3OLM
 
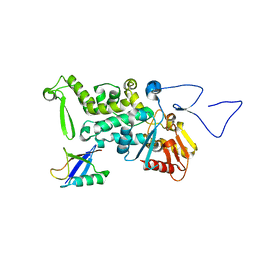 | | Structure and Function of a Ubiquitin Binding Site within the Catalytic Domain of a HECT Ubiquitin Ligase | | Descriptor: | E3 ubiquitin-protein ligase RSP5, Ubiquitin | | Authors: | Kim, H.C, Steffen, A, Chen, J, Huibregtse, J.M. | | Deposit date: | 2010-08-26 | | Release date: | 2011-03-23 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.495 Å) | | Cite: | Structure and function of a HECT domain ubiquitin-binding site.
Embo Rep., 12, 2011
|
|
4LCD
 
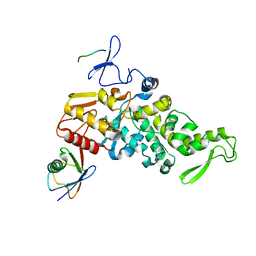 | |
3IHP
 
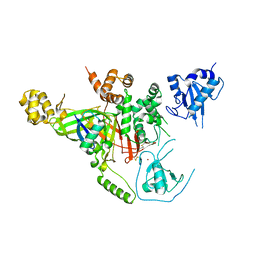 | | Covalent Ubiquitin-Usp5 Complex | | Descriptor: | CHLORIDE ION, ETHANAMINE, Ubiquitin, ... | | Authors: | Walker, J.R, Avvakumov, G.V, Xue, S, Butler-Cole, C, Weigelt, J, Bountra, C, Arrowsmith, C.H, Edwards, A.M, Bochkarev, A, Dhe-Paganon, S, Structural Genomics Consortium (SGC) | | Deposit date: | 2009-07-30 | | Release date: | 2009-12-29 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Covalent Ubiquitin-Usp5 Complex
To be Published
|
|
5GML
 
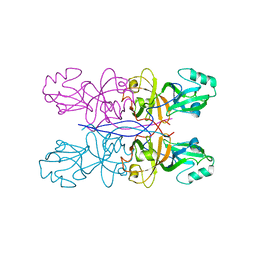 | |
1WDZ
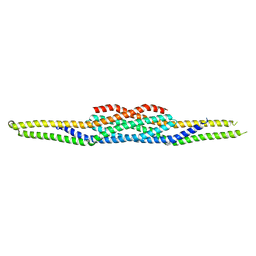 
 | | Crystal structure of RCB domain of IRSp53 | | Descriptor: | insulin receptor substrate p53 | | Authors: | Murayama, K, Suetsugu, S, Seto, A, Shirouzu, M, Takenawa, T, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2004-05-21 | | Release date: | 2005-06-07 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.63 Å) | | Cite: | Crystal structure of RCB domain of IRSp53
TO BE PUBLISHED
|
|
1P5X
 
 | | STRUCTURE OF THE D55N MUTANT OF PHOSPHOLIPASE C FROM BACILLUS CEREUS | | Descriptor: | Phospholipase C, ZINC ION | | Authors: | Antikainen, N.M, Monzingo, A.F, Franklin, C.L, Robertus, J.D, Martin, S.F. | | Deposit date: | 2003-04-28 | | Release date: | 2003-09-30 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Using X-ray crystallography of the Asp55Asn mutant of the phosphatidylcholine-preferring phospholipase C from Bacillus cereus to support the mechanistic role of Asp55 as the general base.
Arch.Biochem.Biophys., 417, 2003
|
|
1P6E
 
 | | STRUCTURE OF THE D55N MUTANT OF PHOSPHOLIPASE C FROM BACILLUS CEREUS IN COMPLEX WITH 1,2-DI-N-PENTANOYL-SN-GLYCERO-3-DITHIOPHOSPHOCHOLINE | | Descriptor: | 1,2-DI-N-PENTANOYL-SN-GLYCERO-3-DITHIOPHOSPHOCHOLINE, Phospholipase C, ZINC ION | | Authors: | Antikainen, N.M, Monzingo, A.F, Franklin, C.L, Robertus, J.D, Martin, S.F. | | Deposit date: | 2003-04-29 | | Release date: | 2003-09-30 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Using X-ray crystallography of the Asp55Asn mutant of the phosphatidylcholine-preferring phospholipase C from Bacillus cereus to support the mechanistic role of Asp55 as the general base.
Arch.Biochem.Biophys., 417, 2003
|
|
1P6D
 
