7RZ0
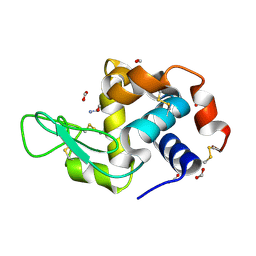 
 | | Hen egg-white lysozyme with ionic liquid ethanolammonium formate 6.7 mol% | | Descriptor: | ETHANOLAMINE, FORMIC ACID, Lysozyme C | | Authors: | Han, Q, Darmanin, C, Drummond, C, Greaves, T. | | Deposit date: | 2021-08-27 | | Release date: | 2023-03-01 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.38 Å) | | Cite: | Probing ion-binding at a protein interface: Modulation of protein properties by ionic liquids.
J Colloid Interface Sci, 650, 2023
|
|
1GVL
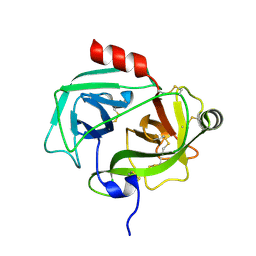 
 | | Human prokallikrein 6 (hK6)/ prozyme/ proprotease M/ proneurosin | | Descriptor: | KALLIKREIN 6 | | Authors: | Gomis-Ruth, F.X, Bayes, A, Sotiropoulou, G, Pampalakis, G, Tsetsenis, T, Villegas, V, Aviles, F.X, Coll, M. | | Deposit date: | 2002-02-14 | | Release date: | 2002-05-16 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | The Structure of Human Prokallikrein 6 Reveals a Novel Activation Mechanism for the Kallikrein Family.
J.Biol.Chem., 277, 2002
|
|
7RZ2
 
 | | Hen egg-white lysozyme with ionic liquid ethanolammonium formate 4 mol% | | Descriptor: | ETHANOLAMINE, FORMIC ACID, Lysozyme C | | Authors: | Han, Q, Darmanin, C, Drummond, C, Greaves, T. | | Deposit date: | 2021-08-27 | | Release date: | 2023-03-01 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.07 Å) | | Cite: | Probing ion-binding at a protein interface: Modulation of protein properties by ionic liquids.
J Colloid Interface Sci, 650, 2023
|
|
7RZ1
 
 | | Hen egg-white lysozyme with ionic liquid ethanolammonium formate 14.4 mol% | | Descriptor: | ETHANOLAMINE, FORMIC ACID, Lysozyme C | | Authors: | Han, Q, Darmanin, C, Drummond, C, Greaves, T. | | Deposit date: | 2021-08-27 | | Release date: | 2023-03-01 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.046 Å) | | Cite: | Probing ion-binding at a protein interface: Modulation of protein properties by ionic liquids.
J Colloid Interface Sci, 650, 2023
|
|
1GWA
 
 | | Triiodide derivative of porcine pancreas elastase | | Descriptor: | CALCIUM ION, ELASTASE 1, IODIDE ION, ... | | Authors: | Evans, G, Bricogne, G. | | Deposit date: | 2002-03-13 | | Release date: | 2002-06-06 | | Last modified: | 2019-05-08 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Triiodide Derivatization and Combinatorial Counter-Ion Replacement: Two Methods for Enhancing Phasing Signal Using Laboratory Cu Kalpha X-Ray Equipment
Acta Crystallogr.,Sect.D, 58, 2002
|
|
1H00
 
 | | CDK2 in complex with a disubstituted 4, 6-bis anilino pyrimidine CDK4 inhibitor | | Descriptor: | (2R)-1-[4-({6-[(2,6-DIFLUOROPHENYL)AMINO]PYRIMIDIN-4-YL}AMINO)PHENOXY]-3-(DIMETHYLAMINO)PROPAN-2-OL, (2S)-1-[4-({6-[(2,6-DIFLUOROPHENYL)AMINO]PYRIMIDIN-4-YL}AMINO)PHENOXY]-3-(DIMETHYLAMINO)PROPAN-2-OL, CELL DIVISION PROTEIN KINASE 2 | | Authors: | Beattie, J.F, Breault, G.A, Ellston, R.P.A, Green, S, Jewsbury, P.J, Midgley, C.J, Naven, R.T, Minshull, C.A, Pauptit, R.A, Tucker, J.A, Pease, J.E. | | Deposit date: | 2002-06-10 | | Release date: | 2003-07-11 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Cyclin-Dependent Kinase 4 Inhibitors as a Treatment for Cancer. Part 1: Identification and Optimisation of Substituted 4,6-Bis Anilino Pyrimidines
Bioorg.Med.Chem.Lett., 13, 2003
|
|
1GXA
 
