9CR2
 
 | | TRiC-ADP-C1 | | Descriptor: | T-complex protein 1 subunit alpha, T-complex protein 1 subunit beta, T-complex protein 1 subunit delta, ... | | Authors: | Jin, M, Cong, Y. | | Deposit date: | 2024-07-20 | | Release date: | 2025-01-22 | | Last modified: | 2025-05-28 | | Method: | ELECTRON MICROSCOPY (4.8 Å) | | Cite: | The conformational landscape of TRiC ring-opening and its underlying stepwise mechanism revealed by cryo-EM.
Qrb Discov, 6, 2025
|
|
9HTA
 
 | | Dispersin from Terribacillus saccharophilus Dispts2 in complex with NAG-thiazoline | | Descriptor: | 3AR,5R,6S,7R,7AR-5-HYDROXYMETHYL-2-METHYL-5,6,7,7A-TETRAHYDRO-3AH-PYRANO[3,2-D]THIAZOLE-6,7-DIOL, Dispersin | | Authors: | Males, A, Moroz, O.V, Blagova, E, Munch, A, Hansen, G.H, Johansen, A.H, Ostergaard, L.H, Segura, D.R, Eddenden, A, Due, A.V, Gudmand, M, Salomon, J, Sorensen, S.R, Franco Cairo, J.P.L, Pache, R.A, Vejborg, R.M, Bhosale, S, Vocadlo, D, Davies, G.J, Wilson, K.S. | | Deposit date: | 2024-12-19 | | Release date: | 2025-03-12 | | Last modified: | 2025-03-19 | | Method: | X-RAY DIFFRACTION (2.17 Å) | | Cite: | Expansion of the diversity of dispersin scaffolds.
Acta Crystallogr D Struct Biol, 81, 2025
|
|
9CSA
 
 | | TRiC-ATP-AlFx | | Descriptor: | T-complex protein 1 subunit alpha, T-complex protein 1 subunit beta, T-complex protein 1 subunit delta, ... | | Authors: | Jin, M, Cong, Y. | | Deposit date: | 2024-07-23 | | Release date: | 2025-01-22 | | Last modified: | 2025-06-04 | | Method: | ELECTRON MICROSCOPY (3.6 Å) | | Cite: | The conformational landscape of TRiC ring-opening and its underlying stepwise mechanism revealed by cryo-EM.
Qrb Discov, 6, 2025
|
|
9CS4
 
 | | TRiC-ADP-S3 | | Descriptor: | T-complex protein 1 subunit alpha, T-complex protein 1 subunit beta, T-complex protein 1 subunit delta, ... | | Authors: | Jin, M, Cong, Y. | | Deposit date: | 2024-07-23 | | Release date: | 2025-01-22 | | Last modified: | 2025-05-21 | | Method: | ELECTRON MICROSCOPY (6.8 Å) | | Cite: | The conformational landscape of TRiC ring-opening and its underlying stepwise mechanism revealed by cryo-EM.
Qrb Discov, 6, 2025
|
|
9CS6
 
 | | TRiC-ADP-S5 state is a conformation when TRiC incubated in 1 mM ADP | | Descriptor: | T-complex protein 1 subunit alpha, T-complex protein 1 subunit beta, T-complex protein 1 subunit delta, ... | | Authors: | Jin, M, Cong, Y. | | Deposit date: | 2024-07-23 | | Release date: | 2025-01-22 | | Last modified: | 2025-03-26 | | Method: | ELECTRON MICROSCOPY (4.1 Å) | | Cite: | The conformational landscape of TRiC ring-opening and its underlying stepwise mechanism revealed by cryo-EM.
Qrb Discov, 6, 2025
|
|
9CS3
 
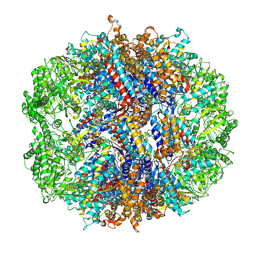 | | TRiC-ADP-S2 state is a conformation when TRiC incubated in 1 mM ADP | | Descriptor: | T-complex protein 1 subunit alpha, T-complex protein 1 subunit beta, T-complex protein 1 subunit delta, ... | | Authors: | Jin, M, Cong, Y. | | Deposit date: | 2024-07-23 | | Release date: | 2025-01-22 | | Last modified: | 2025-05-21 | | Method: | ELECTRON MICROSCOPY (5.6 Å) | | Cite: | The conformational landscape of TRiC ring-opening and its underlying stepwise mechanism revealed by cryo-EM.
Qrb Discov, 6, 2025
|
|
7OXO
 
