1C9T
 
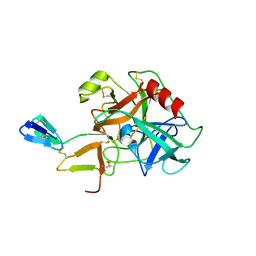 | | COMPLEX OF BDELLASTASIN WITH BOVINE TRYPSIN | | Descriptor: | BDELLASTASIN, TRYPSIN | | Authors: | Rester, U, Bode, W, Moser, M, Parry, M.A, Huber, R, Auerswald, E. | | Deposit date: | 1999-08-03 | | Release date: | 2000-08-03 | | Last modified: | 2017-10-04 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | Structure of the complex of the antistasin-type inhibitor bdellastasin with trypsin and modelling of the bdellastasin-microplasmin system.
J.Mol.Biol., 293, 1999
|
|
1C9U
 
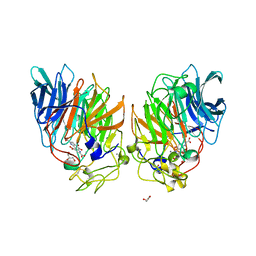 | | CRYSTAL STRUCTURE OF THE SOLUBLE QUINOPROTEIN GLUCOSE DEHYDROGENASE IN COMPLEX WITH PQQ | | Descriptor: | CALCIUM ION, GLYCEROL, PYRROLOQUINOLINE QUINONE, ... | | Authors: | Oubrie, A, Rozeboom, H.J, Dijkstra, B.W. | | Deposit date: | 1999-08-03 | | Release date: | 2000-02-04 | | Last modified: | 2019-08-14 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structure and mechanism of soluble quinoprotein glucose dehydrogenase.
EMBO J., 18, 1999
|
|
1C9V
 
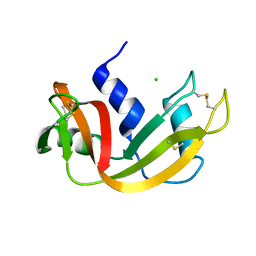 | | H12A VARIANT OF RIBONUCLEASE A | | Descriptor: | CHLORIDE ION, RIBONUCLEASE A | | Authors: | Park, C, Schultz, L.W, Raines, R.T. | | Deposit date: | 1999-08-03 | | Release date: | 2001-06-27 | | Last modified: | 2021-11-03 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Contribution of the active site histidine residues of ribonuclease A to nucleic acid binding.
Biochemistry, 40, 2001
|
|
1C9W
 
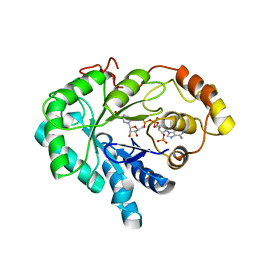 | | CHO REDUCTASE WITH NADP+ | | Descriptor: | CHO REDUCTASE, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE | | Authors: | Ye, Q, Li, X, Hyndman, D, Flynn, T.G, Jia, Z. | | Deposit date: | 1999-08-03 | | Release date: | 2000-01-07 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal structure of CHO reductase, a member of the aldo-keto reductase superfamily.
Proteins, 38, 2000
|
|
1C9X
 
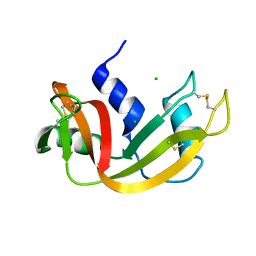 | | H119A VARIANT OF RIBONUCLEASE A | | Descriptor: | CHLORIDE ION, RIBONUCLEASE A | | Authors: | Park, C, Schultz, L.W, Raines, R.T. | | Deposit date: | 1999-08-03 | | Release date: | 2001-06-27 | | Last modified: | 2021-11-03 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Contribution of the active site histidine residues of ribonuclease A to nucleic acid binding.
Biochemistry, 40, 2001
|
|
1C9Y
 
