1ZSE
 
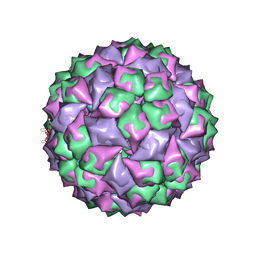 | | RNA stemloop from bacteriophage Qbeta complexed with an N87S mutant MS2 Capsid | | Descriptor: | Coat protein, RNA HAIRPIN | | Authors: | Horn, W.T, Tars, K, Grahn, E, Helgstrand, C, Baron, A.J, Lago, H, Adams, C.J, Peabody, D.S, Phillips, S.E.V, Stonehouse, N.J, Liljas, L, Stockley, P.G. | | Deposit date: | 2005-05-24 | | Release date: | 2006-05-09 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structural Basis of RNA Binding Discrimination between Bacteriophages Qbeta and MS2
Structure, 14, 2006
|
|
2KOC
 
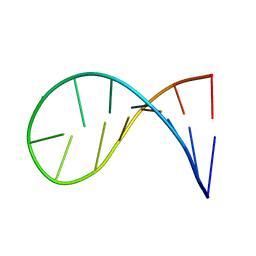 | | NMR solution structure of a 14-mer hairpin RNA with cUUCGg tetraloop | | Descriptor: | RNA (5'-R(P*GP*GP*CP*AP*CP*UP*UP*CP*GP*GP*UP*GP*CP*C)-3') | | Authors: | Nozinovic, S, Fuertig, B, Jonker, H.R.A, Richter, C, Schwalbe, H. | | Deposit date: | 2009-09-17 | | Release date: | 2009-12-01 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | High-resolution NMR structure of an RNA model system: the 14-mer cUUCGg tetraloop hairpin RNA
Nucleic Acids Res., 2009
|
|
2L1F
 
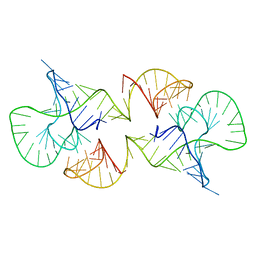 | | Structure of a conserved retroviral RNA packaging element by NMR spectroscopy and cryo-electron tomography | | Descriptor: | RNA (65-MER), RNA (66-MER) | | Authors: | Summers, M.F, Irobalieva, R.N, Tolbert, B, Smalls-Manty, A, Iyalla, K, Loeliger, K, D'Souza, V, Khant, H, Schmid, M, Garcia, E, Telesnitsky, A, Chiu, W, Miyazaki, Y. | | Deposit date: | 2010-07-28 | | Release date: | 2010-10-27 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Structure of a conserved retroviral RNA packaging element by NMR spectroscopy and cryo-electron tomography.
J.Mol.Biol., 404, 2010
|
|
2N2P
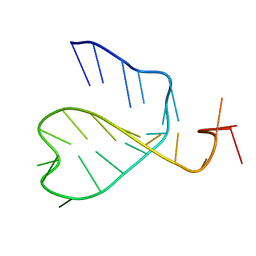 
 | | Solution structure of a double base-pair inversion mutant of murine tumour necrosis factor alpha CDE-23 RNA | | Descriptor: | RNA (5'-R(P*GP*CP*AP*UP*GP*UP*UP*UP*AP*GP*UP*GP*UP*CP*UP*AP*AP*AP*CP*GP*GP*UP*U)-3') | | Authors: | Codutti, L, Leppek, K, Zalesak, J, Windeisen, V, Masiewicz, P, Stoecklin, G, Carlomagno, T. | | Deposit date: | 2015-05-11 | | Release date: | 2015-08-05 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | A Distinct, Sequence-Induced Conformation Is Required for Recognition of the Constitutive Decay Element RNA by Roquin.
Structure, 23, 2015
|
|
2N2O
 