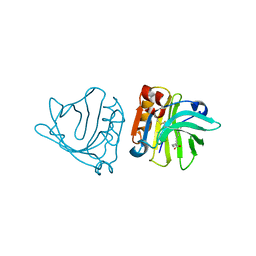 | |
1H0D
 
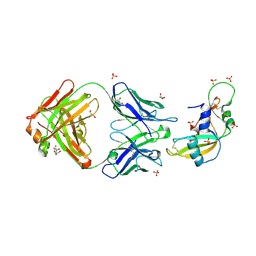 | | Crystal structure of Human Angiogenin in complex with Fab fragment of its monoclonal antibody mAb 26-2F | | Descriptor: | ANGIOGENIN, ANTIBODY FAB FRAGMENT, HEAVY CHAIN, ... | | Authors: | Chavali, G.B, Papageorgiou, A.C, Acharya, K.R. | | Deposit date: | 2002-06-19 | | Release date: | 2003-06-19 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The Crystal Structure of Human Angiogenin in Complex with an Antitumor Neutralizing Antibody
Structure, 11, 2003
|
|
7SLH
 
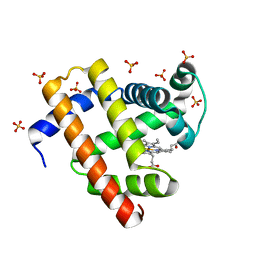 | | Engineered sperm whale myoglobin-based carbene transferase MbBTIC-C3 | | Descriptor: | Myoglobin, PROTOPORPHYRIN IX CONTAINING FE, SULFATE ION | | Authors: | Jenkins, J.L, Vargas, D, Ren, X, Fasan, R. | | Deposit date: | 2021-10-24 | | Release date: | 2023-05-03 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (1.15 Å) | | Cite: | Biocatalytic strategy for the construction of sp 3 -rich polycyclic compounds from directed evolution and computational modelling.
Nat.Chem., 16, 2024
|
|
7SLI
 
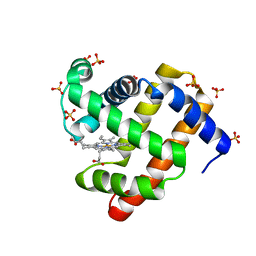 | | Engineered sperm whale myoglobin-based carbene transferase MbBTIC-C2 | | Descriptor: | Myoglobin, PROTOPORPHYRIN IX CONTAINING FE, SULFATE ION | | Authors: | Jenkins, J.L, Vargas, D, Ren, X, Fasan, R. | | Deposit date: | 2021-10-24 | | Release date: | 2023-05-03 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Biocatalytic strategy for the construction of sp 3 -rich polycyclic compounds from directed evolution and computational modelling.
Nat.Chem., 16, 2024
|
|
1GT1
 
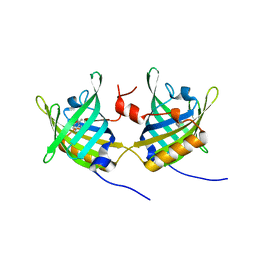 | | Complex of Bovine Odorant Binding Protein with Aminoanthracene and pyrazine | | Descriptor: | (3S)-1-octen-3-ol, 2-ISOBUTYL-3-METHOXYPYRAZINE, ANTHRACEN-1-YLAMINE, ... | | Authors: | Vincent, F, Ramoni, R, Spinelli, S, Grolli, S, Conti, V, Cambillau, C, Tegoni, M. | | Deposit date: | 2002-01-10 | | Release date: | 2003-10-03 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.71 Å) | | Cite: | Crystal Structures of Bovine Odorant-Binding Protein in Complex with Odorant Molecules.
Eur.J.Biochem., 271, 2004
|
|
1H8X
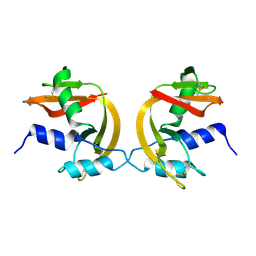 
 | | Domain-swapped Dimer of a Human Pancreatic Ribonuclease Variant | | Descriptor: | RIBONUCLEASE 1 | | Authors: | Canals, A, Pous, J, Guasch, A, Benito, A, Ribo, M, Vilanova, M, Coll, M. | | Deposit date: | 2001-02-16 | | Release date: | 2002-02-14 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The Structure of an Engineered Domain-Swapped Ribonuclease Dimer and its Implications for the Evolution of Proteins Toward Oligomerization
Structure, 9, 2001
|
|
1H1H
 
