1JZQ
 
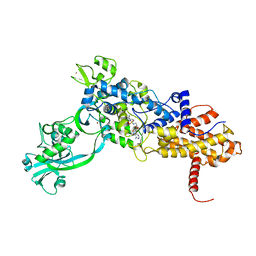 | | Isoleucyl-tRNA synthetase Complexed with Isoleucyl-adenylate analogue | | Descriptor: | Isoleucyl-tRNA synthetase, N-[ISOLEUCINYL]-N'-[ADENOSYL]-DIAMINOSUFONE, ZINC ION | | Authors: | Nakama, T, Nureki, O, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2001-09-17 | | Release date: | 2001-12-21 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structural basis for the recognition of isoleucyl-adenylate and an antibiotic, mupirocin, by isoleucyl-tRNA synthetase.
J.Biol.Chem., 276, 2001
|
|
1NQM
 
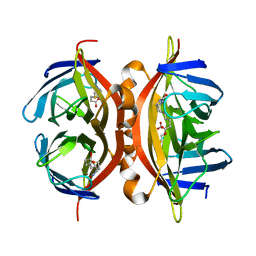 | | Structure of Savm-W120K, streptavidin mutant | | Descriptor: | BIOTIN, Streptavidin | | Authors: | Pazy, Y, Eisenberg-Domovich, Y, Laitinen, O.H, Kulomaa, M.S, Bayer, E.A, Wilchek, M, Livnah, O. | | Deposit date: | 2003-01-22 | | Release date: | 2003-07-15 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Dimer-Tetramer Transition between Solution and Crystalline States of Streptavidin and Avidin Mutants.
J.Bacteriol., 185, 2003
|
|
2B42
 
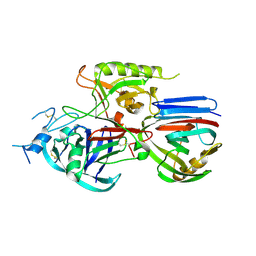 | | Crystal structure of the Triticum xylanse inhibitor-I in complex with bacillus subtilis xylanase | | Descriptor: | Endo-1,4-beta-xylanase A, xylanase inhibitor-I | | Authors: | Sansen, S, Dewilde, M, De Ranter, C.J, Gebruers, K, Brijs, K, Courtin, C.M, Delcour, J.A, Rabijns, A. | | Deposit date: | 2005-09-22 | | Release date: | 2006-09-19 | | Last modified: | 2019-07-24 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Identification of structural determinants for inhibition strength and specificity of wheat xylanase inhibitors TAXI-IA and TAXI-IIA.
Febs J., 276, 2009
|
|
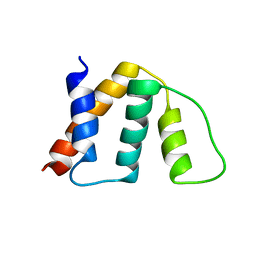
1NTI
 
 | |
1JZ6
 
 | | E. COLI (lacZ) BETA-GALACTOSIDASE IN COMPLEX WITH GALACTO-TETRAZOLE | | Descriptor: | (5R, 6S, 7S, ... | | Authors: | Juers, D.H, Heightman, T.D, Vasella, A, Matthews, B.W. | | Deposit date: | 2001-09-13 | | Release date: | 2001-12-07 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | A Structural View of the Action of Escherichia Coli (Lacz) Beta-Galactosidase
Biochemistry, 40, 2001
|
|
1NWK
 
 | | CRYSTAL STRUCTURE OF MONOMERIC ACTIN IN THE ATP STATE | | Descriptor: | Actin, alpha skeletal muscle, CALCIUM ION, ... | | Authors: | Graceffa, P, Dominguez, R. | | Deposit date: | 2003-02-06 | | Release date: | 2003-10-14 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Crystal structure of monomeric actin in the ATP state. Structural basis of nucleotide-dependent actin dynamics.
J.Biol.Chem., 278, 2003
|
|
2ASM
 