 | | STRUCTURE OF THE D55N MUTANT OF PHOSPHOLIPASE C FROM BACILLUS CEREUS IN COMPLEX WITH (3S)-3,4,DI-N-HEXANOYLOXYBUTYL-1-PHOSPHOCHOLINE | | Descriptor: | (3S)-3,4-DI-N-HEXANOYLOXYBUTYL-1-PHOSPHOCHOLINE, PHOSPHOLIPASE C, ZINC ION | | Authors: | Antikainen, N.M, Monzingo, A.F, Franklin, C.L, Robertus, J.D, Martin, S.F. | | Deposit date: | 2003-04-29 | | Release date: | 2003-09-30 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Using X-ray crystallography of the Asp55Asn mutant of the phosphatidylcholine-preferring phospholipase C from Bacillus cereus to support the mechanistic role of Asp55 as the general base.
Arch.Biochem.Biophys., 417, 2003
|
|
3RNJ
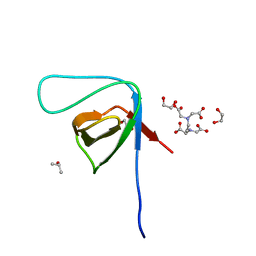 
 | | Crystal structure of the SH3 domain from IRSp53 (BAIAP2) | | Descriptor: | 1,2-ETHANEDIOL, Brain-specific angiogenesis inhibitor 1-associated protein 2, ISOPROPYL ALCOHOL, ... | | Authors: | Simister, P.C, Barilari, M, Muniz, J.R.C, Dente, L, Knapp, S, von Delft, F, Filippakopoulos, P, Vollmar, M, Chaikuad, A, Raynor, J, Tregubova, A, Arrowsmith, C.H, Edwards, A.M, Weigelt, J, Bountra, C, Feller, S.M, Structural Genomics Consortium (SGC) | | Deposit date: | 2011-04-22 | | Release date: | 2011-05-04 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Crystal structure of the SH3 domain from IRSp53 (BAIAP2)
To be Published
|
|
2D5K
 
 | | Crystal structure of Dps from Staphylococcus aureus | | Descriptor: | Dps family protein, GLYCEROL | | Authors: | Tanaka, Y, Yao, M, Watanabe, N, Tanaka, I. | | Deposit date: | 2005-11-02 | | Release date: | 2006-10-17 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Nucleoid compaction by MrgA(Asp56Ala/Glu60Ala) does not contribute to staphylococcal cell survival against oxidative stress and phagocytic killing by macrophages
FEMS Microbiol. Lett., 360, 2014
|
|
2G6Z
 
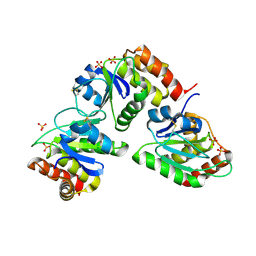 | | Crystal structure of human DUSP5 | | Descriptor: | Dual specificity protein phosphatase 5, SULFATE ION | | Authors: | Kim, S.J, Ryu, S.E. | | Deposit date: | 2006-02-26 | | Release date: | 2007-01-16 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Crystal structure of the catalytic domain of human DUSP5, a dual specificity MAP kinase protein phosphatase
Proteins, 66, 2007
|
|
4XY5
 
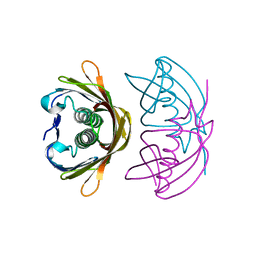 | |
3A3Q
 
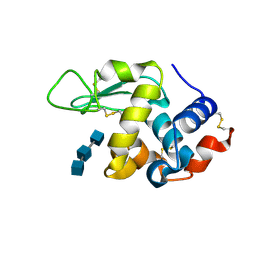 | | Structure of N59D HEN EGG-WHITE LYSOZYME in complex with (GlcNAc)3 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Lysozyme C | | Authors: | Ose, T, Kuroki, K, Matsushima, M, Maenaka, K, Kumagai, I. | | Deposit date: | 2009-06-16 | | Release date: | 2009-11-03 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Importance of the hydrogen bonding network including Asp52 for catalysis, as revealed by Asn59 mutant hen egg-white lysozymes
J.Biochem., 146, 2009
|
|
3A3R
 