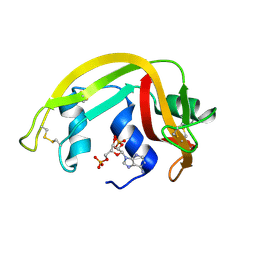 | | Crystal Structure of Eosinophil Cationic Protein in Complex with 2',5'-ADP at 2.0 A resolution Reveals the Details of the Ribonucleolytic Active site | | Descriptor: | ADENOSINE-2'-5'-DIPHOSPHATE, EOSINOPHIL CATIONIC PROTEIN | | Authors: | Mohan, C.G, Boix, E, Evans, H.R, Nikolovski, Z, Nogues, M.V, Cuchillo, C.M, Acharya, K.R. | | Deposit date: | 2002-07-15 | | Release date: | 2002-10-03 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The Crystal Structure of Eosinophil Cationic Protein in Complex with 2'5'-Adp at 2.0 A Resolution Reveals the Details of the Ribonucleolytic Active Site
Biochemistry, 41, 2002
|
|
1GWI
 
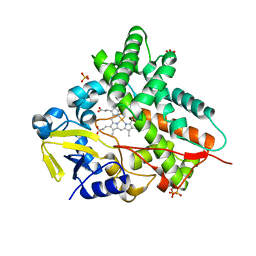 | | The 1.92 A structure of Streptomyces coelicolor A3(2) CYP154C1: A new monooxygenase that functionalizes macrolide ring systems | | Descriptor: | CYTOCHROME P450 154C1, PROTOPORPHYRIN IX CONTAINING FE, SULFATE ION | | Authors: | Podust, L.M, Kim, Y, Arase, M, Neely, B.A, Beck, B.J, Bach, H, Sherman, D.H, Lamb, D.C, Kelly, S.L, Waterman, M.R. | | Deposit date: | 2002-03-15 | | Release date: | 2003-01-29 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.92 Å) | | Cite: | The 1.92 A Structure of Streptomyces Coelicolor A3(2) Cyp154C1: A New Monooxygenase that Functionalizes Macrolide Ring Systems
J.Biol.Chem., 278, 2003
|
|
7SP5
 
 | | Crystal Structure of a Eukaryotic Phosphate Transporter | | Descriptor: | PHOSPHATE ION, Phosphate transporter, nonyl beta-D-glucopyranoside | | Authors: | Stroud, R.M, Pedersen, B.P, Kumar, H, Waight, A.B, Risenmay, A.J, Roe-Zurz, Z, Chau, B.H, Schlessinger, A, Bonomi, M, Harries, W, Sali, A, Johri, A.K, Finer-Moore, J. | | Deposit date: | 2021-11-02 | | Release date: | 2021-11-17 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Crystal structure of a eukaryotic phosphate transporter.
Nature, 496, 2013
|
|
1H5Z
 
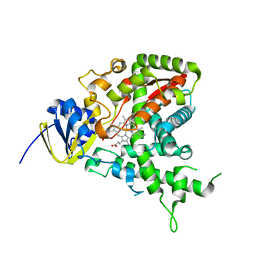 | |
1GV7
 
 | | ARH-I, an angiogenin/RNase A chimera | | Descriptor: | ANGIOGENIN, CITRIC ACID | | Authors: | Holloway, D.E, Shapiro, R, Hares, M.C, Leonidas, D.D, Acharya, K.R. | | Deposit date: | 2002-02-06 | | Release date: | 2002-08-22 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Guest-Host Crosstalk in an Angiogenin-RNase A Chimeric Protein
Biochemistry, 41, 2002
|
|
1HEN
 