 | | Structure of Rabbit Actin In Complex With Reidispongiolide A | | Descriptor: | 1,2-ETHANEDIOL, ADENOSINE-5'-TRIPHOSPHATE, Actin, ... | | Authors: | Allingham, J.S, Zampella, A, D'Auria, M.V, Rayment, I. | | Deposit date: | 2005-08-23 | | Release date: | 2005-10-11 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structures of microfilament destabilizing toxins bound to actin provide insight into toxin design and activity
Proc.Natl.Acad.Sci.Usa, 102, 2005
|
|
1O1B
 
 | | MOLECULAR MODELS OF AVERAGED RIGOR CROSSBRIDGES FROM TOMOGRAMS OF INSECT FLIGHT MUSCLE | | Descriptor: | SKELETAL MUSCLE ACTIN, SKELETAL MUSCLE MYOSIN II, SKELETAL MUSCLE MYOSIN II ESSENTIAL LIGHT CHAIN, ... | | Authors: | Chen, L.F, Winkler, H, Reedy, M.K, Reedy, M.C, Taylor, K.A. | | Deposit date: | 2002-11-15 | | Release date: | 2002-12-04 | | Last modified: | 2023-12-27 | | Method: | ELECTRON MICROSCOPY (70 Å) | | Cite: | Molecular Modeling of Averaged Rigor Crossbridges from Tomograms of Insect Flight Muscle
J.Struct.Biol., 138, 2002
|
|
1JPC
 
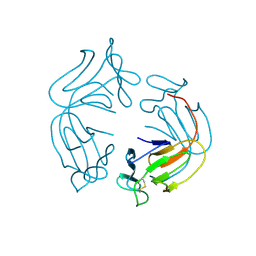 | | MANNOSE-SPECIFIC AGGLUTININ (LECTIN) FROM SNOWDROP (GALANTHUS NIVALIS) BULBS IN COMPLEX WITH MANNOSE-ALPHA1,6-(MANNOSE-ALPHA1,3)-MANNOSE-ALPHA1,6-(MANNOSE-ALPHA1,3)-MANNOSE | | Descriptor: | AGGLUTININ, alpha-D-mannopyranose-(1-3)-[alpha-D-mannopyranose-(1-6)]alpha-D-mannopyranose, alpha-D-mannopyranose-(1-6)-alpha-D-mannopyranose | | Authors: | Wright, C.S, Hester, G. | | Deposit date: | 1996-07-30 | | Release date: | 1997-01-27 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The 2.0 A structure of a cross-linked complex between snowdrop lectin and a branched mannopentaose: evidence for two unique binding modes.
Structure, 4, 1996
|
|
2AJF
 
 | | Structure of SARS coronavirus spike receptor-binding domain complexed with its receptor | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Angiotensin-converting enzyme-Related Carboxypeptidase (Ace2), CHLORIDE ION, ... | | Authors: | Li, F, Li, W, Farzan, M, Harrison, S.C. | | Deposit date: | 2005-08-01 | | Release date: | 2005-09-20 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Structure of SARS coronavirus spike receptor-binding domain complexed with receptor.
Science, 309, 2005
|
|
2AKI
 
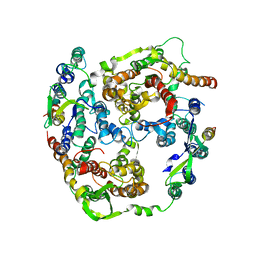 | | Normal mode-based flexible fitted coordinates of a translocating SecYEG protein-conducting channel into the cryo-EM map of a SecYEG-nascent chain-70S ribosome complex from E. coli | | Descriptor: | Preprotein translocase secE subunit, Preprotein translocase secY subunit, Protein-export membrane protein secG | | Authors: | Mitra, K, Schaffitzel, C, Shaikh, T, Tama, F, Jenni, S, Brooks III, C.L, Ban, N, Frank, J. | | Deposit date: | 2005-08-03 | | Release date: | 2005-11-15 | | Last modified: | 2024-02-14 | | Method: | ELECTRON MICROSCOPY (14.9 Å) | | Cite: | Structure of the E. coli protein-conducting channel bound to a translating ribosome.
Nature, 438, 2005
|
|
1O1T
 