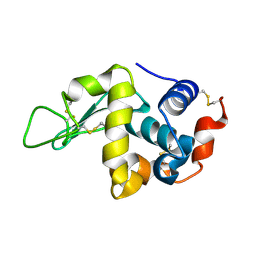 | | Structure of N59D HEN EGG-WHITE LYSOZYME | | Descriptor: | Lysozyme C | | Authors: | Ose, T, Kuroki, K, Matsushima, M, Maenaka, K, Kumagai, I. | | Deposit date: | 2009-06-16 | | Release date: | 2009-11-03 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Importance of the hydrogen bonding network including Asp52 for catalysis, as revealed by Asn59 mutant hen egg-white lysozymes
J.Biochem., 146, 2009
|
|
8C61
 
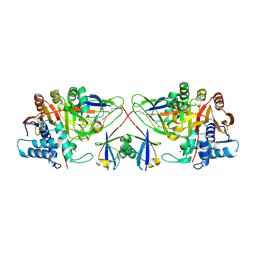 | |
6P2I
 
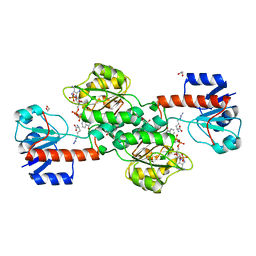 | | Acyclic imino acid reductase (Bsp5) in complex with NADPH and D-Arg | | Descriptor: | 1,2-ETHANEDIOL, D-ARGININE, Glycerate dehydrogenase, ... | | Authors: | Guo, J, Higgins, M.A, Daniel-Ivad, P, Ryan, K.S. | | Deposit date: | 2019-05-21 | | Release date: | 2019-07-24 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.63 Å) | | Cite: | An Asymmetric Reductase That Intercepts Acyclic Imino Acids Producedin Situby a Partner Oxidase.
J.Am.Chem.Soc., 141, 2019
|
|
7O46
 
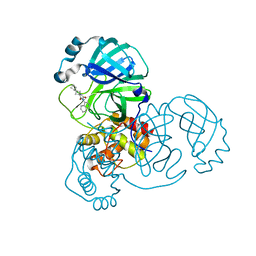 | | Crystal Structure of SARS-CoV-2 main protease (Nsp5) in complex with compound 17 | | Descriptor: | 2-cyclobutyl-7-isoquinolin-4-yl-5,7-diazaspiro[3.4]octane-6,8-dione, 3C-like proteinase nsp5 | | Authors: | Talibov, V.O. | | Deposit date: | 2021-04-05 | | Release date: | 2022-02-23 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.23 Å) | | Cite: | Ultralarge Virtual Screening Identifies SARS-CoV-2 Main Protease Inhibitors with Broad-Spectrum Activity against Coronaviruses.
J.Am.Chem.Soc., 144, 2022
|
|
8FIG
 
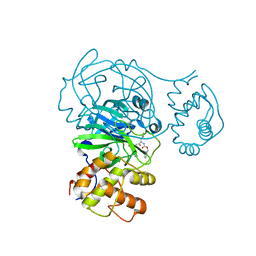 | |
4AQK
 
 | | Inositol 1,3,4,5,6-pentakisphosphate 2-kinase in complex with ADP and IP6 | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, INOSITOL HEXAKISPHOSPHATE, INOSITOL-PENTAKISPHOSPHATE 2-KINASE, ... | | Authors: | I Banos-Sanz, J, Sanz-Aparicio, J, Gonzalez, B. | | Deposit date: | 2012-04-18 | | Release date: | 2012-05-02 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Expression, Purification, Crystallization and Preliminary X-Ray Diffraction Analysis of the Apo Form of Insp5 2-K from Arabidopsis Thaliana.
Acta Crystallogr.,Sect.F, 68, 2012
|
|
8OKB
 