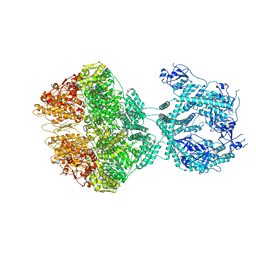 | | human LonP1, R-state, incubated in AMPPCP | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, Lon protease homolog, mitochondrial | | Authors: | Abrahams, J.P, Mohammed, I, Schmitz, K.A, Schenck, N, Maier, T. | | Deposit date: | 2021-06-22 | | Release date: | 2021-12-22 | | Last modified: | 2024-07-17 | | Method: | ELECTRON MICROSCOPY (3.9 Å) | | Cite: | Catalytic cycling of human mitochondrial Lon protease.
Structure, 30, 2022
|
|
9IOL
 
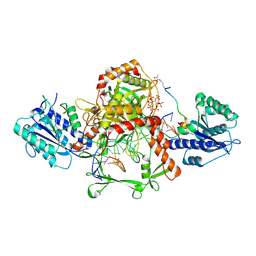 | | Cryo-EM structure of the complex of DNA, Ku70/80, and laXLF. | | Descriptor: | (2S)-2-HYDROXYPROPANOIC ACID, DNA (5'-D(P*AP*GP*GP*TP*CP*GP*AP*CP*GP*AP*AP*TP*CP*GP*GP*CP*AP*GP*CP*G)-3'), DNA (5'-D(P*CP*GP*CP*TP*GP*CP*CP*GP*AP*TP*TP*CP*GP*TP*CP*GP*AP*CP*CP*T)-3'), ... | | Authors: | Liang, S. | | Deposit date: | 2024-07-09 | | Release date: | 2025-07-09 | | Last modified: | 2025-07-30 | | Method: | ELECTRON MICROSCOPY (3.46 Å) | | Cite: | Lactylation of XLF promotes non-homologous end-joining repair and chemoresistance in cancer.
Mol.Cell, 85, 2025
|
|
5KNM
 
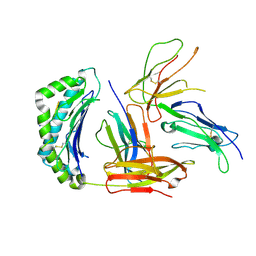 | |
7OWE
 
 | | Odinarchaeota Adenylate kinase (OdinAK) in complex with inhibitor Ap5a | | Descriptor: | Adenylate kinase, BIS(ADENOSINE)-5'-PENTAPHOSPHATE | | Authors: | Grundstrom, C, Aberg-Zingmark, E, Verma, A, Wolf-Watz, M, Sauer, U.H, Sauer-Eriksson, A.E. | | Deposit date: | 2021-06-18 | | Release date: | 2022-09-28 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | Insights into the evolution of enzymatic specificity and catalysis: From Asgard archaea to human adenylate kinases.
Sci Adv, 8, 2022
|
|
5KYC
 
 | |
5KYF
 
 | |
5KYE
 
 | |
5KYB
 
 | |
5KYD
 
 | |
7R8N
 
 | | Structure of the SARS-CoV-2 S 6P trimer in complex with neutralizing antibody C051 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, C051 Fab Heavy Chain, ... | | Authors: | Barnes, C.O, Bjorkman, P.J. | | Deposit date: | 2021-06-26 | | Release date: | 2021-08-04 | | Last modified: | 2024-11-20 | | Method: | ELECTRON MICROSCOPY (3.55 Å) | | Cite: | Affinity maturation of SARS-CoV-2 neutralizing antibodies confers potency, breadth, and resilience to viral escape mutations.
Immunity, 54, 2021
|
|
7R8O
 
 | | Structure of the SARS-CoV-2 S 6P trimer in complex with neutralizing antibody C548 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | DeLaitsch, A.T, Barnes, C.O, Bjorkman, P.J. | | Deposit date: | 2021-06-26 | | Release date: | 2021-08-04 | | Last modified: | 2024-10-16 | | Method: | ELECTRON MICROSCOPY (3.5 Å) | | Cite: | Affinity maturation of SARS-CoV-2 neutralizing antibodies confers potency, breadth, and resilience to viral escape mutations.
Immunity, 54, 2021
|
|
7R8L
 
