133L
 
 | |
7QZL
 
 | | Amine Dehydrogenase from Cystobacter fuscus (CfusAmDH) W145A mutant with NADP+ and pentylamine | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, AMYLAMINE, Amine Dehydrogenase, ... | | Authors: | Bennett, M, Ducrot, L, Vaxelaire-Vergne, C, Grogan, G. | | Deposit date: | 2022-01-31 | | Release date: | 2022-12-07 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Cover Feature: Expanding the Substrate Scope of Native Amine Dehydrogenases through In Silico Structural Exploration and Targeted Protein Engineering (ChemCatChem 22/2022)
Chemcatchem, 14, 2022
|
|
1VRB
 
 | |
1VRM
 
 | |
7QZH
 
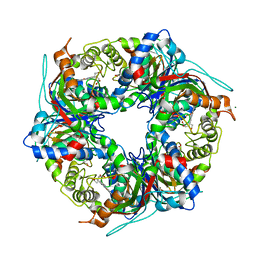 | | SFX structure of dye-type peroxidase DtpB D152A variant in the ferric state | | Descriptor: | Dyp-type peroxidase family, MAGNESIUM ION, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Lucic, M, Worrall, J.A.R, Hough, M.A, Shilova, A, Axford, D.A, Owen, R.L, Tosha, T, Sugimoto, H, Owada, S. | | Deposit date: | 2022-01-31 | | Release date: | 2022-12-07 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.92 Å) | | Cite: | Serial Femtosecond Crystallography Reveals the Role of Water in the One- or Two-Electron Redox Chemistry of Compound I in the Catalytic Cycle of the B-Type Dye-Decolorizing Peroxidase DtpB.
Acs Catalysis, 12, 2022
|
|
134L
 
 | |
7QZE
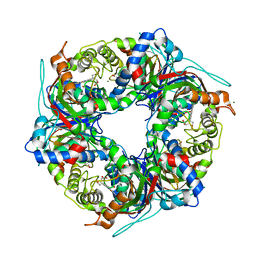 
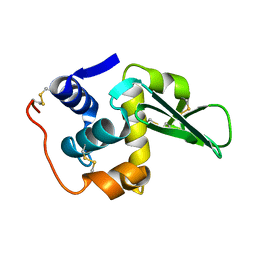
 | | SFX structure of dye-type peroxidase DtpB D152A variant in the ferryl state | | Descriptor: | Dyp-type peroxidase family, MAGNESIUM ION, OXYGEN ATOM, ... | | Authors: | Lucic, M, Worrall, J.A.R, Hough, M.A, Shilova, A, Axford, D.A, Owen, R.L, Tosha, T, Sugimoto, H, Owada, S. | | Deposit date: | 2022-01-31 | | Release date: | 2022-12-07 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Serial Femtosecond Crystallography Reveals the Role of Water in the One- or Two-Electron Redox Chemistry of Compound I in the Catalytic Cycle of the B-Type Dye-Decolorizing Peroxidase DtpB.
Acs Catalysis, 12, 2022
|
|
1VS1
 
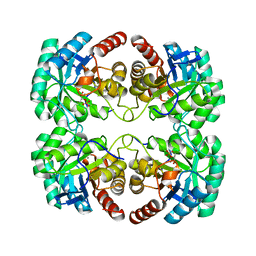 | | Crystal structure of 3-deoxy-D-arabino-heptulosonate-7-phosphate synthase (DAHP synthase) from Aeropyrum pernix in complex with Mn2+ and PEP | | Descriptor: | 3-deoxy-7-phosphoheptulonate synthase, MANGANESE (II) ION, PHOSPHOENOLPYRUVATE | | Authors: | Shumilin, I.A, Zhou, L, Wu, J, Woodard, R.W, Bauerle, R, Kretsinger, R.H. | | Deposit date: | 2006-03-09 | | Release date: | 2006-05-16 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal structure of 3-deoxy-D-arabino-heptulosonate-7-phosphate synthase (DAHP synthase) from Aeropyrum pernix in complex with Mn2+ and PEP
TO BE PUBLISHED
|
|
7QZF
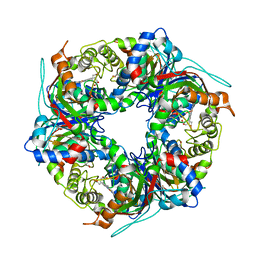 
 | | SFX structure of dye-type peroxidase DtpB D152A/N245A variant in the ferric state | | Descriptor: | Dyp-type peroxidase family, MAGNESIUM ION, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Lucic, M, Worrall, J.A.R, Hough, M.A, Shilova, A, Owen, R.L, Axford, D, Tosha, T, Sugimoto, H, Owada, S. | | Deposit date: | 2022-01-31 | | Release date: | 2022-12-07 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Serial Femtosecond Crystallography Reveals the Role of Water in the One- or Two-Electron Redox Chemistry of Compound I in the Catalytic Cycle of the B-Type Dye-Decolorizing Peroxidase DtpB.
Acs Catalysis, 12, 2022
|
|
1W7S
 