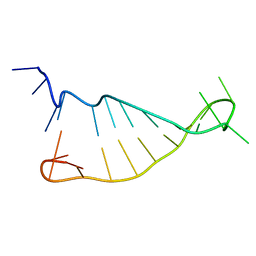 | | Structure of murine tumour necrosis factor alpha CDE RNA | | Descriptor: | RNA (5'-R(P*GP*CP*AP*UP*GP*UP*UP*UP*UP*CP*UP*GP*UP*GP*AP*AP*AP*AP*CP*GP*GP*UP*U)-3') | | Authors: | Codutti, L, Leppek, K, Zalesak, J, Windeisen, V, Masiewicz, P, Stoecklin, G, Carlomagno, T. | | Deposit date: | 2015-05-11 | | Release date: | 2015-08-05 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | A Distinct, Sequence-Induced Conformation Is Required for Recognition of the Constitutive Decay Element RNA by Roquin.
Structure, 23, 2015
|
|
1ELH
 
 | |
1F5U
 
 | |
1XSG
 
 | |
1R9F
 
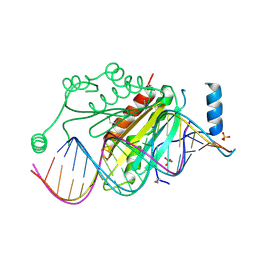 | | Crystal structure of p19 complexed with 19-bp small interfering RNA | | Descriptor: | 5'-R(*CP*GP*UP*AP*CP*GP*CP*GP*GP*AP*AP*UP*AP*CP*UP*UP*CP*GP*AP*UP*U)-3', 5'-R(*UP*CP*GP*AP*AP*GP*UP*AP*UP*UP*CP*CP*GP*CP*GP*UP*AP*CP*GP*UP*U)-3', Core protein P19, ... | | Authors: | Ye, K, Malinina, L, Patel, D.J. | | Deposit date: | 2003-10-28 | | Release date: | 2004-01-27 | | Last modified: | 2021-10-27 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Recognition of small interfering RNA by a viral suppressor of RNA
Nature, 426, 2003
|
|
1XSH
 
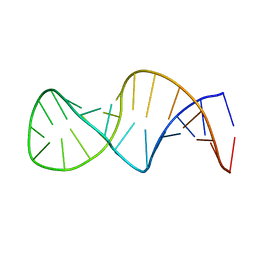 | |
299D
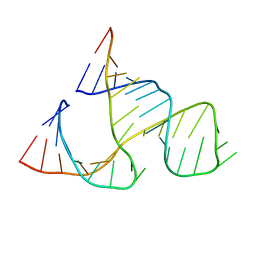 
 | | CAPTURING THE STRUCTURE OF A CATALYTIC RNA INTERMEDIATE: THE HAMMERHEAD RIBOZYME | | Descriptor: | RNA HAMMERHEAD RIBOZYME | | Authors: | Scott, W.G, Murray, J.B, Arnold, J.R.P, Stoddard, B.L, Klug, A. | | Deposit date: | 1996-12-14 | | Release date: | 1997-01-24 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Capturing the structure of a catalytic RNA intermediate: the hammerhead ribozyme.
Science, 274, 1996
|
|
3ELV
 
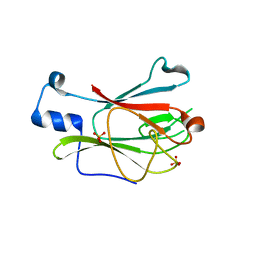 | | Crystal Structure of Full-Length Yeast Pml1p | | Descriptor: | Pre-mRNA leakage protein 1, SULFATE ION | | Authors: | Trowitzsch, S, Weber, G, L hrmann, R, Wahl, M.C. | | Deposit date: | 2008-09-23 | | Release date: | 2009-02-10 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal structure of the Pml1p subunit of the yeast precursor mRNA retention and splicing complex.
J.Mol.Biol., 385, 2009
|
|
6LSO
 