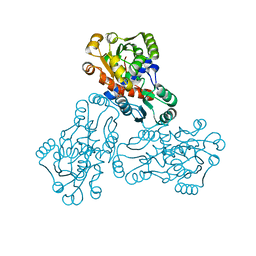 | | HUMAN ORNITHINE TRANSCARBAMYLASE: CRYSTALLOGRAPHIC INSIGHTS INTO SUBSTRATE RECOGNITION AND CATALYTIC MECHANISM | | Descriptor: | NORVALINE, ORNITHINE CARBAMOYLTRANSFERASE, PHOSPHORIC ACID MONO(FORMAMIDE)ESTER | | Authors: | Shi, D, Yu, X, Morizono, H, Tuchman, M, Allewell, N.M. | | Deposit date: | 1999-08-03 | | Release date: | 2000-06-06 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structure of human ornithine transcarbamylase complexed with carbamoyl phosphate and L-norvaline at 1.9 A resolution.
Proteins, 39, 2000
|
|
1C9Z
 
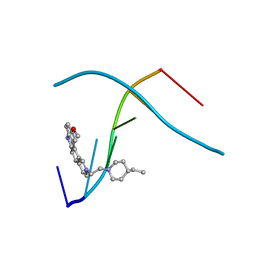 | | D232-CGTACG | | Descriptor: | 1,3-DI[[[10-METHOXY-7H-PYRIDO[4,3-C]CARBAZOL-2-IUMYL]-ETHYL]-PIPERIDIN-4-YL]-PROPANE, 5'-D(*CP*GP*TP*AP*CP*G)-3' | | Authors: | Williams, L.D. | | Deposit date: | 1999-08-03 | | Release date: | 2000-04-02 | | Last modified: | 2024-04-17 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Effects of cationic charge on three-dimensional structures of intercalative complexes: structure of a bis-intercalated DNA complex solved by MAD phasing.
Curr.Med.Chem., 7, 2000
|
|
1CA0
 
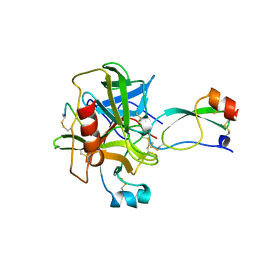 | | BOVINE CHYMOTRYPSIN COMPLEXED TO APPI | | Descriptor: | BOVINE CHYMOTRYPSIN, PROTEASE INHIBITOR DOMAIN OF ALZHEIMER'S AMYLOID BETA-PROTEIN PRECURSOR | | Authors: | Scheidig, A.J, Kossiakoff, A.A. | | Deposit date: | 1997-01-23 | | Release date: | 1997-07-23 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal structures of bovine chymotrypsin and trypsin complexed to the inhibitor domain of Alzheimer's amyloid beta-protein precursor (APPI) and basic pancreatic trypsin inhibitor (BPTI): engineering of inhibitors with altered specificities.
Protein Sci., 6, 1997
|
|
1CA1
 
 | |
1CA2
 
 | |
1CA3
 
 | |
1CA4
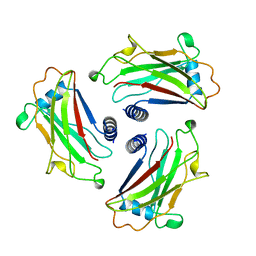 
 | | STRUCTURE OF TNF RECEPTOR ASSOCIATED FACTOR 2 (TRAF2) | | Descriptor: | PROTEIN (TNF RECEPTOR ASSOCIATED FACTOR 2) | | Authors: | Park, Y.C, Burkitt, V, Villa, A.R, Tong, L, Wu, H. | | Deposit date: | 1999-02-23 | | Release date: | 1999-04-12 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structural basis for self-association and receptor recognition of human TRAF2.
Nature, 398, 1999
|
|
1CA5
 
 | | INTERCALATION SITE OF HYPERTHERMOPHILE CHROMOSOMAL PROTEIN SSO7D/SAC7D BOUND TO DNA | | Descriptor: | 5'-D(*GP*TP*GP*AP*TP*CP*AP*C)-3', CHROMOSOMAL PROTEIN SAC7D | | Authors: | Su, S, Gao, Y.-G, Robinson, H, Shriver, J.W, Wang, A.H.-J. | | Deposit date: | 1999-02-23 | | Release date: | 2000-02-23 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structures of the chromosomal proteins Sso7d/Sac7d bound to DNA containing T-G mismatched base-pairs
J.Mol.Biol., 303, 2000
|
|
1CA6
 | | INTERCALATION SITE OF HYPERTHERMOPHILE CHROMOSOMAL PROTEIN SSO7D/SAC7D BOUND TO DNA | | Descriptor: | 5'-D(*GP*TP*GP*AP*TP*CP*GP*C)-3', CHROMOSOMAL PROTEIN SAC7D | | Authors: | Su, S, Gao, Y.-G, Robinson, H, Shriver, J.W, Wang, A.H.-J. | | Deposit date: | 1999-02-23 | | Release date: | 2000-02-23 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structures of the chromosomal proteins Sso7d/Sac7d bound to DNA containing T-G mismatched base-pairs
J.Mol.Biol., 303, 2000
|
|
1CA7
 