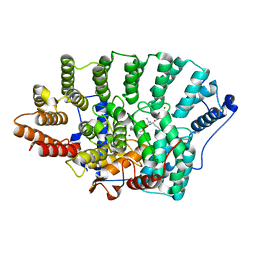 | | Structure of FPT bound to the CVIM-FPP product | | Descriptor: | MAGNESIUM ION, N-ACETYL-S-[(2E,6E)-3,7,11-TRIMETHYLDODECA-2,6,10-TRIENYL]-L-CYSTEINYL-D-VALYL-L-ISOLEUCYL-L-METHIONINE, Protein farnesyltransferase alpha subunit, ... | | Authors: | Turek-Etienne, T.C, Strickland, C.L, Distefano, M.D. | | Deposit date: | 2003-01-14 | | Release date: | 2003-05-06 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Biochemical and Structural Studies with Prenyl Diphosphate Analogues Provide Insights into
Isoprenoid Recognition by Protein Farnesyl Transferase
Biochemistry, 42, 2003
|
|
2B12
 
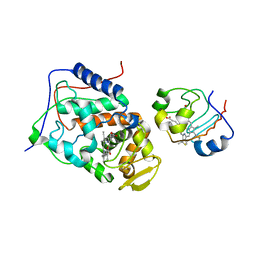 | |
1JZ5
 
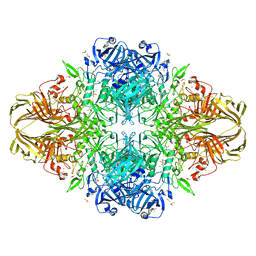 | |
1JTS
 
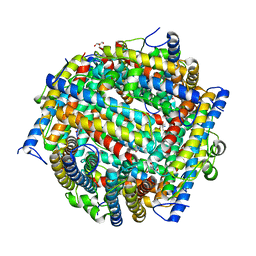 | |
2B1J
 
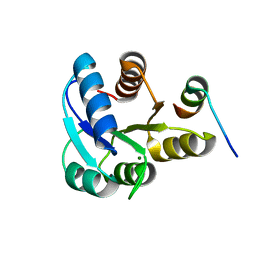 | |
2B2H
 
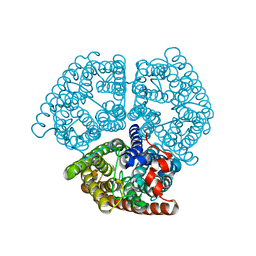 | |
1K2U
 
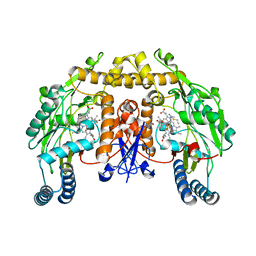 | | Structure of rat brain nNOS heme domain complexed with S-ethyl-N-[4-(trifluoromethyl)phenyl] isothiourea | | Descriptor: | 5,6,7,8-TETRAHYDROBIOPTERIN, ACETATE ION, PROTOPORPHYRIN IX CONTAINING FE, ... | | Authors: | Li, H, Martasek, P, Masters, B.S.S, Poulos, T.L, Raman, C.S. | | Deposit date: | 2001-09-28 | | Release date: | 2003-03-04 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structure of rat brain nNOS heme domain
To be Published
|
|
1JYN
 
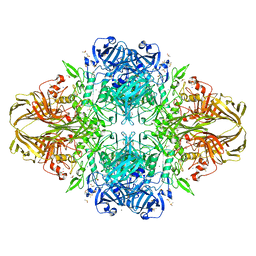 | |
2ASP
 
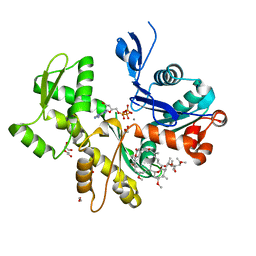 | | Structure of Rabbit Actin In Complex With Reidispongiolide C | | Descriptor: | 1,2-ETHANEDIOL, ADENOSINE-5'-TRIPHOSPHATE, Actin, ... | | Authors: | Allingham, J.S, Zampella, A, D'Auria, M.V, Rayment, I. | | Deposit date: | 2005-08-23 | | Release date: | 2005-10-11 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.64 Å) | | Cite: | Structures of microfilament destabilizing toxins bound to actin provide insight into toxin design and activity
Proc.Natl.Acad.Sci.Usa, 102, 2005
|
|
2AN2
 
