2D1X
 
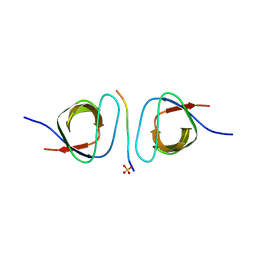 | | The crystal structure of the cortactin-SH3 domain and AMAP1-peptide complex | | Descriptor: | SULFATE ION, cortactin isoform a, proline rich region from development and differentiation enhancing factor 1 | | Authors: | Hashimoto, S, Hirose, M, Hashimoto, A, Morishige, M, Yamada, A, Hosaka, H, Akagi, K, Ogawa, E, Oneyama, C, Agatsuma, T, Okada, M, Kobayashi, H, Wada, H, Nakano, H, Ikegami, T, Nakagawa, A, Sabe, H. | | Deposit date: | 2005-09-01 | | Release date: | 2006-04-25 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Targeting AMAP1 and cortactin binding bearing an atypical src homology 3/proline interface for prevention of breast cancer invasion and metastasis.
Proc.Natl.Acad.Sci.Usa, 103, 2006
|
|
2DA0
 
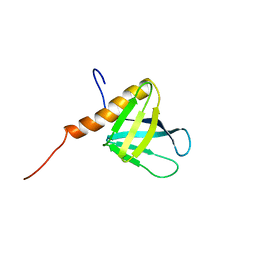 | | Solution structure of the PH domain of PIP2-dependent ARF1 GTPase-activating protein from human | | Descriptor: | 130-kDa phosphatidylinositol 4,5-biphosphate-dependent ARF1 GTPase-activating protein | | Authors: | Li, H, Tomizawa, T, Koshiba, S, Inoue, M, Kigawa, T, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2005-12-13 | | Release date: | 2006-06-13 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the PH domain of PIP2-dependent ARF1 GTPase-activating protein from human
To be Published
|
|
2RQT
 
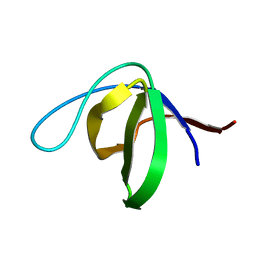 | | Solution structure of the human DDEF1 SH3 domain | | Descriptor: | Arf-GAP with SH3 domain, ANK repeat and PH domain-containing protein 1 | | Authors: | Kaieda, S, Matsui, C, Mimori-Kiyosue, Y, Ikegami, T. | | Deposit date: | 2009-12-14 | | Release date: | 2010-07-07 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Structural basis of the recognition of the SAMP motif of adenomatous polyposis coli by the Src-homology 3 domain.
Biochemistry, 49, 2010
|
|
2RQU
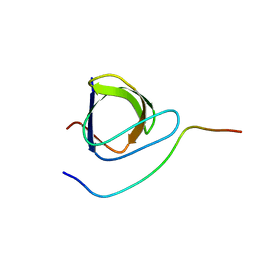 
 | | Solution structure of the complex between the DDEF1 SH3 domain and the APC SAMP1 motif | | Descriptor: | 19-mer from Adenomatous polyposis coli protein, DDEF1_SH3 | | Authors: | Kaieda, S, Matsui, C, Mimori-Kiyosue, Y, Ikegami, T. | | Deposit date: | 2009-12-14 | | Release date: | 2010-07-07 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Structural basis of the recognition of the SAMP motif of adenomatous polyposis coli by the Src-homology 3 domain.
Biochemistry, 49, 2010
|
|
2ED1
 
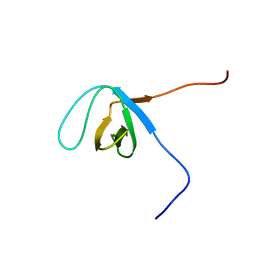 | | Solution structure of the SH3 domain of 130 kDa phosphatidylinositol 4,5-biphosphate-dependent ARF1 GTPase-activating protein | | Descriptor: | 130 kDa phosphatidylinositol 4,5-biphosphate-dependent ARF1 GTPase-activating protein | | Authors: | Abe, H, Tochio, N, Miyamoto, K, Saito, K, Kigawa, T, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2007-02-14 | | Release date: | 2008-02-26 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the SH3 domain of 130 kDa phosphatidylinositol 4,5-biphosphate-dependent ARF1 GTPase-activating protein
To be Published
|
|
