3IOB
 
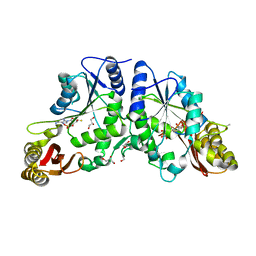 | | Crystal Structure of Mycobacterium Tuberculosis Pantothenate Synthetase at 1.80 Ang resolution in complex with 5'-deoxy-5'-thioadenosine | | Descriptor: | 5'-thioadenosine, ETHANOL, GLYCEROL, ... | | Authors: | Ciulli, A, Scott, D.E, Abell, C. | | Deposit date: | 2009-08-14 | | Release date: | 2009-12-01 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | A Fragment-Based Approach to Probing Adenosine Recognition Sites by Using Dynamic Combinatorial Chemistry
Chembiochem, 10, 2009
|
|
4Q1N
 
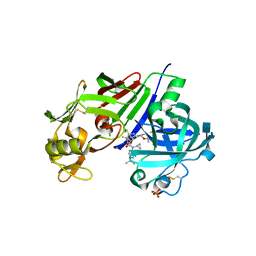 | | Structure-based design of 4-hydroxy-3,5-substituted piperidines as direct renin inhibitors | | Descriptor: | (3S,4R,5R)-N-cyclopropyl-N'-[(2R)-1-ethoxy-4-methylpentan-2-yl]-4-hydroxy-N-[5-(propan-2-yl)pyridin-2-yl]piperidine-3,5-dicarboxamide, 2-acetamido-2-deoxy-beta-D-glucopyranose, DIMETHYL SULFOXIDE, ... | | Authors: | Schiering, N, D'Arcy, A, Irie, O, Yokokawa, F. | | Deposit date: | 2014-04-04 | | Release date: | 2014-08-06 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (2.09 Å) | | Cite: | Structure-based design of substituted piperidines as a new class of highly efficacious oral direct Renin inhibitors.
ACS Med Chem Lett, 5, 2014
|
|
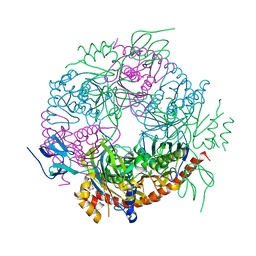
4BA1
 
 | |
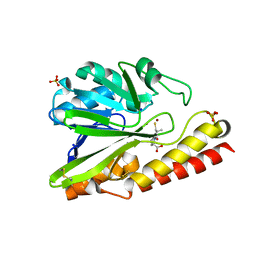
5AYA
 
 | |
5XUR
 
 | | Crystal Structure of Rv2466c C22S Mutant | | Descriptor: | 1,2-ETHANEDIOL, CHLORIDE ION, Thioredoxin-like reductase Rv2466c | | Authors: | Zhang, X, Li, H. | | Deposit date: | 2017-06-25 | | Release date: | 2018-03-14 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.996 Å) | | Cite: | Identification of a Mycothiol-Dependent Nitroreductase from Mycobacterium tuberculosis.
ACS Infect Dis, 4, 2018
|
|
6E99
 
 | |
2WB4
 
 | | activated diguanylate cyclase PleD in complex with c-di-GMP | | Descriptor: | 9,9'-[(2R,3R,3aS,5S,7aR,9R,10R,10aS,12S,14aR)-3,5,10,12-tetrahydroxy-5,12-dioxidooctahydro-2H,7H-difuro[3,2-d:3',2'-j][1,3,7,9,2,8]tetraoxadiphosphacyclododecine-2,9-diyl]bis(2-amino-1,9-dihydro-6H-purin-6-one), BERYLLIUM TRIFLUORIDE ION, DIGUANYLATE CYCLASE, ... | | Authors: | Wassmann, P, Schirmer, T. | | Deposit date: | 2009-02-20 | | Release date: | 2010-04-21 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Crystal Structure of Activated Pled, Identification of Dimerization and Catalysis Relevant Regulatory Mechanisms
To be Published
|
|
6KNL
 
