9DE5
 
 | |
8EKZ
 
 | | Human PU.1 ETS-Domain (165-270) Bound to d(AATAAGGAGAAGTAGG) | | Descriptor: | DNA (5'-D(*AP*AP*TP*AP*AP*GP*GP*AP*GP*AP*AP*GP*TP*AP*GP*G)-3'), DNA (5'-D(*TP*CP*CP*TP*AP*CP*TP*TP*CP*TP*CP*CP*TP*TP*AP*T)-3'), SODIUM ION, ... | | Authors: | Terrell, J.R, Poon, G.M.K. | | Deposit date: | 2022-09-22 | | Release date: | 2023-07-05 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.42 Å) | | Cite: | DNA selection by the master transcription factor PU.1.
Cell Rep, 42, 2023
|
|
8EMD
 
 | |
8E5Y
 
 | |
8EE9
 
 | | Human PU.1 ETS-Domain (165-270) Bound to d(AATAAAAGGAATGGGG) | | Descriptor: | DNA (5'-D(*AP*AP*TP*AP*AP*AP*AP*GP*GP*AP*AP*TP*GP*GP*GP*G)-3'), DNA (5'-D(*TP*CP*CP*CP*CP*AP*TP*TP*CP*CP*TP*TP*TP*TP*AP*T)-3'), Transcription factor PU.1 | | Authors: | Terrell, J.R, Poon, G.M.K. | | Deposit date: | 2022-09-06 | | Release date: | 2023-07-05 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.22 Å) | | Cite: | DNA selection by the master transcription factor PU.1.
Cell Rep, 42, 2023
|
|
8EK8
 
 | | Human PU.1 ETS-Domain (165-270) Bound to d(AATAAAAGGAGAAGGG) | | Descriptor: | DNA (5'-D(*AP*TP*AP*AP*AP*AP*GP*GP*AP*GP*AP*AP*GP*GP*G)-3'), DNA (5'-D(P*CP*CP*CP*TP*TP*CP*TP*CP*CP*TP*TP*TP*TP*AP*T)-3'), Transcription factor PU.1 | | Authors: | Terrell, J.R, Poon, G.M.K. | | Deposit date: | 2022-09-20 | | Release date: | 2023-07-05 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.63 Å) | | Cite: | DNA selection by the master transcription factor PU.1.
Cell Rep, 42, 2023
|
|
8EBH
 
 | | Human PU.1 ETS-Domain (165-270) Bound to d(AATAAGCGGAATGGGG) | | Descriptor: | DNA (5'-D(*AP*AP*TP*AP*AP*GP*CP*GP*GP*AP*AP*TP*GP*GP*GP*G)-3'), DNA (5'-D(*TP*CP*CP*CP*CP*AP*TP*TP*CP*CP*GP*CP*TP*TP*AP*T)-3'), Transcription factor PU.1 | | Authors: | Terrell, J.R, Poon, G.M.K. | | Deposit date: | 2022-08-31 | | Release date: | 2023-07-05 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.33 Å) | | Cite: | DNA selection by the master transcription factor PU.1.
Cell Rep, 42, 2023
|
|
8EJ6
 
 | | Human PU.1 ETS-Domain (165-270) Bound to d(AATAAGAGGAATGGGG) | | Descriptor: | DNA (5'-D(*AP*AP*TP*AP*AP*GP*AP*GP*GP*AP*AP*TP*GP*GP*GP*G)-3'), DNA (5'-D(*TP*CP*CP*CP*CP*AP*TP*TP*CP*CP*TP*CP*TP*TP*AP*T)-3'), Transcription factor PU.1 | | Authors: | Terrell, J.R, Poon, G.M.K. | | Deposit date: | 2022-09-16 | | Release date: | 2023-07-05 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.39 Å) | | Cite: | DNA selection by the master transcription factor PU.1.
Cell Rep, 42, 2023
|
|
5HY1
 
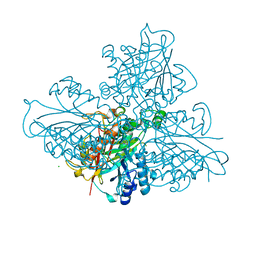 | | high resolution structure of barbiturase | | Descriptor: | 1,3,5-triazine-2,4,6-triol, Barbiturase, CHLORIDE ION, ... | | Authors: | Peat, T.S, Scott, C, Balotra, S, Wilding, M, Newman, J. | | Deposit date: | 2016-02-01 | | Release date: | 2017-02-01 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.01 Å) | | Cite: | High-Resolution X-Ray Structures of Two Functionally Distinct Members of the Cyclic Amide Hydrolase Family of Toblerone Fold Enzymes.
Appl. Environ. Microbiol., 83, 2017
|
|
5ZJN
 