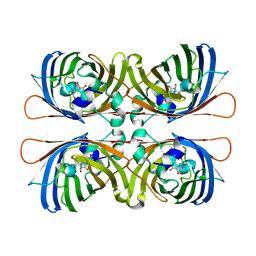 | |
1W8K
 
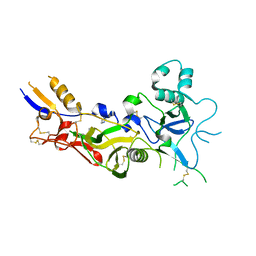 | | Crystal structure of apical membrane antigen 1 from Plasmodium vivax | | Descriptor: | APICAL MEMBRANE ANTIGEN 1, IMIDAZOLE | | Authors: | Pizarro, J.C, Vulliez-Le Normand, B, Chesne-Seck, M.-L, Kocken, C.H.M, Thomas, A.W, Bentley, G.A. | | Deposit date: | 2004-09-23 | | Release date: | 2005-03-01 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal Structure of the Malaria Vaccine Candidate Apical Membrane Antigen 1
Science, 308, 2005
|
|
7QZG
 
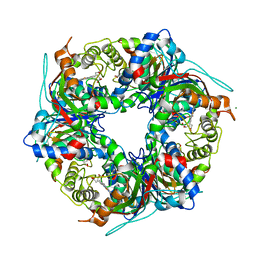 | | SFX structure of dye-type peroxidase DtpB N245A variant in the ferric state | | Descriptor: | Dyp-type peroxidase family, MAGNESIUM ION, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Lucic, M, Worrall, J.A.R, Hough, M.A, Shilova, A, Axford, D.A, Owen, R.L, Tosha, T, Sugimoto, H, Owada, S. | | Deposit date: | 2022-01-31 | | Release date: | 2022-12-07 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Serial Femtosecond Crystallography Reveals the Role of Water in the One- or Two-Electron Redox Chemistry of Compound I in the Catalytic Cycle of the B-Type Dye-Decolorizing Peroxidase DtpB.
Acs Catalysis, 12, 2022
|
|
5GP4
 
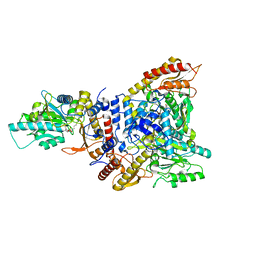 | | Lactobacillus brevis CGMCC 1306 Glutamate decarboxylase | | Descriptor: | Glutamate decarboxylase, PYRIDOXAL-5'-PHOSPHATE | | Authors: | Mei, L, Huang, J. | | Deposit date: | 2016-07-31 | | Release date: | 2017-08-02 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.16 Å) | | Cite: | Lactobacillus brevis CGMCC 1306 glutamate decarboxylase: Crystal structure and functional analysis.
Biochem. Biophys. Res. Commun., 503, 2018
|
|
5GQZ
 
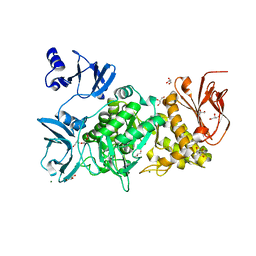 | |
7R59
 
 | | PARP2 catalytic domain in complex with OUL245 | | Descriptor: | GLYCEROL, Poly [ADP-ribose] polymerase 2, [1,2,4]triazolo[3,4-b][1,3]benzothiazol-6-ol | | Authors: | Galera-Prat, A, Maksimainen, M.M, Lehtio, L. | | Deposit date: | 2022-02-10 | | Release date: | 2023-01-25 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | [1,2,4]Triazolo[3,4- b ]benzothiazole Scaffold as Versatile Nicotinamide Mimic Allowing Nanomolar Inhibition of Different PARP Enzymes.
J.Med.Chem., 66, 2023
|
|
7REY
 