 | |
5B6T
 
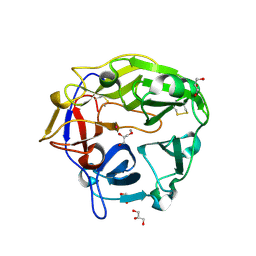 | | Catalytic domain of Coprinopsis cinerea GH62 alpha-L-arabinofuranosidase complexed with Pb | | Descriptor: | CALCIUM ION, GLYCEROL, Glycosyl hydrolase family 62 protein, ... | | Authors: | Tonozuka, T. | | Deposit date: | 2016-06-01 | | Release date: | 2016-09-07 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.48 Å) | | Cite: | Structure of the Catalytic Domain of alpha-L-Arabinofuranosidase from Coprinopsis cinerea, CcAbf62A, Provides Insights into Structure-Function Relationships in Glycoside Hydrolase Family 62
Appl. Biochem. Biotechnol., 181, 2017
|
|
5XRV
 
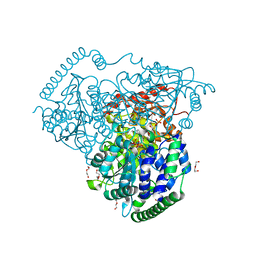 | | Crystal Structure of Transketolase in complex with TPP_V and fructose-6-phosphate from Pichia Stipitis | | Descriptor: | 2-[(5S)-3-[(4-azanyl-2-methyl-pyrimidin-5-yl)methyl]-4-methylidene-1,3-thiazolidin-5-yl]ethyl phosphono hydrogen phosphate, CALCIUM ION, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Li, T.L, Hsu, N.S, Wang, Y.L. | | Deposit date: | 2017-06-09 | | Release date: | 2018-06-13 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Evidence of Diradicals Involved in the Yeast Transketolase Catalyzed Keto-Transferring Reactions.
Chembiochem, 19, 2018
|
|
7E6C
 
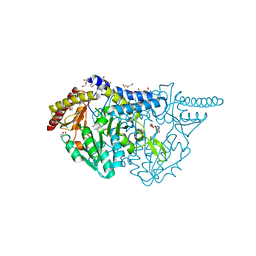 | | Crystal structure of L-cycloserine-bound form of cysteine desulfurase SufS C361A from Bacillus subtilis | | Descriptor: | (5-hydroxy-6-methyl-4-{[(3-oxo-2,3-dihydro-1,2-oxazol-4-yl)amino]methyl}pyridin-3-yl)methyl dihydrogen phosphate, 1,2-ETHANEDIOL, Cysteine desulfurase SufS, ... | | Authors: | Nakamura, R, Takahashi, Y, Fujishiro, T. | | Deposit date: | 2021-02-22 | | Release date: | 2022-03-02 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.73 Å) | | Cite: | Cycloserine enantiomers inhibit PLP-dependent cysteine desulfurase SufS via distinct mechanisms.
Febs J., 289, 2022
|
|
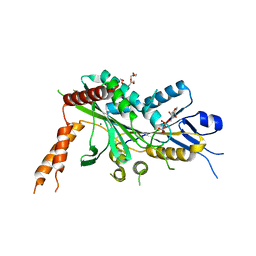
6A20
 | | Crystal Structure of auto-inhibited Kinesin-3 KIF13B | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, HEXAETHYLENE GLYCOL, Kinesin family member 13B, ... | | Authors: | Ren, J.Q, Wang, S, Feng, W. | | Deposit date: | 2018-06-08 | | Release date: | 2018-11-21 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Coiled-coil 1-mediated fastening of the neck and motor domains for kinesin-3 autoinhibition.
Proc. Natl. Acad. Sci. U.S.A., 115, 2018
|
|
3IGP
 