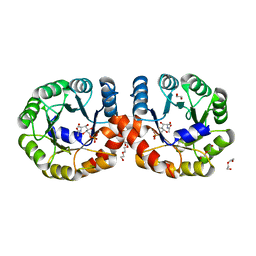 | |
8IY6
 
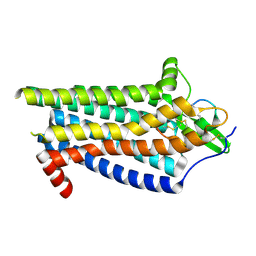 | | ETB-Gi complex bound to Endotheline-1, focused on receptor | | Descriptor: | Endothelin type B receptor, Endothelin-1 | | Authors: | Sano, F.K, Akasaka, H, Shihoya, W, Nureki, O. | | Deposit date: | 2023-04-04 | | Release date: | 2023-08-16 | | Last modified: | 2025-07-02 | | Method: | ELECTRON MICROSCOPY (3.13 Å) | | Cite: | Cryo-EM structure of the endothelin-1-ET B -G i complex.
Elife, 12, 2023
|
|
5N7D
 
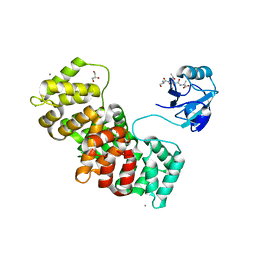 | | MAGI-1 complexed with a RSK1 peptide | | Descriptor: | CALCIUM ION, GLYCEROL, Membrane-associated guanylate kinase, ... | | Authors: | Gogl, G, Nyitray, L. | | Deposit date: | 2017-02-20 | | Release date: | 2017-11-08 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Dynamic control of RSK complexes by phosphoswitch-based regulation.
FEBS J., 285, 2018
|
|
7VKW
 
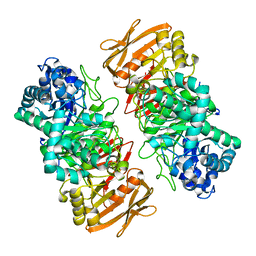 | | The apo structure of beta-1,2-glucosyltransferase from Ignavibacterium album | | Descriptor: | beta-1,2-glucosyltransferase | | Authors: | Kobayashi, K, Shimizu, H, Tanaka, N, Kuramochi, K, Nakai, H, Nakajima, M, Taguchi, H. | | Deposit date: | 2021-10-01 | | Release date: | 2022-03-09 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Characterization and structural analyses of a novel glycosyltransferase acting on the beta-1,2-glucosidic linkages.
J.Biol.Chem., 298, 2022
|
|
8VKM
 
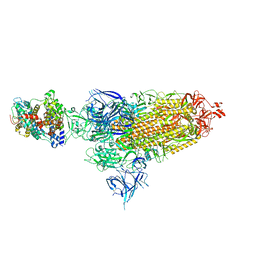 | | Cryo-EM structure of SARS-CoV-2 XBB.1.5 spike protein in complex with mouse ACE2 (conformation 1) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Angiotensin-converting enzyme 2, ... | | Authors: | Zhu, X, Mannar, D, Saville, J, Poloni, C, Bezeruk, A, Tidey, K, Ahmed, S, Tuttle, K, Vahdatihassani, F, Cholak, S, Cook, L, Steiner, T.S, Subramaniam, S. | | Deposit date: | 2024-01-09 | | Release date: | 2024-02-14 | | Last modified: | 2025-02-26 | | Method: | ELECTRON MICROSCOPY (2.83 Å) | | Cite: | Altered receptor binding, antibody evasion and retention of T cell recognition by the SARS-CoV-2 XBB.1.5 spike protein.
Nat Commun, 15, 2024
|
|
8VKL
 
