6USM
 
 | |
8PKN
 
 | |
9EBO
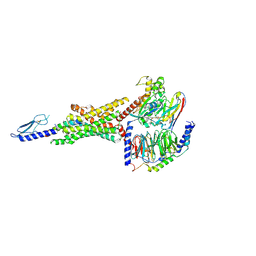 
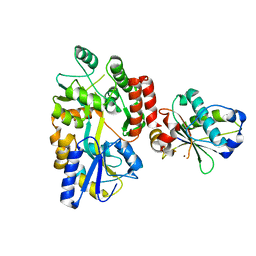
 | | Peptide 2 (GLP-1 (ACPC18)) bound to GLP-1R/Gs complex (conformer 1) | | Descriptor: | Glucagon-like peptide 1 receptor, Glucagon-like peptide 1(7-36), Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, ... | | Authors: | Cary, B.P, Hager, M.V, Mariam, Z, Morris, R.K, Belousoff, M.J, Deganutti, G, Wootten, D, Sexton, P.M, Gellman, S.H. | | Deposit date: | 2024-11-12 | | Release date: | 2025-03-26 | | Last modified: | 2025-04-16 | | Method: | ELECTRON MICROSCOPY (3.13 Å) | | Cite: | Prolonged signaling of backbone-modified glucagon-like peptide- 1 analogues with diverse receptor trafficking.
Proc.Natl.Acad.Sci.USA, 122, 2025
|
|
9EBN
 
 | | Peptide 1 (GLP-1 (Aib16, ACPC18)) bound to GLP-1R/Gs complex | | Descriptor: | Glucagon-like peptide 1 receptor, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1, ... | | Authors: | Cary, B.P, Hager, M.V, Mariam, Z, Morris, R.K, Belousoff, M.J, Deganutti, G, Wootten, D, Sexton, P.M, Gellman, S.H. | | Deposit date: | 2024-11-12 | | Release date: | 2025-03-26 | | Last modified: | 2025-04-16 | | Method: | ELECTRON MICROSCOPY (3.44 Å) | | Cite: | Prolonged signaling of backbone-modified glucagon-like peptide- 1 analogues with diverse receptor trafficking.
Proc.Natl.Acad.Sci.USA, 122, 2025
|
|
9EBQ
 
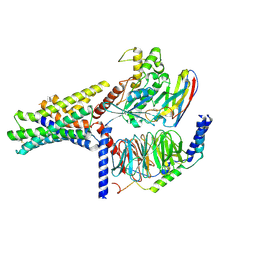 | | Peptide 2 (GLP-1 (ACPC18)) bound to GLP-1R/Gs complex (conformer 2) | | Descriptor: | Glucagon-like peptide 1 receptor, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1, ... | | Authors: | Cary, B.P, Hager, M.V, Mariam, Z, Morris, R.K, Belousoff, M.J, Deganutti, G, Wootten, D, Sexton, P.M, Gellman, S.H. | | Deposit date: | 2024-11-12 | | Release date: | 2025-03-26 | | Last modified: | 2025-04-16 | | Method: | ELECTRON MICROSCOPY (3.16 Å) | | Cite: | Prolonged signaling of backbone-modified glucagon-like peptide- 1 analogues with diverse receptor trafficking.
Proc.Natl.Acad.Sci.USA, 122, 2025
|
|
8VSM
 
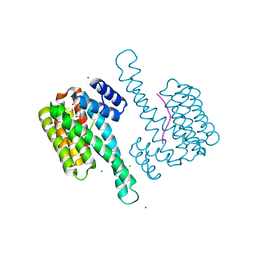 | | Ternary structure of 14-3-3 sigma, ARAF phosphopeptide (pS214) and compound 78 (1124378) | | Descriptor: | 1-[8-(4-bromophenyl)sulfonyl-5-oxa-2,8-diazaspiro[3.5]nonan-2-yl]-2-chloranyl-ethanone, 14-3-3 protein sigma, CHLORIDE ION, ... | | Authors: | Vickery, H.R, Virta, J.M, Pennings, M, Konstantinidou, M, van den Oetelaar, M, Neitz, R.J, Ottmann, C, Brunsveld, L, Arkin, M.R. | | Deposit date: | 2024-01-24 | | Release date: | 2025-03-19 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Ternary structure of 14-3-3 sigma, ARAF phosphopeptide (pS214) and compound 78 (1124378)
To Be Published
|
|
8VSN
 
