1QE6
 
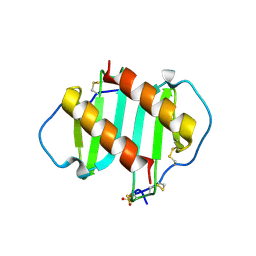 | | INTERLEUKIN-8 WITH AN ADDED DISULFIDE BETWEEN RESIDUES 5 AND 33 (L5C/H33C) | | Descriptor: | INTERLEUKIN-8 VARIANT, SULFATE ION | | Authors: | Gerber, N, Lowman, H, Artis, D.R, Eigenbrot, C. | | Deposit date: | 1999-07-13 | | Release date: | 2000-03-22 | | Last modified: | 2018-01-31 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Receptor-binding conformation of the "ELR" motif of IL-8: X-ray structure of the L5C/H33C variant at 2.35 A resolution.
Proteins, 38, 2000
|
|
1QLI
 
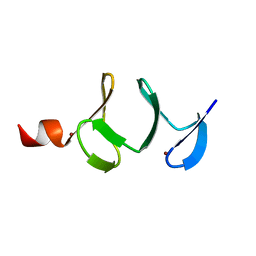 | | QUAIL CYSTEINE AND GLYCINE-RICH PROTEIN, NMR, MINIMIZED AVERAGE STRUCTURE | | Descriptor: | CYSTEINE AND GLYCINE-RICH PROTEIN, ZINC ION | | Authors: | Konrat, R, Weiskirchen, R, Krautler, B, Bister, K. | | Deposit date: | 1997-02-17 | | Release date: | 1997-08-20 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the carboxyl-terminal LIM domain from quail cysteine-rich protein CRP2.
J.Biol.Chem., 272, 1997
|
|
1QD3
 
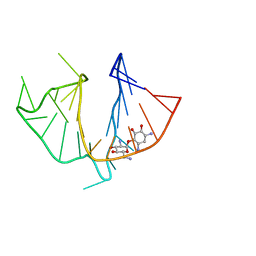 | | HIV-1 TAR RNA/NEOMYCIN B COMPLEX | | Descriptor: | 2,6-diamino-2,6-dideoxy-alpha-D-glucopyranose, 2,6-diamino-2,6-dideoxy-beta-L-idopyranose-(1-3)-alpha-D-ribofuranose, 2-DEOXYSTREPTAMINE, ... | | Authors: | Faber, C, Sticht, H, Roesch, P. | | Deposit date: | 1999-07-07 | | Release date: | 2000-07-12 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | Structural rearrangements of HIV-1 Tat-responsive RNA upon binding of neomycin B.
J.Biol.Chem., 275, 2000
|
|
1QFB
 
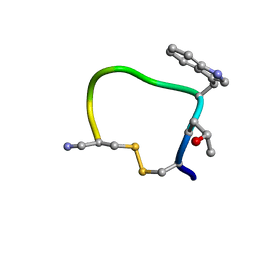 | | THE CYCLIC PEPTIDE CONTRYPHAN-R FROM CONUS RADIATUS | | Descriptor: | PROTEIN (CONTRYPHAN-R) | | Authors: | Pallaghy, P.K, Melnikova, A.P, Jimenez, E.C, Olivera, B.M, Norton, R.S. | | Deposit date: | 1999-04-08 | | Release date: | 1999-09-29 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | Solution structure of contryphan-R, a naturally occurring disulfide-bridged octapeptide containing D-tryptophan: comparison with protein loops.
Biochemistry, 38, 1999
|
|
1KAT
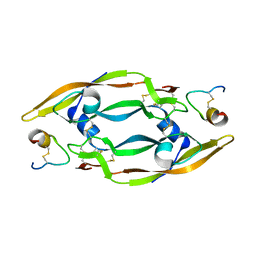 
 | | Solution Structure of a Phage-Derived Peptide Antagonist in Complex with Vascular Endothelial Growth Factor | | Descriptor: | Phage-Derived Peptide Antagonist, Vascular Endothelial Growth Factor | | Authors: | Pan, B, Li, B, Russell, S.J, Tom, J.Y.K, Cochran, A.G, Fairbrother, W.J. | | Deposit date: | 2001-11-02 | | Release date: | 2002-11-02 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of a Phage-derived Peptide Antagonist in Complex with Vascular Endothelial Growth Factor
J.Mol.Biol., 316, 2002
|
|
1MXL
 