 | | SARS-CoV2 NSP5 in complex with a peptidomimetic ligand | | Descriptor: | 3C-like proteinase nsp5, methyl (4~{S})-4-[[(2~{S})-4-methyl-2-(phenylmethoxycarbonylamino)pentanoyl]amino]-5-[(3~{S})-2-oxidanylidenepyrrolidin-3-yl]pentanoate | | Authors: | Calderone, V. | | Deposit date: | 2023-03-28 | | Release date: | 2024-01-17 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.31 Å) | | Cite: | Development of a GC-376 Based Peptidomimetic PROTAC as a Degrader of 3-Chymotrypsin-like Protease of SARS-CoV-2.
Acs Med.Chem.Lett., 15, 2024
|
|
8OKC
 
 | | SARS-CoV2 NSP5 in complex with a GC-376 based peptidomimetic PROTAC | | Descriptor: | (phenylmethyl) ~{N}-[(2~{R})-1-[[(~{Z},2~{S})-5-[4-[[1-[2-[(3~{R})-2,6-bis(oxidanylidene)piperidin-3-yl]-6-fluoranyl-1,3-bis(oxidanylidene)isoindol-5-yl]piperidin-4-yl]methyl]piperazin-1-yl]-5-oxidanylidene-1-[(3~{R})-2-oxidanylidenepyrrolidin-3-yl]pent-3-en-2-yl]amino]-4-methyl-1-oxidanylidene-pentan-2-yl]carbamate, 3C-like proteinase nsp5 | | Authors: | Calderone, V. | | Deposit date: | 2023-03-28 | | Release date: | 2024-01-17 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Development of a GC-376 Based Peptidomimetic PROTAC as a Degrader of 3-Chymotrypsin-like Protease of SARS-CoV-2.
Acs Med.Chem.Lett., 15, 2024
|
|
3RLE
 
 | | Crystal Structure of GRASP55 GRASP domain (residues 7-208) | | Descriptor: | Golgi reassembly-stacking protein 2 | | Authors: | Truschel, S.T, Sengupta, D, Foote, A, Heroux, A, Macbeth, M.R, Linstedt, A.D. | | Deposit date: | 2011-04-19 | | Release date: | 2011-05-04 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.649 Å) | | Cite: | Structure of the Membrane-tethering GRASP Domain Reveals a Unique PDZ Ligand Interaction That Mediates Golgi Biogenesis.
J.Biol.Chem., 286, 2011
|
|
8EJ7
 
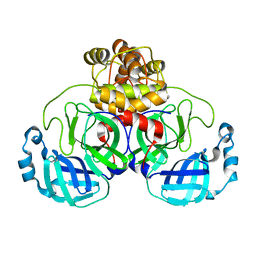 | |
6J2H
 
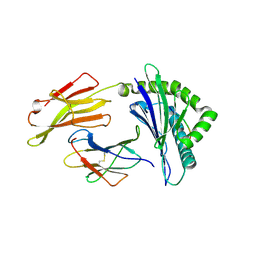 | | Crystal structure of bat (Pteropus Alecto) MHC class I Ptal-N*01:01 mutant (Met52 Asp53 Leu54 deleted) in complex with Hendra virus-derived peptide HeV1 | | Descriptor: | Beta-2-microglobulin, HeV1, Ptal-N*01:01 (Met52 Asp53 Leu54 deleted) | | Authors: | Lu, D, Liu, K.F, Yue, C, Lu, Q, Cheng, H, Chai, Y, Qi, J.X, Gao, G.F, Liu, W.J. | | Deposit date: | 2019-01-01 | | Release date: | 2019-09-18 | | Last modified: | 2019-12-04 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Peptide presentation by bat MHC class I provides new insight into the antiviral immunity of bats.
Plos Biol., 17, 2019
|
|
6ZEI
 
 | | Structure of PP1-IRSp53 S455E chimera [PP1(7-304) + linker (G/S)x9 + IRSp53(449-465)] bound to Phactr1 (516-580) | | Descriptor: | GLYCEROL, MANGANESE (II) ION, PHOSPHATE ION, ... | | Authors: | Mouilleron, S, Treisman, R, Fedoryshchak, R, Lee, R, Butler, A.M, Prechova, M. | | Deposit date: | 2020-06-16 | | Release date: | 2020-09-30 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.39 Å) | | Cite: | Molecular basis for substrate specificity of the Phactr1/PP1 phosphatase holoenzyme.
Elife, 9, 2020
|
|
