7ECO
 
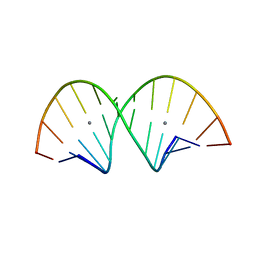 | | RNA duplex containing U-Ag-U base pairs | | Descriptor: | RNA (5'-R(*GP*GP*GP*UP*CP*CP*GP*GP*UP*CP*CP*C)-3'), SILVER ION | | Authors: | Kondo, J, Tsudura, A. | | Deposit date: | 2021-03-12 | | Release date: | 2022-03-16 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (1.81 Å) | | Cite: | RNA duplex containing U-Ag-U base pairs
To Be Published
|
|
7ECP
 
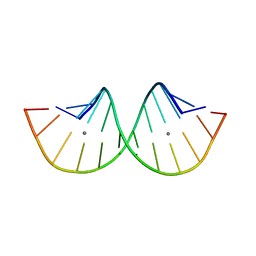 | | RNA duplex containing C-Ag-U | | Descriptor: | RNA (5'-R(*GP*GP*GP*CP*CP*CP*GP*GP*UP*CP*CP*C)-3'), SILVER ION | | Authors: | Kondo, J, Tsudura, A. | | Deposit date: | 2021-03-12 | | Release date: | 2022-03-16 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (2.91 Å) | | Cite: | RNA duplex containing C-Ag-U
To Be Published
|
|
7ECN
 
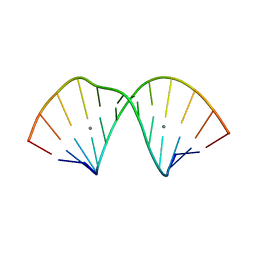 | | RNA duplex containing C-Ag-A base pairs | | Descriptor: | RNA (5'-R(*GP*GP*GP*CP*CP*CP*GP*GP*AP*CP*CP*C)-3'), SILVER ION | | Authors: | Kondo, J, Tsudura, A. | | Deposit date: | 2021-03-12 | | Release date: | 2022-03-16 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | RNA duplex containing C-Ag-A base pairs
To Be Published
|
|
7EDT
 
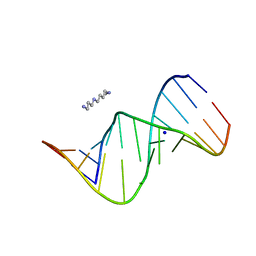 | | RNA duplex containing CC mispairs | | Descriptor: | RNA (5'-R(*GP*GP*AP*CP*UP*CP*CP*GP*GP*(5BU)P*CP*C)-3'), SODIUM ION, SPERMINE | | Authors: | Kondo, J, Tada, Y. | | Deposit date: | 2021-03-17 | | Release date: | 2022-03-23 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | RNA duplex containing CC mispairs
To Be Published
|
|
7EFH
 
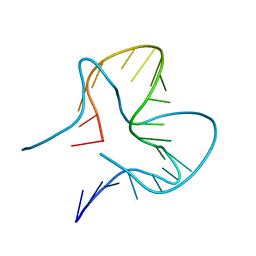 | | RNA kink-turn motif | | Descriptor: | RNA (5'-R(*GP*GP*CP*GP*AP*AP*GP*AP*AP*CP*CP*GP*GP*GP*GP*AP*GP*CP*C)-3') | | Authors: | Kondo, J, Nagashima, M. | | Deposit date: | 2021-03-21 | | Release date: | 2022-03-23 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | RNA kink-turn motif
To Be Published
|
|
7EFG
 
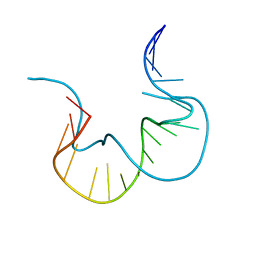 | | RNA kink-turn motif | | Descriptor: | RNA (5'-R(*GP*GP*CP*GP*AP*AP*GP*AP*AP*CP*CP*GP*GP*GP*GP*AP*GP*CP*C)-3') | | Authors: | Kondo, J, Nagashima, M. | | Deposit date: | 2021-03-21 | | Release date: | 2022-04-13 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | RNA kink-turn motif
To Be Published
|
|
6EC8
 