 | |
7T3X
 
 | | Structure of unphosphorylated Pediculus humanus (Ph) PINK1 D334A mutant | | Descriptor: | Serine/threonine-protein kinase PINK1 | | Authors: | Gan, Z.Y, Leis, A, Dewson, G, Glukhova, A, Komander, D. | | Deposit date: | 2021-12-09 | | Release date: | 2021-12-22 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (3.53 Å) | | Cite: | Activation mechanism of PINK1.
Nature, 602, 2022
|
|
7SUF
 
 | | Structure of CHK1 10-pt. mutant complex with LRRK2 inhibitor 06 | | Descriptor: | 1,2-ETHANEDIOL, 8-cyclopropyl-N-[5-methyl-1-(oxan-4-yl)-1H-pyrazol-4-yl]quinazolin-2-amine, Serine/threonine-protein kinase Chk1 | | Authors: | Palte, R.L. | | Deposit date: | 2021-11-17 | | Release date: | 2022-01-12 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.48 Å) | | Cite: | Structure-Guided Discovery of Aminoquinazolines as Brain-Penetrant and Selective LRRK2 Inhibitors.
J.Med.Chem., 65, 2022
|
|
7SUJ
 
 | | Structure of CHK1 10-pt. mutant complex with LRRK2 inhibitor 24 | | Descriptor: | (3R,4R)-4-{4-[6-chloro-2-({1-[(1R)-2,2-difluorocyclopropyl]-5-methyl-1H-pyrazol-4-yl}amino)quinazolin-7-yl]piperidin-1-yl}-4-methyloxolan-3-ol, Serine/threonine-protein kinase Chk1 | | Authors: | Palte, R.L. | | Deposit date: | 2021-11-17 | | Release date: | 2022-01-12 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.299 Å) | | Cite: | Structure-Guided Discovery of Aminoquinazolines as Brain-Penetrant and Selective LRRK2 Inhibitors.
J.Med.Chem., 65, 2022
|
|
7T4K
 
 | | Structure of dimeric phosphorylated Pediculus humanus (Ph) PINK1 with kinked alpha-C helix in chain B | | Descriptor: | Serine/threonine-protein kinase PINK1, putative | | Authors: | Gan, Z.Y, Leis, A, Dewson, G, Glukhova, A, Komander, D. | | Deposit date: | 2021-12-10 | | Release date: | 2022-01-12 | | Last modified: | 2022-02-23 | | Method: | ELECTRON MICROSCOPY (3.25 Å) | | Cite: | Activation mechanism of PINK1.
Nature, 602, 2022
|
|
7T4N
 
 | | Structure of dimeric unphosphorylated Pediculus humanus (Ph) PINK1 D357A mutant | | Descriptor: | Serine/threonine-protein kinase PINK1, putative | | Authors: | Gan, Z.Y, Leis, A, Dewson, G, Glukhova, A, Komander, D. | | Deposit date: | 2021-12-10 | | Release date: | 2022-01-12 | | Last modified: | 2024-02-28 | | Method: | ELECTRON MICROSCOPY (2.35 Å) | | Cite: | Activation mechanism of PINK1.
Nature, 602, 2022
|
|
7SUH
 
 | | Structure of CHK1 10-pt. mutant complex with LRRK2 inhibitor 15 | | Descriptor: | 1-[5-chloro-4-({6-chloro-7-[1-(oxetan-3-yl)piperidin-4-yl]quinazolin-2-yl}amino)-1H-pyrazol-1-yl]-2-methylpropan-2-ol, Serine/threonine-protein kinase Chk1 | | Authors: | Palte, R.L. | | Deposit date: | 2021-11-17 | | Release date: | 2022-01-12 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.46 Å) | | Cite: | Structure-Guided Discovery of Aminoquinazolines as Brain-Penetrant and Selective LRRK2 Inhibitors.
J.Med.Chem., 65, 2022
|
|
7T4L
 
 | | Structure of dimeric phosphorylated Pediculus humanus (Ph) PINK1 with extended alpha-C helix in chain B | | Descriptor: | Serine/threonine-protein kinase PINK1, putative | | Authors: | Gan, Z.Y, Leis, A, Dewson, G, Glukhova, A, Komander, D. | | Deposit date: | 2021-12-10 | | Release date: | 2022-01-12 | | Last modified: | 2022-02-23 | | Method: | ELECTRON MICROSCOPY (3.28 Å) | | Cite: | Activation mechanism of PINK1.
Nature, 602, 2022
|
|