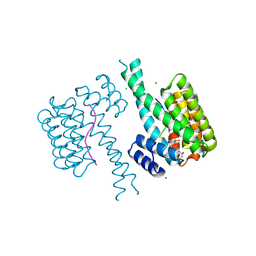 | | Ternary structure of 14-3-3 sigma, ARAF phosphopeptide (pS214) and compound 79 (1124379) | | Descriptor: | 14-3-3 protein sigma, 2-chloranyl-1-[8-(4-iodophenyl)sulfonyl-5-oxa-2,8-diazaspiro[3.5]nonan-2-yl]ethanone, CHLORIDE ION, ... | | Authors: | Vickery, H.R, Virta, J.M, Pennings, M, Konstantinidou, M, van den Oetelaar, M, Neitz, R.J, Ottmann, C, Brunsveld, L, Arkin, M.R. | | Deposit date: | 2024-01-24 | | Release date: | 2025-03-19 | | Method: | X-RAY DIFFRACTION (1.91 Å) | | Cite: | Ternary structure of 14-3-3 sigma, ARAF phosphopeptide (pS214) and compound 79 (1124379)
To Be Published
|
|
8VSL
 
 | | Binary structure of 14-3-3 sigma and ARAF phosphopeptide (pS214) | | Descriptor: | 14-3-3 protein sigma, CHLORIDE ION, MAGNESIUM ION, ... | | Authors: | Vickery, H.R, Virta, J.M, Pennings, M, Konstantinidou, M, van den Oetelaar, M, Neitz, R.J, Ottmann, C, Brunsveld, L, Arkin, M.R. | | Deposit date: | 2024-01-24 | | Release date: | 2025-03-19 | | Method: | X-RAY DIFFRACTION (1.42 Å) | | Cite: | Binary structure of 14-3-3 sigma and ARAF phosphopeptide (pS214)
To Be Published
|
|
8VSO
 
 | | Ternary structure of 14-3-3 sigma, BRAF phosphopeptide (pS365) and compound 78 (1124378) | | Descriptor: | 1-[8-(4-bromophenyl)sulfonyl-5-oxa-2,8-diazaspiro[3.5]nonan-2-yl]-2-chloranyl-ethanone, 14-3-3 protein sigma, CHLORIDE ION, ... | | Authors: | Vickery, H.R, Virta, J.M, Pennings, M, Konstantinidou, M, van den Oetelaar, M, Neitz, R.J, Ottmann, C, Brunsveld, L, Arkin, M.R. | | Deposit date: | 2024-01-24 | | Release date: | 2025-03-19 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Ternary structure of 14-3-3 sigma, BRAF phosphopeptide (pS365) and compound 78 (1124378)
To Be Published
|
|
2KTV
 
 | |
8S6B
 
 | |
8PP2
 
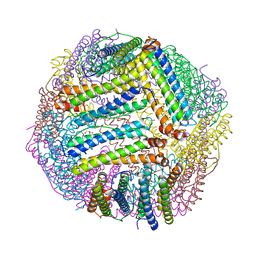 | |
5ZVQ
 
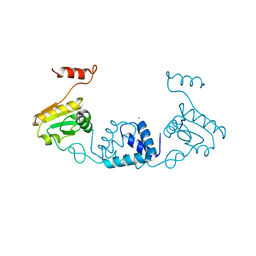 | |
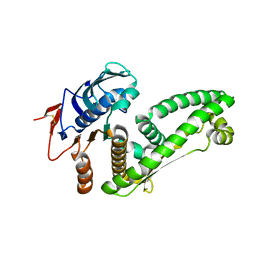
5ZWU
 
 | |
5IG5
 
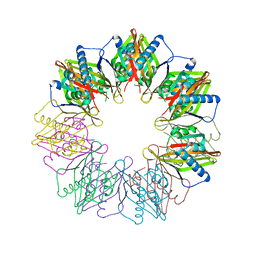 | |
5IG0
 
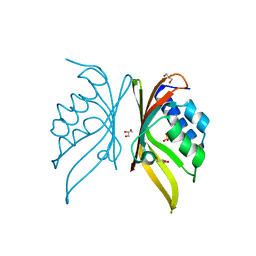 | | Crystal structure of S. rosetta CaMKII hub | | Descriptor: | CAMK/CAMK2 protein kinase, GLYCEROL, SULFATE ION | | Authors: | Bhattacharyya, M, Gee, C.L, Barros, T, Kuriyan, J. | | Deposit date: | 2016-02-26 | | Release date: | 2016-03-23 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Molecular mechanism of activation-triggered subunit exchange in Ca(2+)/calmodulin-dependent protein kinase II.
Elife, 5, 2016
|
|
5IG4
 
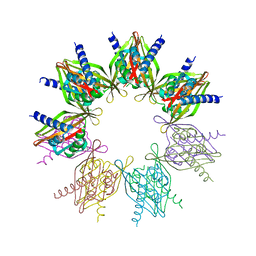 | | Crystal structure of N. vectensis CaMKII-A hub | | Descriptor: | GLYCEROL, Predicted protein | | Authors: | Bhattacharyya, M, Pappireddi, N, Gee, C.L, Barros, T, Kuriyan, J. | | Deposit date: | 2016-02-26 | | Release date: | 2016-03-23 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Molecular mechanism of activation-triggered subunit exchange in Ca(2+)/calmodulin-dependent protein kinase II.
Elife, 5, 2016
|
|
5IG1
 