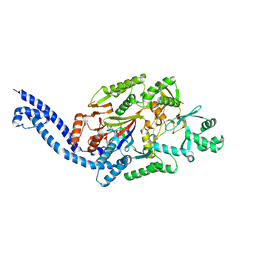 | |
4HI6
 
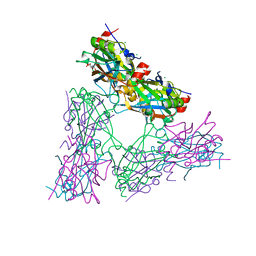 | |
4HIY
 | |
4HIU
 
 | |
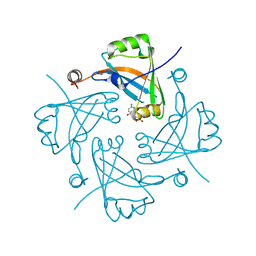
1WDV
 
 | | Crystal structure of hypothetical protein APE2540 | | Descriptor: | hypothetical protein APE2540 | | Authors: | Murayama, K, Kato-Murayama, M, Shirouzu, M, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2004-05-18 | | Release date: | 2004-11-18 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structure of a putative trans-editing enzyme for prolyl-tRNA synthetase from Aeropyrum pernix K1 at 1.7 A resolution.
Acta Crystallogr.,Sect.F, 61, 2005
|
|
4HIW
 
 | |
4HI5
 | |
4HIT
 
 | |
5VGW
 
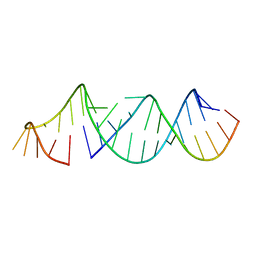 | | RNA hairpin structure containing tetraloop/receptor motif | | Descriptor: | RNA (5'-R(*CP*UP*GP*CP*UP*GP*GP*CP*UP*AP*AP*GP*GP*CP*CP*CP*GP*AP*AP*AP*GP*G)-3'), RNA (5'-R(P*GP*CP*UP*AP*UP*GP*CP*CP*UP*GP*CP*UP*G)-3') | | Authors: | Zhang, W, Szostak, J.W. | | Deposit date: | 2017-04-11 | | Release date: | 2018-02-21 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.42 Å) | | Cite: | Structural Rationale for the Enhanced Catalysis of Nonenzymatic RNA Primer Extension by a Downstream Oligonucleotide.
J. Am. Chem. Soc., 140, 2018
|
|
5UZ6
 
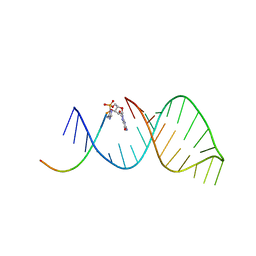 | | RNA hairpin structure containing 2-MeImp-oligo analogue | | Descriptor: | RNA (25-MER), RNA (5'-D(*(8OS))-R(P*CP*AP*CP*CP*UP*CP*A)-3') | | Authors: | Zhang, W, Oh, S.S, Szostak, J.W. | | Deposit date: | 2017-02-24 | | Release date: | 2018-02-21 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structural Rationale for the Enhanced Catalysis of Nonenzymatic RNA Primer Extension by a Downstream Oligonucleotide.
J. Am. Chem. Soc., 140, 2018
|
|
4WKM
 
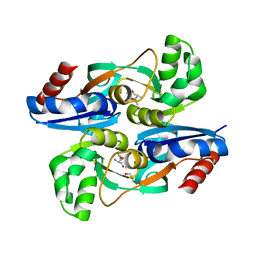 | | AmpR effector binding domain from Citrobacter freundii bound to UDP-MurNAc-pentapeptide | | Descriptor: | ALA-FGA-API-DAL-DAL, GLYCEROL, LysR family transcriptional regulator, ... | | Authors: | Vadlamani, G, Reeve, T.M, Mark, B.L. | | Deposit date: | 2014-10-02 | | Release date: | 2014-12-17 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | The beta-Lactamase Gene Regulator AmpR Is a Tetramer That Recognizes and Binds the d-Ala-d-Ala Motif of Its Repressor UDP-N-acetylmuramic Acid (MurNAc)-pentapeptide.
J.Biol.Chem., 290, 2015
|
|
7BYY
 