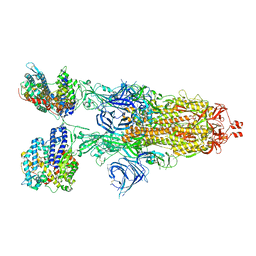 | | Cryo-EM structure of SARS-CoV-2 XBB.1.5 spike protein in complex with mouse ACE2 (conformation 2) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Angiotensin-converting enzyme 2, ... | | Authors: | Zhu, X, Mannar, D, Saville, J, Poloni, C, Bezeruk, A, Tidey, K, Ahmed, S, Tuttle, K, Vahdatihassani, F, Cholak, S, Cook, L, Steiner, T.S, Subramaniam, S. | | Deposit date: | 2024-01-09 | | Release date: | 2024-02-14 | | Last modified: | 2025-02-26 | | Method: | ELECTRON MICROSCOPY (2.91 Å) | | Cite: | Altered receptor binding, antibody evasion and retention of T cell recognition by the SARS-CoV-2 XBB.1.5 spike protein.
Nat Commun, 15, 2024
|
|
9R3I
 
 | | Structure of liver pyruvate kinase in complex with fluorescent probe 4c | | Descriptor: | 1,6-di-O-phosphono-beta-D-fructofuranose, 4-[4-[[7-(dimethylamino)-2,1,3-benzoxadiazol-4-yl]sulfonyl]piperazin-1-yl]sulfonyl-5-pyridin-3-yl-benzene-1,2-diol, Isoform L-type of Pyruvate kinase PKLR | | Authors: | Bogucka, A, Nilsson, O, Grotli, M, Hyvonen, M. | | Deposit date: | 2025-05-05 | | Release date: | 2025-08-13 | | Method: | X-RAY DIFFRACTION (2.579 Å) | | Cite: | Fluorescent binding assay for allosteric ligands of liver pyruvate kinase.
Eur.J.Med.Chem., 298, 2025
|
|
7W2B
 
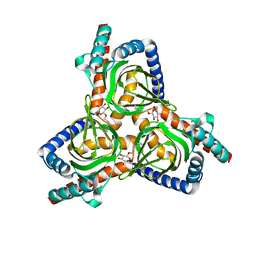 | |
8VKN
 
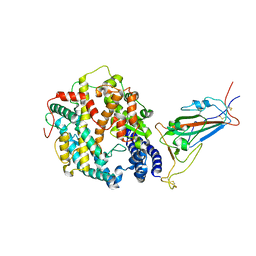 | | Cryo-EM structure of SARS-CoV-2 XBB.1.5 spike protein in complex with mouse ACE2 (focused refinement of RBD and mouse ACE2) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Angiotensin-converting enzyme 2, ... | | Authors: | Zhu, X, Mannar, D, Saville, J, Poloni, C, Bezeruk, A, Tidey, K, Ahmed, S, Tuttle, K, Vahdatihassani, F, Cholak, S, Cook, L, Steiner, T.S, Subramaniam, S. | | Deposit date: | 2024-01-09 | | Release date: | 2024-02-14 | | Last modified: | 2025-02-26 | | Method: | ELECTRON MICROSCOPY (2.93 Å) | | Cite: | Altered receptor binding, antibody evasion and retention of T cell recognition by the SARS-CoV-2 XBB.1.5 spike protein.
Nat Commun, 15, 2024
|
|
9BZP
 
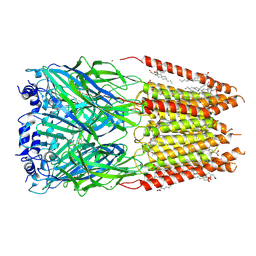 | | Homomeric alpha3 glycine receptor in the presence of 0.1 millimolar glycine and 1 micromolar zinc in a desensitized state | | Descriptor: | 1,2-DIMYRISTOYL-SN-GLYCERO-3-PHOSPHOCHOLINE, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, GLYCINE, ... | | Authors: | Kindig, K, Gibbs, E, Chakrapani, S. | | Deposit date: | 2024-05-24 | | Release date: | 2024-11-13 | | Last modified: | 2025-05-28 | | Method: | ELECTRON MICROSCOPY (2.88 Å) | | Cite: | Mechanisms underlying modulation of human GlyR alpha 3 by Zn 2+ and pH.
Sci Adv, 10, 2024
|
|
9QAN
 
 | | Human angiotensin-1 converting enzyme C-domain in complex with ramiprilat | | Descriptor: | 1,2-ETHANEDIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose, Angiotensin-converting enzyme, ... | | Authors: | Gregory, K.S, Acharya, K.R. | | Deposit date: | 2025-02-28 | | Release date: | 2025-09-03 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Molecular basis of domain-specific angiotensin I-converting enzyme inhibition by the antihypertensive drugs enalaprilat, ramiprilat, trandolaprilat, quinaprilat and perindoprilat.
Febs J., 2025
|
|
9BWJ
 