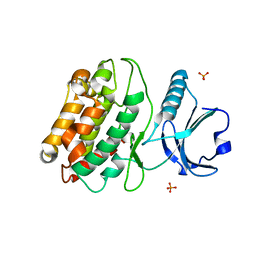 | | Crystal structure of S. rosetta CaMKII kinase domain | | Descriptor: | CAMK/CAMK2 protein kinase, PHOSPHATE ION | | Authors: | Bhattacharyya, M, Gee, C.L, Barros, T, Kuriyan, J. | | Deposit date: | 2016-02-26 | | Release date: | 2016-03-23 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Molecular mechanism of activation-triggered subunit exchange in Ca(2+)/calmodulin-dependent protein kinase II.
Elife, 5, 2016
|
|
3TF2
 
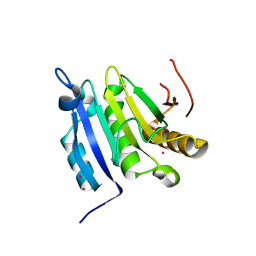 | | Crystal structure of the cap free human translation initiation factor eIF4E | | Descriptor: | Eukaryotic translation initiation factor 4E, UNKNOWN ATOM OR ION | | Authors: | Siddiqui, N, Tempel, W, Nedyalkova, L, Wernimont, A.K, Arrowsmith, C.H, Edwards, A.M, Bountra, C, Weigelt, J, Borden, K.L.B, Park, H, Structural Genomics Consortium (SGC) | | Deposit date: | 2011-08-15 | | Release date: | 2011-08-31 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structural insights into the allosteric effects of 4EBP1 on the eukaryotic translation initiation factor eIF4E.
J.Mol.Biol., 415, 2012
|
|
6BGE
 
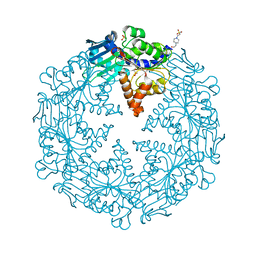 | | HELICOBACTER PYLORI ATPASE, HP0525, IN COMPLEX WITH 1G2 COMPOUND | | Descriptor: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, 4-[(pyridin-2-yl)oxy]benzoic acid, GLYCEROL, ... | | Authors: | Arya, T, Casu, B, Baron, C. | | Deposit date: | 2017-10-27 | | Release date: | 2018-10-31 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | CagAlpha in complex with hexamer inhibitor
To Be Published
|
|
4BTS
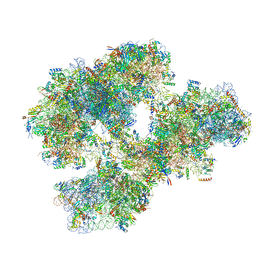 
 | | THE CRYSTAL STRUCTURE OF THE EUKARYOTIC 40S RIBOSOMAL SUBUNIT IN COMPLEX WITH EIF1 AND EIF1A | | Descriptor: | 18S ribosomal RNA, 40S RIBOSOMAL PROTEIN RACK1, 40S RIBOSOMAL PROTEIN RPS10E, ... | | Authors: | Weisser, M, Voigts-Hoffmann, F, Rabl, J, Leibundgut, M, Ban, N. | | Deposit date: | 2013-06-19 | | Release date: | 2013-07-17 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (3.703 Å) | | Cite: | The crystal structure of the eukaryotic 40S ribosomal subunit in complex with eIF1 and eIF1A.
Nat. Struct. Mol. Biol., 20, 2013
|
|
5JBS
 
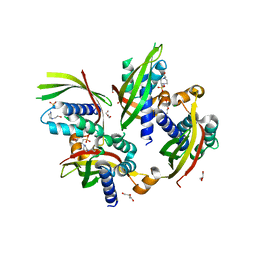 | | Conformational changes during monomer-to-dimer transition of Brucella suis VirB8 | | Descriptor: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, CHLORIDE ION, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Arya, T, Sharifahmadian, M, Sygusch, J, Baron, B. | | Deposit date: | 2016-04-13 | | Release date: | 2017-03-01 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | NMR analyses, X-ray crystallography and small-molecule probing reveal conformational shifts during monomer-to-dimer transition of Brucella suis VirB8
To Be Published
|
|
6E8F
 
 | |
6EB5
 | |
2GOO
 
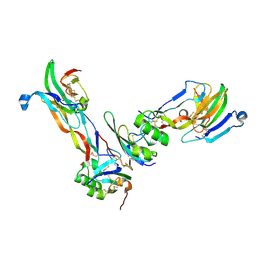 | | Ternary Complex of BMP-2 bound to BMPR-Ia-ECD and ActRII-ECD | | Descriptor: | 2-acetamido-2-deoxy-alpha-D-glucopyranose, Activin receptor type 2A, Bone morphogenetic protein 2, ... | | Authors: | Allendorph, G.P, Choe, S. | | Deposit date: | 2006-04-13 | | Release date: | 2006-05-09 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structure of the ternary signaling complex of a TGF-beta superfamily member.
Proc.Natl.Acad.Sci.Usa, 103, 2006
|
|