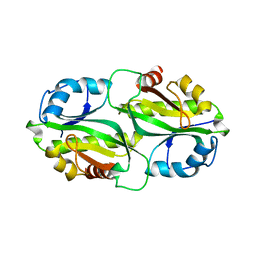 | | Crystal structure of bacterial toxin | | Descriptor: | Acetyltransferase | | Authors: | Zhang, C, Yashiro, Y, Tomita, K. | | Deposit date: | 2020-04-25 | | Release date: | 2020-06-03 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.799 Å) | | Cite: | Substrate specificities of Escherichia coli ItaT that acetylates aminoacyl-tRNAs.
Nucleic Acids Res., 48, 2020
|
|
9BLJ
 | |
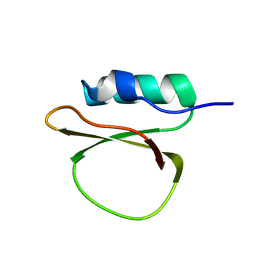
1RZW
 
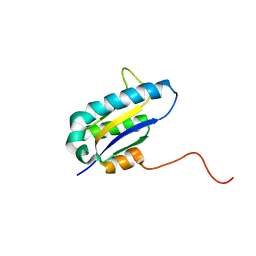 | | The Solution Structure of the Archaeglobus fulgidis protein AF2095. Northeast Structural Genomics Consortium target GR4 | | Descriptor: | Protein AF2095(GR4) | | Authors: | Powers, R, Acton, T.B, Huang, Y.J, Liu, J, Ma, L, Rost, B, Chiang, Y, Cort, J.R, Kennedy, M.A, Montelione, G.T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2003-12-29 | | Release date: | 2004-11-16 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution structure of Archaeglobus fulgidis peptidyl-tRNA hydrolase (Pth2) provides evidence for an extensive conserved family of Pth2 enzymes in archea, bacteria, and eukaryotes
Protein Sci., 14, 2005
|
|
8VU8
 
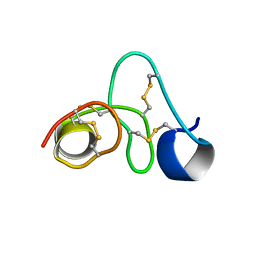 | |
8VU7
 
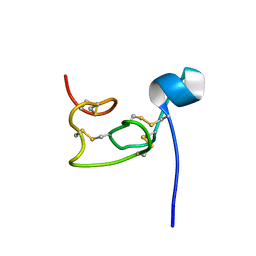 | |
8VU6
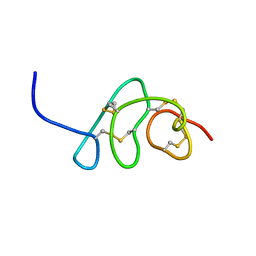 
 | |
8CE4
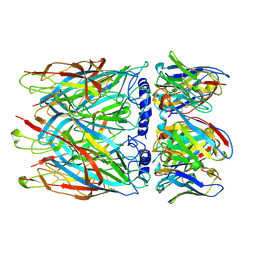 
 | | Human alpha7 nicotinic receptor in complex with the E3 nanobody | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Nanobody E3, ... | | Authors: | Prevost, M.S, Barilone, N, Dejean de la Batie, G, Pons, S, Ayme, G, England, P, Gielen, M, Bontems, F, Pehau-Arnaudet, G, Maskos, U, Lafaye, P, Corringer, P.-J. | | Deposit date: | 2023-02-01 | | Release date: | 2023-10-11 | | Last modified: | 2025-07-09 | | Method: | ELECTRON MICROSCOPY (2.7 Å) | | Cite: | An original potentiating mechanism revealed by the cryo-EM structures of the human alpha 7 nicotinic receptor in complex with nanobodies.
Nat Commun, 14, 2023
|
|
8C9X
 
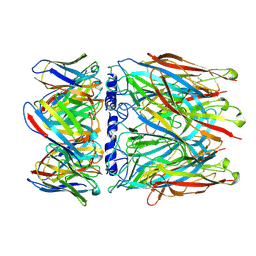 | | human alpha7 nicotinic receptor in complex with the C4 nanobody | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Nanobody C4, ... | | Authors: | Prevost, M.S, Barilone, N, Dejean de la Batie, G, Pons, S, Ayme, G, England, P, Gielen, M, Bontems, F, Pehau-Arnaudet, G, Maskos, U, Lafaye, P, Corringer, P.-J. | | Deposit date: | 2023-01-23 | | Release date: | 2023-10-11 | | Last modified: | 2025-07-02 | | Method: | ELECTRON MICROSCOPY (2.3 Å) | | Cite: | An original potentiating mechanism revealed by the cryo-EM structures of the human alpha 7 nicotinic receptor in complex with nanobodies.
Nat Commun, 14, 2023
|
|