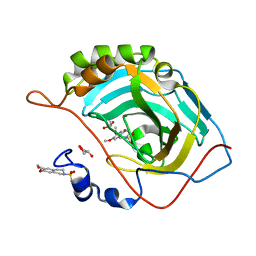 | | Structure of inhibitor binding to Carbonic Anhydrase II | | Descriptor: | 6,7-dimethoxy-3,4-dihydroisoquinoline-2(1H)-sulfonamide, Carbonic anhydrase 2, GLYCEROL, ... | | Authors: | Gitto, R, Agnello, S, Brynda, J, Mader, P, Supuran, C.T, Chimirri, A. | | Deposit date: | 2009-07-28 | | Release date: | 2010-03-09 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Identification of 3,4-Dihydroisoquinoline-2(1H)-sulfonamides as Potent Carbonic Anhydrase Inhibitors: Synthesis, Biological Evaluation, and Enzyme-Ligand X-ray Studies.
J.Med.Chem., 2010
|
|
4Q62
 
 | | Crystal Structure of Leucine-rich repeat- and Coiled coil-containing Protein from Legionella pneumophila | | Descriptor: | 1,2-ETHANEDIOL, Leucine-rich repeat-and coiled coil-containing protein, SULFATE ION | | Authors: | Kim, Y, Hatzos-Skintges, C, Jedrzejczak, R, Joachimiak, A, Midwest Center for Structural Genomics (MCSG), Program for the Characterization of Secreted Effector Proteins (PCSEP) | | Deposit date: | 2014-04-20 | | Release date: | 2014-05-07 | | Last modified: | 2024-11-27 | | Method: | X-RAY DIFFRACTION (1.898 Å) | | Cite: | Crystal Structure of Leucine-rich repeat- and Coiled coil-containing Protein from
Legionella pneumophila
To be Published
|
|
4XM7
 
 | | Anthrax toxin lethal factor with ligand-induced binding pocket | | Descriptor: | 1,2-ETHANEDIOL, Lethal factor, N~2~-[(4-fluoro-3-methoxyphenyl)sulfonyl]-N-hydroxy-N~2~-(2-methylpropyl)-D-valinamide, ... | | Authors: | Maize, K.M, Finzel, B.C. | | Deposit date: | 2015-01-14 | | Release date: | 2015-11-11 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Ligand-induced expansion of the S1' site in the anthrax toxin lethal factor.
Febs Lett., 589, 2015
|
|
3IKD
 
 | | Structure-Based Design of Novel PIN1 Inhibitors (I) | | Descriptor: | (2R)-2-[(1-benzothiophen-2-ylcarbonyl)amino]-3-phenylpropyl phosphate, Peptidyl-prolyl cis-trans isomerase NIMA-interacting 1 | | Authors: | Matthews, D, Greasley, S, Ferre, R, Parge, H. | | Deposit date: | 2009-08-05 | | Release date: | 2009-09-22 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure-based design of novel human Pin1 inhibitors (I).
Bioorg.Med.Chem.Lett., 19, 2009
|
|
5XN7
 
 | |
4XY9
 
 | | Crystal Structure of the first bromodomain of human BRD4 in complex with a 2-amine-9H-purine ligand | | Descriptor: | 1,2-ETHANEDIOL, 6-(5-bromo-2-methoxyphenyl)-9H-purin-2-amine, Bromodomain-containing protein 4 | | Authors: | Picaud, S, von Delft, F, Edwards, A.M, Arrowsmith, C.H, Bountra, C, Filippakopoulos, P, Structural Genomics Consortium (SGC) | | Deposit date: | 2015-02-02 | | Release date: | 2015-03-11 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.83 Å) | | Cite: | 9H-Purine Scaffold Reveals Induced-Fit Pocket Plasticity of the BRD9 Bromodomain.
J.Med.Chem., 58, 2015
|
|
3F5J
 
 | | Semi-active E176Q mutant of rice BGlu1, a plant exoglucanase/beta-glucosidase | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, Beta-glucosidase, SULFATE ION, ... | | Authors: | Chuenchor, W, Ketudat Cairns, J.R, Pengthaisong, S, Robinson, R.C, Yuvaniyama, J, Chen, C.-J. | | Deposit date: | 2008-11-03 | | Release date: | 2009-11-03 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | The structural basis of oligosaccharide binding by rice BGlu1 beta-glucosidase
J.Struct.Biol., 173, 2011
|
|
3IPB
 
