, C.X, Nwosu, C, Cunningham, K, Ma, L.-C, Xiao, R, Liu, J, Baran, M.C, Swapna, G.V.T, Acton, T.B, Burkhard, R, Montelione, G.T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2006-12-14 | | Release date: | 2007-03-06 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | Northeast Structural Genomics Consortium Target SiR5
To be Published
|
|
2PHE
 
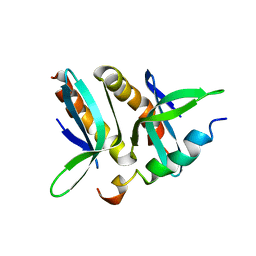 | | Model for VP16 binding to PC4 | | Descriptor: | Alpha trans-inducing protein, TRANSCRIPTIONAL COACTIVATOR PC4 | | Authors: | Jonker, H.R.A, Wechselberger, R.W, Boelens, R, Folkers, G.E, Kaptein, R. | | Deposit date: | 2007-04-11 | | Release date: | 2007-04-24 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structural Properties of the Promiscuous VP16 Activation Domain
Biochemistry, 44, 2005
|
|
2ND0
 
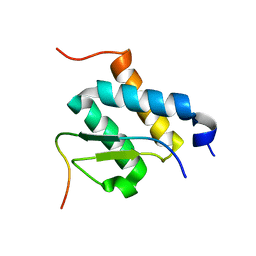 | |
2NCZ
 
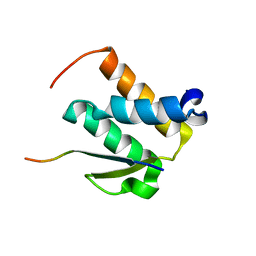 | |
2N8U
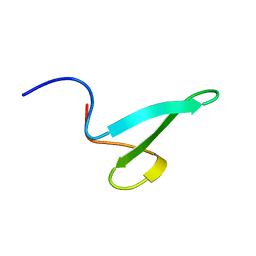 
 | |
2N8S
 
 | |
2N8T
 
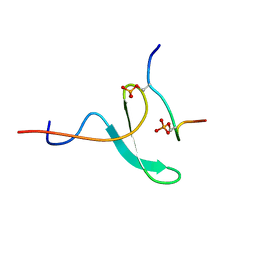 | |
2N9U
 
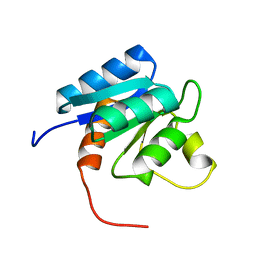 | |
2NBJ
 
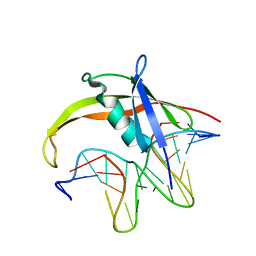 | | DNA-archeal MC1 protein complex structure by NMR | | Descriptor: | Chromosomal protein MC1, DNA (5'-D(*AP*AP*AP*AP*AP*CP*AP*CP*AP*CP*AP*CP*CP*CP*A)-3'), DNA (5'-D(P*TP*GP*GP*GP*TP*GP*TP*GP*TP*GP*TP*TP*TP*TP*T)-3') | | Authors: | Paquet, F, Loth, K, Landon, C. | | Deposit date: | 2016-02-25 | | Release date: | 2017-03-01 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | First 3D structure of an atypical protein-DNA complex
To be Published
|
|
2LHH
 
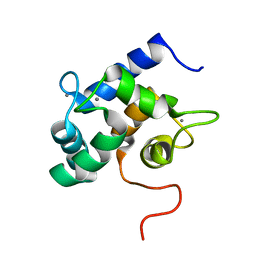 | | Solution structure of Ca2+-bound yCaM | | Descriptor: | CALCIUM ION, Calmodulin | | Authors: | Ogura, K, Takahashi, K, Kobashigawa, Y, Yoshida, R, Itoh, H, Yazawa, M, Inagaki, F. | | Deposit date: | 2011-08-10 | | Release date: | 2012-08-29 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Solution structures of yeast Saccharomyces cerevisiae calmodulin in calcium- and target peptide-bound states reveal similarities and differences to vertebrate calmodulin.
Genes Cells, 17, 2012
|
|
2M1M
 
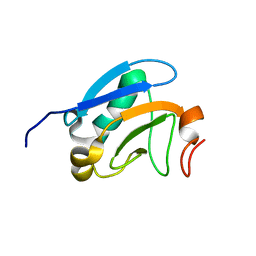 | | Solution structure of the PsIAA4 oligomerization domain reveals interaction modes for transcription factors in early auxin response | | Descriptor: | Auxin-induced protein IAA4 | | Authors: | Kovermann, M, Dinesh, D.C, Gopalswamy, M, Abel, S, Balbach, J. | | Deposit date: | 2012-12-03 | | Release date: | 2013-12-11 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the PsIAA4 oligomerization domain reveals interaction modes for transcription factors in early auxin response.
Proc.Natl.Acad.Sci.USA, 112, 2015
|
|
2M4K
 
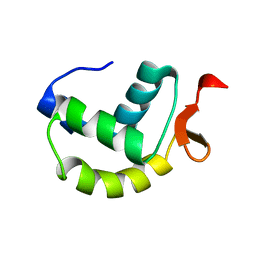 | | Solution structure of the delta subunit of RNA polymerase from Bacillus subtilis | | Descriptor: | DNA-directed RNA polymerase subunit delta | | Authors: | Papouskova, V, Novacek, J, Kaderavek, P, Zidek, L, Rabatinova, A, Sanderova, H, Krasny, L, Sklenar, V. | | Deposit date: | 2013-02-07 | | Release date: | 2013-10-09 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Structural Study of the Partially Disordered Full-Length delta Subunit of RNA Polymerase from Bacillus subtilis.
Chembiochem, 14, 2013
|
|
2MJN
 
 | |
2MNH
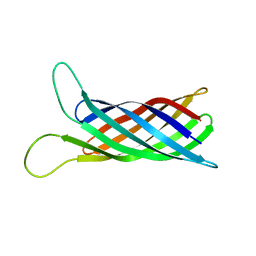 
 | | Refined structure of outer membrane protein x in nanodisc by measuring residual dipolar couplings | | Descriptor: | Outer membrane protein X | | Authors: | Bibow, S, Carneiro, M.G, Sabo, T.M, Schwiegk, C, Becker, S, Riek, R, Lee, D. | | Deposit date: | 2014-04-05 | | Release date: | 2015-03-18 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Measuring membrane protein bond orientations in nanodiscs via residual dipolar couplings.
Protein Sci., 23, 2014
|
|
2N9B
 
 | |
1DPU
 
 | | SOLUTION STRUCTURE OF THE C-TERMINAL DOMAIN OF HUMAN RPA32 COMPLEXED WITH UNG2(73-88) | | Descriptor: | REPLICATION PROTEIN A (RPA32) C-TERMINAL DOMAIN, URACIL DNA GLYCOSYLASE (UNG2) | | Authors: | Mer, G, Edwards, A.M, Chazin, W.J. | | Deposit date: | 1999-12-27 | | Release date: | 2000-11-10 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structural basis for the recognition of DNA repair proteins UNG2, XPA, and RAD52 by replication factor RPA.
Cell(Cambridge,Mass.), 103, 2000
|
|