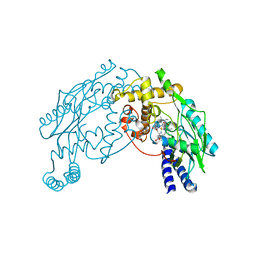 | | P332G, A333S Double mutant of the Bacillus subtilis Nitric Oxide Synthase | | Descriptor: | 5,6,7,8-TETRAHYDROBIOPTERIN, ARGININE, P332G A333S double mutant of Nitric Oxide Synthase from Bacillus subtilis, ... | | Authors: | Pant, K, Crane, B.R. | | Deposit date: | 2005-08-11 | | Release date: | 2006-08-29 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structure of a loose dimer: an intermediate in nitric oxide synthase assembly
J.Mol.Biol., 352, 2005
|
|
1OC5
 | | D405N mutant of the CELLOBIOHYDROLASE CEL6A FROM HUMICOLA INSOLENS in complex with methyl-cellobiosyl-4-deoxy-4-thio-beta-D-cellobioside | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, CELLOBIOHYDROLASE II, GLYCEROL, ... | | Authors: | Varrot, A, Frandsen, T.P, Von Ossowski, I, Boyer, V, Driguez, H, Schulein, M, Davies, G.J. | | Deposit date: | 2003-02-06 | | Release date: | 2003-07-10 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structural Basis for Ligand Binding and Processivity in Cellobiohydrolase Cel6A from Humicola Insolens
Structure, 11, 2003
|
|
1OCN
 
 | | Mutant D416A of the CELLOBIOHYDROLASE CEL6A FROM HUMICOLA INSOLENS in complex with a cellobio-derived isofagomine at 1.3 angstrom resolution | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 5-HYDROXYMETHYL-3,4-DIHYDROXYPIPERIDINE, CALCIUM ION, ... | | Authors: | Varrot, A, Macdonald, J, Stick, R.V, Pell, G, Gilbert, H.J, Davies, G.J. | | Deposit date: | 2003-02-09 | | Release date: | 2003-05-22 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.31 Å) | | Cite: | Distortion of a Cellobio-Derived Isofagomine Highlights the Potential Conformational Itinerary of Inverting Beta-Glucosidases
Chem.Commun.(Camb.), 21, 2003
|
|
1OCB
 
 | | Structure of the wild-type cellobiohydrolase Cel6A from Humicolas insolens in complex with a fluorescent substrate | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 4-amino-4-deoxy-alpha-D-glucopyranose-(1-4)-beta-D-glucopyranose-(1-4)-beta-D-glucopyranose-(1-4)-methyl 4-thio-beta-D-glucopyranoside, CELLOBIOHYDROLASE II, ... | | Authors: | Varrot, A, Frandsen, T.P, Von Ossowski, I, Boyer, V, Driguez, H, Schulein, M, Davies, G.J. | | Deposit date: | 2003-02-07 | | Release date: | 2003-07-10 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Structural Basis for Ligand Binding and Processivity in Cellobiohydrolase Cel6A from Humicola Insolens
Structure, 11, 2003
|
|
1JI1
 
 | | Crystal Structure Analysis of Thermoactinomyces vulgaris R-47 alpha-Amylase 1 | | Descriptor: | ALPHA-AMYLASE I, CALCIUM ION | | Authors: | Kamitori, S, Abe, A, Ohtaki, A, Kaji, A, Tonozuka, T, Sakano, Y. | | Deposit date: | 2001-06-28 | | Release date: | 2002-06-05 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Crystal structures and structural comparison of Thermoactinomyces vulgaris R-47 alpha-amylase 1 (TVAI) at 1.6 A resolution and alpha-amylase 2 (TVAII) at 2.3 A resolution.
J.Mol.Biol., 318, 2002
|
|