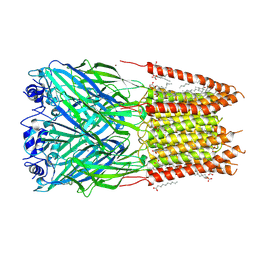 | | Homomeric alpha3 glycine receptor in the presence of 0.1 mM glycine and 0.1 mM zinc in an apo state | | Descriptor: | 1,2-DIMYRISTOYL-SN-GLYCERO-3-PHOSPHOCHOLINE, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Glycine receptor subunit alpha-3, ... | | Authors: | Kindig, K, Gibbs, E, Chakrapani, S. | | Deposit date: | 2024-05-21 | | Release date: | 2024-11-13 | | Last modified: | 2025-05-28 | | Method: | ELECTRON MICROSCOPY (2.21 Å) | | Cite: | Mechanisms underlying modulation of human GlyR alpha 3 by Zn 2+ and pH.
Sci Adv, 10, 2024
|
|
8ORP
 
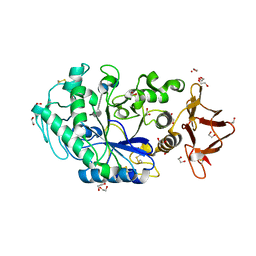 | | Crystal structure of Drosophila melanogaster alpha-amylase in complex with the inhibitor acarbose | | Descriptor: | 1,2-ETHANEDIOL, 4,6-dideoxy-4-{[(1S,2S,3S,4R,5R)-2,3,4-trihydroxy-5-(hydroxymethyl)cyclohexyl]amino}-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose, 4,6-dideoxy-4-{[(1S,2S,3S,4R,5R)-2,3,4-trihydroxy-5-(hydroxymethyl)cyclohexyl]amino}-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose-(1-4)-beta-D-glucopyranose, ... | | Authors: | Aghajari, N, Haser, R. | | Deposit date: | 2023-04-16 | | Release date: | 2023-08-16 | | Last modified: | 2025-10-01 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural and Functional Characterization of Drosophila melanogaster alpha-Amylase.
Molecules, 28, 2023
|
|
9C6E
 
 | | High-resolution structure of bovine (3-367)Arrestin-1 in a pre-activated conformation | | Descriptor: | GLYCEROL, GLYCOLIC ACID, S-arrestin, ... | | Authors: | Salom, D, Palczewski, K, Kiser, P.D. | | Deposit date: | 2024-06-07 | | Release date: | 2024-11-06 | | Last modified: | 2025-01-29 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Insights into the Activation and Self-Association of Arrestin-1.
Biochemistry, 64, 2025
|
|
9BWG
 
 | | Homomeric alpha3 glycine receptor in the presence of 0.1 mM glycine and 0.1 mM zinc in a desensitized state | | Descriptor: | 1,2-DIMYRISTOYL-SN-GLYCERO-3-PHOSPHOCHOLINE, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, GLYCINE, ... | | Authors: | Kindig, K, Gibbs, E, Chakrapani, S. | | Deposit date: | 2024-05-21 | | Release date: | 2024-11-13 | | Last modified: | 2025-05-28 | | Method: | ELECTRON MICROSCOPY (2.59 Å) | | Cite: | Mechanisms underlying modulation of human GlyR alpha 3 by Zn 2+ and pH.
Sci Adv, 10, 2024
|
|
8QYX
 
 | | Human 60S ribosomal subunit | | Descriptor: | 1,4-DIAMINOBUTANE, 28S rRNA, 5.8S rRNA, ... | | Authors: | Wiechert, F, Schacherl, M, Sprink, T. | | Deposit date: | 2023-10-26 | | Release date: | 2024-12-11 | | Last modified: | 2025-01-22 | | Method: | ELECTRON MICROSCOPY (1.78 Å) | | Cite: | Visualizing the modification landscape of the human 60S ribosomal subunit at close to atomic resolution.
Nucleic Acids Res., 53, 2025
|
|