 | |
1CA8
 
 | | Thrombin inhibitors with rigid tripeptidyl aldehydes | | Descriptor: | 2-{(3S)-3-[(benzylsulfonyl)amino]-2-oxopiperidin-1-yl}-N-{(2S)-1-[(3S)-1-carbamimidoylpiperidin-3-yl]-3-oxopropan-2-yl}acetamide, HIRUGEN, SODIUM ION, ... | | Authors: | Krishnan, R, Zhang, E, Hakansson, K, Arni, R.K, Tulinsky, A, Lim-Wilby, M.S.L, Levy, O.E, Semple, J.E, Brunck, T.K. | | Deposit date: | 1998-04-27 | | Release date: | 1999-04-27 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Highly selective mechanism-based thrombin inhibitors: structures of thrombin and trypsin inhibited with rigid peptidyl aldehydes
Biochemistry, 37, 1998
|
|
1CA9
 
 | | STRUCTURE OF TNF RECEPTOR ASSOCIATED FACTOR 2 IN COMPLEX WITH A PEPTIDE FROM TNF-R2 | | Descriptor: | PROTEIN (TNF RECEPTOR ASSOCIATED FACTOR 2), PROTEIN (TNF-R2) | | Authors: | Park, Y.C, Burkitt, V, Villa, A.R, Tong, L, Wu, H. | | Deposit date: | 1999-02-25 | | Release date: | 1999-04-12 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structural basis for self-association and receptor recognition of human TRAF2.
Nature, 398, 1999
|
|
1CAA
 
 | | X-RAY CRYSTAL STRUCTURES OF THE OXIDIZED AND REDUCED FORMS OF THE RUBREDOXIN FROM THE MARINE HYPERTHERMOPHILIC ARCHAEBACTERIUM PYROCOCCUS FURIOSUS | | Descriptor: | FE (III) ION, RUBREDOXIN | | Authors: | Day, M.W, Hsu, B.T, Joshua-Tor, L, Park, J.B, Zhou, Z.H, Adams, M.W.W, Rees, D.C. | | Deposit date: | 1992-05-18 | | Release date: | 1993-10-31 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | X-ray crystal structures of the oxidized and reduced forms of the rubredoxin from the marine hyperthermophilic archaebacterium Pyrococcus furiosus.
Protein Sci., 1, 1992
|
|
1CAD
 
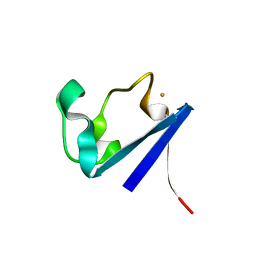 | | X-RAY CRYSTAL STRUCTURES OF THE OXIDIZED AND REDUCED FORMS OF THE RUBREDOXIN FROM THE MARINE HYPERTHERMOPHILIC ARCHAEBACTERIUM PYROCOCCUS FURIOSUS | | Descriptor: | FE (III) ION, RUBREDOXIN | | Authors: | Day, M.W, Hsu, B.T, Joshua-Tor, L, Park, J.B, Zhou, Z.H, Adams, M.W.W, Rees, D.C. | | Deposit date: | 1992-05-18 | | Release date: | 1993-10-31 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | X-ray crystal structures of the oxidized and reduced forms of the rubredoxin from the marine hyperthermophilic archaebacterium Pyrococcus furiosus.
Protein Sci., 1, 1992
|
|
1CAG
 