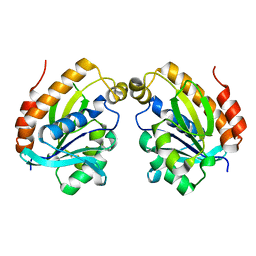 | |
7JRS
 
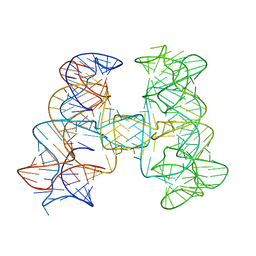 | |
7JRT
 
 | |
1S77
 
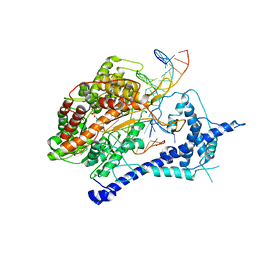 | | T7 RNAP product pyrophosphate elongation complex | | Descriptor: | DNA (5'-D(*GP*CP*CP*GP*TP*GP*CP*GP*CP*AP*TP*TP*CP*GP*CP*CP*GP*TP*GP*TP*T)-3'), DNA (5'-D(*TP*TP*TP*AP*CP*GP*TP*TP*GP*CP*GP*CP*AP*CP*GP*GP*C)-3'), DNA-directed RNA polymerase, ... | | Authors: | Yin, Y.W, Steitz, T.A. | | Deposit date: | 2004-01-29 | | Release date: | 2004-03-23 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.69 Å) | | Cite: | The structural mechanism of translocation and helicase activity in T7 RNA polymerase.
Cell(Cambridge,Mass.), 116, 2004
|
|
6Q8V
 
 | |
438D
 
 | |
255D
 
 | |
3PHP
 
 | | STRUCTURE OF THE 3' HAIRPIN OF THE TYMV PSEUDOKNOT: PREFORMATION IN RNA FOLDING | | Descriptor: | RNA (5'-R(*GP*GP*UP*UP*CP*CP*GP*AP*GP*GP*GP*UP*CP*AP*UP*CP*GP*GP*AP*AP*CP*CP*A) -3') | | Authors: | Kolk, M.H, Van Der Graaf, M, Wijmenga, S.S, Pleij, C.W.A, Heus, H.A, Hilbers, C.W. | | Deposit date: | 1998-10-13 | | Release date: | 1998-10-21 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | Structure of the 3'-hairpin of the TYMV pseudoknot: preformation in RNA folding.
EMBO J., 17, 1998
|
|
405D
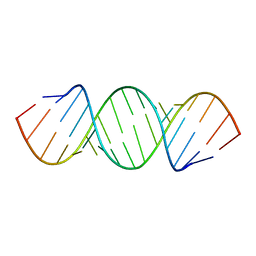 
 | |
1S76
 
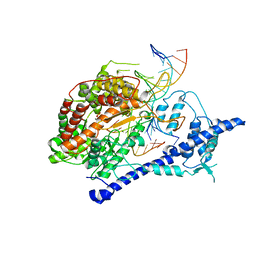 | | T7 RNA polymerase alpha beta methylene ATP elongation complex | | Descriptor: | DIPHOSPHOMETHYLPHOSPHONIC ACID ADENOSYL ESTER, DNA (5'-D(P*GP*CP*CP*GP*TP*GP*CP*GP*CP*AP*TP*TP*CP*GP*CP*CP*GP*TP*GP*TP*T)-3'), DNA (5'-D(P*TP*TP*TP*AP*CP*GP*TP*TP*GP*CP*GP*CP*AP*CP*GP*GP*C)-3'), ... | | Authors: | Yin, Y.W, Steitz, T.A. | | Deposit date: | 2004-01-29 | | Release date: | 2004-03-23 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.88 Å) | | Cite: | The structural mechanism of translocation and helicase activity in T7 RNA polymerase.
Cell(Cambridge,Mass.), 116, 2004
|
|
5C5W
 
 | | 1.25 A resolution structure of an RNA 20-mer | | Descriptor: | RNA (5'-R(P*CP*CP*UP*GP*AP*GP*UP*UP*CP*AP*AP*UP*UP*CP*UP*AP*GP*CP*G)-3') | | Authors: | Stewart, M, Valkov, E. | | Deposit date: | 2015-06-22 | | Release date: | 2015-10-14 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.25 Å) | | Cite: | 1.25 angstrom resolution structure of an RNA 20-mer that binds to the TREX2 complex.
Acta Crystallogr.,Sect.F, 71, 2015
|
|
1L3D
 
 | |
2E7Y
 
 | | High resolution structure of T. maritima tRNase Z | | Descriptor: | S-1,2-PROPANEDIOL, SULFATE ION, ZINC ION, ... | | Authors: | Ishii, R, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2007-01-15 | | Release date: | 2007-09-11 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.97 Å) | | Cite: | The structure of the flexible arm of Thermotoga maritima tRNase Z differs from those of homologous enzymes
Acta Crystallogr.,Sect.F, 63, 2007
|
|