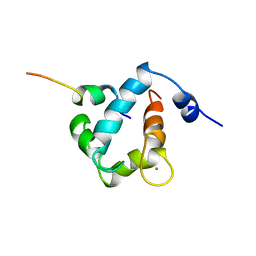 | |
1QH2
 
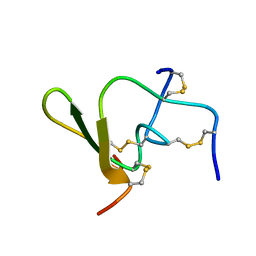 | |
1QFA
 
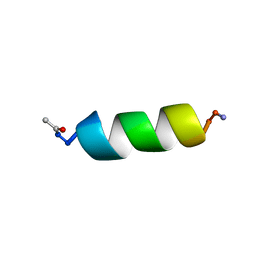 | | STRUCTURE OF A NEUROPEPTIDE Y Y2 AGONIST | | Descriptor: | PROTEIN (NEUROPEPTIDE Y) | | Authors: | Barnham, K.J, Catalfamo, F, Pallaghy, P.K, Howlett, G.J, Norton, R.S. | | Deposit date: | 1999-04-08 | | Release date: | 2000-04-08 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | Helical structure and self-association in a 13 residue neuropeptide Y Y2 receptor agonist: relationship to biological activity.
Biochim.Biophys.Acta, 1435, 1999
|
|
1PZR
 
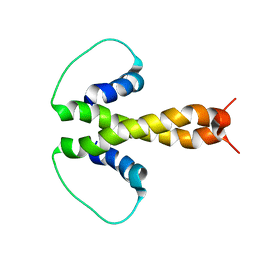 | | Structure of fused docking domains from the erythromycin polyketide synthase (DEBS), a model for the interaction between DEBS2 and DEBS3: the B domain | | Descriptor: | Erythronolide synthase | | Authors: | Broadhurst, R.W, Nietlispach, D, Wheatcroft, M.P, Leadlay, P.F, Weissman, K.J. | | Deposit date: | 2003-07-14 | | Release date: | 2004-02-24 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | The structure of docking domains in modular polyketide synthases.
Chem.Biol., 10, 2003
|
|
1PZQ
 
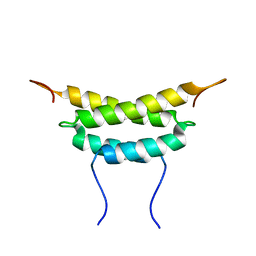 | | Structure of fused docking domains from the erythromycin polyketide synthase (DEBS), a model for the interaction between DEBS 2 and DEBS 3: The A domain | | Descriptor: | Erythronolide synthase | | Authors: | Broadhurst, R.W, Nietlispach, D, Wheatcroft, M.P, Leadlay, P.F, Weissman, K.J. | | Deposit date: | 2003-07-14 | | Release date: | 2004-02-24 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | The structure of docking domains in modular polyketide synthases.
Chem.Biol., 10, 2003
|
|
1J5D
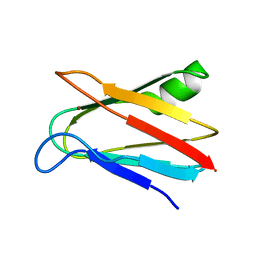 
 | | SOLUTION STRUCTURE OF OXIDIZED PARAMAGNETIC CU(II) PLASTOCYANIN FROM SYNECHOCYSTIS PCC6803-MINIMIZED AVERAGE STRUCTURE | | Descriptor: | COPPER (II) ION, PLASTOCYANIN | | Authors: | Bertini, I, Ciurli, S, Dikiy, A, Fernandez, C.O, Luchinat, C, Safarov, N, Shumilin, S, Vila, A.J. | | Deposit date: | 2002-04-02 | | Release date: | 2002-04-10 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | The first solution structure of a paramagnetic copper(II) protein: the case of oxidized plastocyanin from the cyanobacterium Synechocystis PCC6803.
J.Am.Chem.Soc., 123, 2001
|
|
1QTG
 
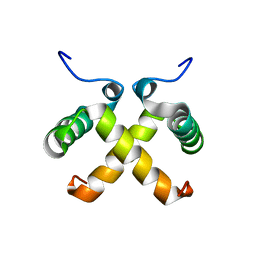 | | AVERAGED NMR MODEL OF SWITCH ARC, A DOUBLE MUTANT OF ARC REPRESSOR | | Descriptor: | Transcriptional repressor arc | | Authors: | Cordes, M.H.J, Walsh, N.P, McKnight, C.J, Sauer, R.T. | | Deposit date: | 1999-06-27 | | Release date: | 1999-07-12 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Evolution of a protein fold in vitro.
Science, 284, 1999
|
|
1NMF
 
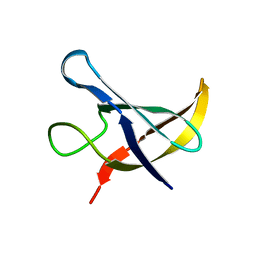 | |
1NTC
 
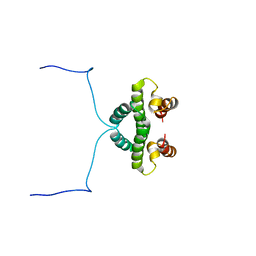 | |
1NMG
 
 | |
1NMJ
 
 | |
1Q3T
 
 | | Solution structure and function of an essential CMP kinase of Streptococcus pneumoniae | | Descriptor: | Cytidylate kinase | | Authors: | Yu, L, Mack, J, Hajduk, P.J, Kakavas, S.J, Saiki, A.Y, Lerner, C.G, Olejniczak, E.T. | | Deposit date: | 2003-07-31 | | Release date: | 2004-08-03 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution structure and function of an essential CMP kinase of Streptococcus pneumoniae
Protein Sci., 12, 2003
|
|
1QGB
 