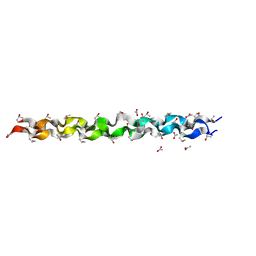 | | CRYSTAL AND MOLECULAR STRUCTURE OF A COLLAGEN-LIKE PEPTIDE AT 1.9 ANGSTROM RESOLUTION | | Descriptor: | ACETIC ACID, COLLAGEN-LIKE PEPTIDE | | Authors: | Bella, J, Eaton, M, Brodsky, B, Berman, H.M. | | Deposit date: | 1994-03-29 | | Release date: | 1994-11-01 | | Last modified: | 2024-06-05 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Crystal and molecular structure of a collagen-like peptide at 1.9 A resolution.
Science, 266, 1994
|
|
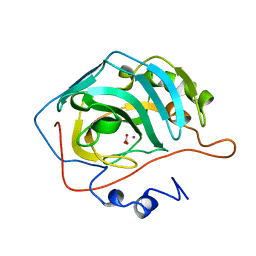
1CAH
 
 | |
1CAI
 
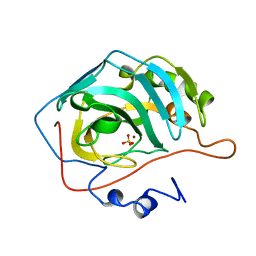 | | STRUCTURAL ANALYSIS OF THE ZINC HYDROXIDE-THR 199-GLU 106 HYDROGEN BONDING NETWORK IN HUMAN CARBONIC ANHYDRASE II | | Descriptor: | CARBONIC ANHYDRASE II, SULFATE ION, ZINC ION | | Authors: | Xue, Y, Liljas, A, Jonsson, B.-H, Lindskog, S. | | Deposit date: | 1992-09-17 | | Release date: | 1993-10-31 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural analysis of the zinc hydroxide-Thr-199-Glu-106 hydrogen-bond network in human carbonic anhydrase II.
Proteins, 17, 1993
|
|
1CAJ
 
 | | STRUCTURAL ANALYSIS OF THE ZINC HYDROXIDE-THR 199-GLU 106 HYDROGEN BONDING NETWORK IN HUMAN CARBONIC ANHYDRASE II | | Descriptor: | CARBONIC ANHYDRASE II, SULFATE ION, ZINC ION | | Authors: | Xue, Y, Liljas, A, Jonsson, B.-H, Lindskog, S. | | Deposit date: | 1992-09-17 | | Release date: | 1993-10-31 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural analysis of the zinc hydroxide-Thr-199-Glu-106 hydrogen-bond network in human carbonic anhydrase II.
Proteins, 17, 1993
|
|
1CAK
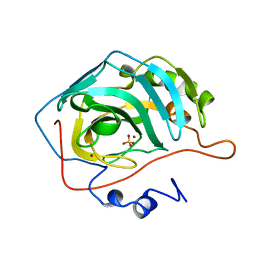 
 | | STRUCTURAL ANALYSIS OF THE ZINC HYDROXIDE-THR 199-GLU 106 HYDROGEN BONDING NETWORK IN HUMAN CARBONIC ANHYDRASE II | | Descriptor: | CARBONIC ANHYDRASE II, SULFATE ION, ZINC ION | | Authors: | Xue, Y, Liljas, A, Jonsson, B.-H, Lindskog, S. | | Deposit date: | 1992-09-17 | | Release date: | 1993-10-31 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural analysis of the zinc hydroxide-Thr-199-Glu-106 hydrogen-bond network in human carbonic anhydrase II.
Proteins, 17, 1993
|
|
1CAL
 
 | | STRUCTURAL ANALYSIS OF THE ZINC HYDROXIDE-THR 199-GLU 106 HYDROGEN BONDING NETWORK IN HUMAN CARBONIC ANHYDRASE II | | Descriptor: | CARBONIC ANHYDRASE II, ZINC ION | | Authors: | Xue, Y, Liljas, A, Jonsson, B.-H, Lindskog, S. | | Deposit date: | 1992-09-17 | | Release date: | 1993-10-31 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structural analysis of the zinc hydroxide-Thr-199-Glu-106 hydrogen-bond network in human carbonic anhydrase II.
Proteins, 17, 1993
|
|