 | | Human Transthyretin (TTR) complexed with a palindromic bivalent amyloid inhibitor (11 carbon linker). | | Descriptor: | 2,2'-{undecane-1,11-diylbis[oxy(3,5-dichlorobenzene-4,1-diyl)imino]}dibenzoic acid, Transthyretin | | Authors: | Kolstoe, S.E, Wood, S.P, Pepys, M.B. | | Deposit date: | 2009-08-17 | | Release date: | 2010-11-17 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Trapping of palindromic ligands within native transthyretin prevents amyloid formation.
Proc.Natl.Acad.Sci.USA, 107, 2010
|
|
2W1J
 
 | | CRYSTAL STRUCTURE OF SORTASE C-1 (SRTC-1) from Streptococcus pneumoniae | | Descriptor: | GLYCEROL, PUTATIVE SORTASE | | Authors: | Manzano, C, Contreras-Martel, C, El Mortaji, L, Izore, T, Fenel, D, Vernet, T, Schoehn, G, Di Guilmi, A.M, Dessen, A. | | Deposit date: | 2008-10-17 | | Release date: | 2008-12-23 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.24 Å) | | Cite: | Sortase-Mediated Pilus Fiber Biogenesis in Streptococcus Pneumoniae.
Structure, 16, 2008
|
|
3FFG
 
 | | Factor XA in complex with the inhibitor (R)-6-(2'-((3- HYDROXYPYRROLIDIN-1-YL)METHYL)BIPHENYL-4-YL)-1-(3-(5-OXO-4,5-DIHYDRO-1H-1,2,4-TRIAZOL-3-YL)PHENYL)-3-(TRIFLUOROMETHYL)-5,6-DIHYDRO-1H-PYRAZOLO[3,4-C]PYRIDIN- 7(4H)-ONE | | Descriptor: | (R)-6-(2'-((3-hydroxypyrrolidin-1-yl)methyl)biphenyl-4-yl)-1-(3-(5-oxo-4,5-dihydro-1h-1,2,4-triazol-3-yl)phenyl)-3-(trifluoromethyl)-5,6-dihydro-1h-pyrazolo[3,4-c]pyridin-7(4h)-one, Coagulation factor X, heavy chain, ... | | Authors: | Alexander, R.A. | | Deposit date: | 2008-12-03 | | Release date: | 2009-12-08 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (1.54 Å) | | Cite: | Phenyltriazolinones as potent factor Xa inhibitors.
Bioorg.Med.Chem.Lett., 20, 2010
|
|
5BST
 
 | |
3FGW
 
 | | One chain form of the 66.3 kDa protein | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, GLYCEROL, IODIDE ION, ... | | Authors: | Lakomek, K, Dickmanns, A, Ficner, R. | | Deposit date: | 2008-12-08 | | Release date: | 2009-09-15 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Initial insight into the function of the lysosomal 66.3 kDa protein from mouse by means of X-ray crystallography
Bmc Struct.Biol., 9, 2009
|
|
3IOE
 
 | | Crystal Structure of Mycobacterium Tuberculosis Pantothenate Synthetase at 1.95 Ang resolution in complex with 5'-deoxy-5'-((R)-3,4-dihydroxybutylthio)-adenosine | | Descriptor: | 5'-S-[(3R)-3,4-dihydroxybutyl]-5'-thioadenosine, ETHANOL, GLYCEROL, ... | | Authors: | Ciulli, A, Scott, D.E, Abell, C. | | Deposit date: | 2009-08-14 | | Release date: | 2009-12-01 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | A Fragment-Based Approach to Probing Adenosine Recognition Sites by Using Dynamic Combinatorial Chemistry
Chembiochem, 10, 2009
|
|