 | | SOLUTION STRUCTURE OF THE N-TERMINAL F1 MODULE PAIR FROM HUMAN FIBRONECTIN | | Descriptor: | PROTEIN (FIBRONECTIN) | | Authors: | Potts, J.R, Bright, J.R, Bolton, D, Pickford, A.R, Campbell, I.D. | | Deposit date: | 1999-04-21 | | Release date: | 1999-12-08 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the N-terminal F1 module pair from human fibronectin.
Biochemistry, 38, 1999
|
|
1QCE
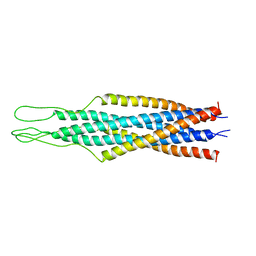 
 | |
2MZB
 
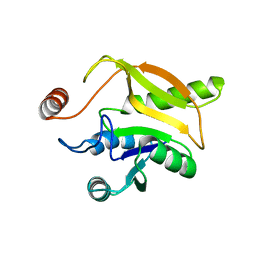 | | Solution structural studies of GTP:adenosylcobinamide-phosphate guanylyltransferase (CobY) from Methanocaldococcus jannaschii | | Descriptor: | Adenosylcobinamide-phosphate guanylyltransferase | | Authors: | Singarapu, K, Otte, M, Tonelli, M, Westler, W.M, Escalante-Semerena, J.C, Markley, J.L. | | Deposit date: | 2015-02-11 | | Release date: | 2015-11-11 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Solution Structural Studies of GTP:Adenosylcobinamide-Phosphateguanylyl Transferase (CobY) from Methanocaldococcus jannaschii.
Plos One, 10, 2015
|
|
1UC6
 
 | | Solution Structure of the Carboxyl Terminal Domain of the Ciliary Neurotrophic Factor Receptor | | Descriptor: | Ciliary Neurotrophic Factor Receptor alpha | | Authors: | Man, D, He, W, Sze, K.H, Ke, G, Smith, D.K, Ip, N.Y, Zhu, G. | | Deposit date: | 2003-04-08 | | Release date: | 2004-08-10 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the C-terminal domain of the ciliary neurotrophic factor (CNTF) receptor and ligand free associations among components of the CNTF receptor complex
J.Biol.Chem., 278, 2003
|
|
1S6W
 
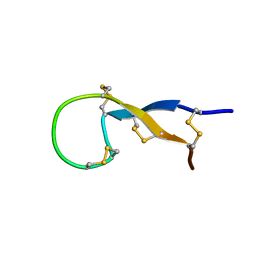 | | Solution Structure of hybrid white striped bass hepcidin | | Descriptor: | Hepcidin | | Authors: | Babon, J.J, Singh, S, Pennington, M.W, Norton, R.S, Westerman, M.E. | | Deposit date: | 2004-01-28 | | Release date: | 2004-12-14 | | Last modified: | 2022-03-02 | | Method: | SOLUTION NMR | | Cite: | Bass hepcidin synthesis, solution structure, antimicrobial activities and synergism, and in vivo hepatic response to bacterial infections.
J.Biol.Chem., 280, 2005
|
|
1T0Y
 
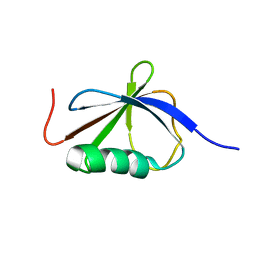 | | Solution Structure of a Ubiquitin-Like Domain from Tubulin-binding Cofactor B | | Descriptor: | tubulin folding cofactor B | | Authors: | Lytle, B.L, Peterson, F.C, Qui, S.H, Luo, M, Volkman, B.F, Markley, J.L, Center for Eukaryotic Structural Genomics (CESG) | | Deposit date: | 2004-04-13 | | Release date: | 2004-04-27 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of a Ubiquitin-like Domain from Tubulin-binding Cofactor B.
J.Biol.Chem., 279, 2004
|
|
1TTE
 
 | |
1T5N
 
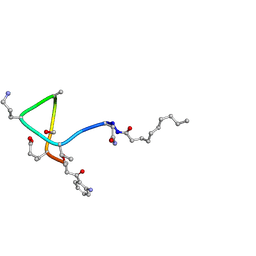 | | Structural transitions as determinants of calcium-dependent antibiotic daptomycin | | Descriptor: | DAPTOMYCIN, DECANOIC ACID | | Authors: | Jung, D, Rozek, A, Okon, M, Hancock, R.E. | | Deposit date: | 2004-05-04 | | Release date: | 2004-08-31 | | Last modified: | 2019-11-06 | | Method: | SOLUTION NMR | | Cite: | Structural Transitions as Determinants of the Action of the Calcium-Dependent Antibiotic Daptomycin.
Chem.Biol., 11, 2004
|
|