 | | MYCOBACTERIUM ABSCESSUS TRNA METHYLTRANSFERASE IN APO FORM | | Descriptor: | SODIUM ION, tRNA (guanine-N(1)-)-methyltransferase | | Authors: | Prucha, G.R, Ismail, M, Suske, A, Das, B, Oz, M, Perez, A, Bolen, R, Jayaraman, S, Stojanoff, V, Halloran, J. | | Deposit date: | 2021-07-13 | | Release date: | 2023-01-18 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.87 Å) | | Cite: | Crystal structure of divalent Mg2+ dependent Mycobacterium abscessus tRNA (m1 G37) Methyltransferase (TrmD)
To Be Published
|
|
7R3Z
 
 | | Tankyrase 2 catalytic domain in complex with OUL40 | | Descriptor: | 6-methyl-[1,2,4]triazolo[3,4-b][1,3]benzothiazole, GLYCEROL, Poly [ADP-ribose] polymerase tankyrase-2, ... | | Authors: | Murthy, S, Venkannagari, H, Maksimainen, M.M, Lehtio, L. | | Deposit date: | 2022-02-08 | | Release date: | 2023-01-25 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | [1,2,4]Triazolo[3,4- b ]benzothiazole Scaffold as Versatile Nicotinamide Mimic Allowing Nanomolar Inhibition of Different PARP Enzymes.
J.Med.Chem., 66, 2023
|
|
7R5D
 
 | | PARP15 catalytic domain in complex with OUL234 | | Descriptor: | 6-propan-2-yl-[1,2,4]triazolo[3,4-b][1,3]benzothiazole, DIMETHYL SULFOXIDE, Protein mono-ADP-ribosyltransferase PARP15 | | Authors: | Maksimainen, M.M, Lehtio, L. | | Deposit date: | 2022-02-10 | | Release date: | 2023-01-25 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | [1,2,4]Triazolo[3,4- b ]benzothiazole Scaffold as Versatile Nicotinamide Mimic Allowing Nanomolar Inhibition of Different PARP Enzymes.
J.Med.Chem., 66, 2023
|
|
1VES
 
 | | Structure of New Antigen Receptor variable domain from sharks | | Descriptor: | New Antigen Receptor variable domain | | Authors: | Streltsov, V.A. | | Deposit date: | 2004-04-05 | | Release date: | 2004-08-24 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.18 Å) | | Cite: | Structural evidence for evolution of shark Ig new antigen receptor variable domain antibodies from a cell-surface receptor
Proc.Natl.Acad.Sci.USA, 101, 2004
|
|
1VKI
 
 | |
6DZH
 
 | | HRAS G13D bound to GDP (H13GDP) | | Descriptor: | 2,3-DIHYDROXY-1,4-DITHIOBUTANE, CALCIUM ION, GLYCEROL, ... | | Authors: | Johnson, C.W, Mattos, C. | | Deposit date: | 2018-07-04 | | Release date: | 2019-07-10 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Isoform-Specific Destabilization of the Active Site Reveals a Molecular Mechanism of Intrinsic Activation of KRas G13D.
Cell Rep, 28, 2019
|
|
7R3L
 
 | | PARP14 catalytic domain in complex with OUL40 | | Descriptor: | 6-methyl-[1,2,4]triazolo[3,4-b][1,3]benzothiazole, CHLORIDE ION, Poly [ADP-ribose] polymerase 14, ... | | Authors: | Maksimainen, M.M, Murthy, S, Lehtio, L. | | Deposit date: | 2022-02-07 | | Release date: | 2023-01-25 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | [1,2,4]Triazolo[3,4- b ]benzothiazole Scaffold as Versatile Nicotinamide Mimic Allowing Nanomolar Inhibition of Different PARP Enzymes.
J.Med.Chem., 66, 2023
|
|
1VL5
 
 | |
7R3O
 
 | | PARP15 catalytic domain in complex with OUL40 | | Descriptor: | 6-methyl-[1,2,4]triazolo[3,4-b][1,3]benzothiazole, DIMETHYL SULFOXIDE, Protein mono-ADP-ribosyltransferase PARP15 | | Authors: | Maksimainen, M.M, Murthy, S, Lehtio, L. | | Deposit date: | 2022-02-07 | | Release date: | 2023-01-25 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | [1,2,4]Triazolo[3,4- b ]benzothiazole Scaffold as Versatile Nicotinamide Mimic Allowing Nanomolar Inhibition of Different PARP Enzymes.
J.Med.Chem., 66, 2023
|
|
1VLU
 
 | |