 | | Structure of the SARS-CoV-2 RBD in complex with neutralizing antibody C099 and CR3022 | | Descriptor: | C099 Fab Heavy Chain, C099 Fab Light Chain, CR3022 Fab heavy chain, ... | | Authors: | Barnes, C.O, Bjorkman, P.J. | | Deposit date: | 2021-06-26 | | Release date: | 2021-08-04 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Affinity maturation of SARS-CoV-2 neutralizing antibodies confers potency, breadth, and resilience to viral escape mutations.
Immunity, 54, 2021
|
|
7R8M
 
 | | Structure of the SARS-CoV-2 S 6P trimer in complex with neutralizing antibody C032 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | DeLaitsch, A.T, Barnes, C.O, Bjorkman, P.J. | | Deposit date: | 2021-06-26 | | Release date: | 2021-08-04 | | Last modified: | 2024-11-13 | | Method: | ELECTRON MICROSCOPY (3.4 Å) | | Cite: | Affinity maturation of SARS-CoV-2 neutralizing antibodies confers potency, breadth, and resilience to viral escape mutations.
Immunity, 54, 2021
|
|
9GFT
 
 | | Structure of the HrpA-bound E. coli disome, Class I | | Descriptor: | 16S ribosomal RNA, 23S RNA, 50S ribosomal protein L11, ... | | Authors: | Esser, H.F, Berninghausen, O, Becker, T, Beckmann, R. | | Deposit date: | 2024-08-12 | | Release date: | 2025-02-12 | | Last modified: | 2025-03-19 | | Method: | ELECTRON MICROSCOPY (3.1 Å) | | Cite: | The RNA helicase HrpA rescues collided ribosomes in E. coli.
Mol.Cell, 85, 2025
|
|
9GGR
 
 | | Structure of the HrpA-bound E. coli disome, Class II | | Descriptor: | 16S ribosomal RNA, 23S ribosomal RNA, 50S ribosomal protein L11, ... | | Authors: | Esser, H.F, Berninghausen, O, Becker, T, Beckmann, R. | | Deposit date: | 2024-08-14 | | Release date: | 2025-02-12 | | Last modified: | 2025-03-19 | | Method: | ELECTRON MICROSCOPY (3.2 Å) | | Cite: | The RNA helicase HrpA rescues collided ribosomes in E. coli.
Mol.Cell, 85, 2025
|
|
9FZB
 
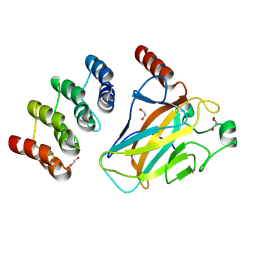 | | Human p53 DNA-binding domain bound to DARPin C10-H82R | | Descriptor: | 1,2-ETHANEDIOL, Cellular tumor antigen p53, DARPin C10-H82R, ... | | Authors: | Yuksel, B, Balourdas, D.I, Muenick, P, Knapp, S, Doetsch, V, Joerger, A.C, Structural Genomics Consortium (SGC) | | Deposit date: | 2024-07-05 | | Release date: | 2025-04-02 | | Last modified: | 2025-05-28 | | Method: | X-RAY DIFFRACTION (1.44 Å) | | Cite: | DARPin-induced reactivation of p53 in HPV-positive cells.
Nat.Struct.Mol.Biol., 32, 2025
|
|
9J89
 
 | | zbp1 nucleic acid complex | | Descriptor: | DNA (5'-D(P*CP*AP*CP*GP*CP*A)-3'), MAGNESIUM ION, RNA (5'-R(P*UP*GP*CP*GP*UP*G)-3'), ... | | Authors: | Gao, A.M, Zhou, C. | | Deposit date: | 2024-08-20 | | Release date: | 2025-06-25 | | Method: | X-RAY DIFFRACTION (1.58 Å) | | Cite: | ZBP1 senses spliceosome stress through Z-RNA:DNA hybrid recognition.
Mol.Cell, 85, 2025
|
|
9J8G
 
 | | mouse zbp1 hybrid complex | | Descriptor: | DNA (5'-D(P*CP*AP*CP*GP*CP*A)-3'), RNA (5'-R(P*UP*GP*CP*GP*UP*G)-3'), Z-DNA-binding protein 1 | | Authors: | Gao, A.M, Zhoiu, C. | | Deposit date: | 2024-08-21 | | Release date: | 2025-06-25 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | ZBP1 senses spliceosome stress through Z-RNA:DNA hybrid recognition.
Mol.Cell, 85, 2025
|
|
5IUE
 
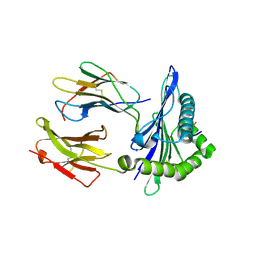 | |
