1RDI
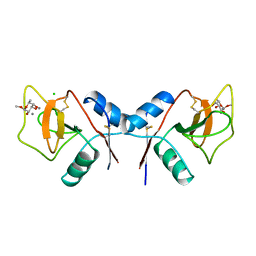 
 | | MANNOSE-BINDING PROTEIN, SUBTILISIN DIGEST FRAGMENT COMPLEX WITH ALPHA-METHYL-L-FUCOPYRANOSIDE | | Descriptor: | CALCIUM ION, CHLORIDE ION, MANNOSE-BINDING PROTEIN-C, ... | | Authors: | Ng, K.K.-S, Drickamer, K, Weis, W.I. | | Deposit date: | 1995-09-05 | | Release date: | 1996-03-08 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural analysis of monosaccharide recognition by rat liver mannose-binding protein.
J.Biol.Chem., 271, 1996
|
|
1CBY
 
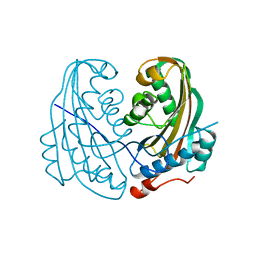 | | DELTA-ENDOTOXIN | | Descriptor: | DELTA-ENDOTOXIN CYTB | | Authors: | Li, J. | | Deposit date: | 1995-09-05 | | Release date: | 1996-10-14 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structure of the mosquitocidal delta-endotoxin CytB from Bacillus thuringiensis sp. kyushuensis and implications for membrane pore formation.
J.Mol.Biol., 257, 1996
|
|
1RNG
 
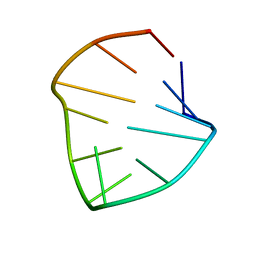 | |
232D
 
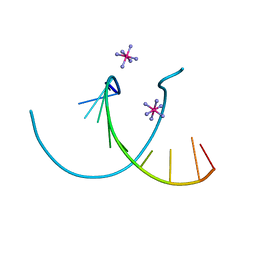 | |
1ANJ
 
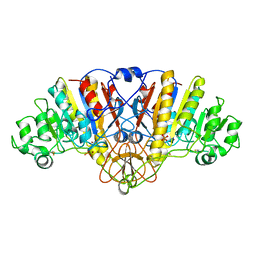 | | ALKALINE PHOSPHATASE (K328H) | | Descriptor: | ALKALINE PHOSPHATASE, PHOSPHATE ION, ZINC ION | | Authors: | Murphy, J.E, Tibbitts, T.T, Kantrowitz, E.R. | | Deposit date: | 1995-09-06 | | Release date: | 1996-01-29 | | Last modified: | 2021-11-03 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Mutations at positions 153 and 328 in Escherichia coli alkaline phosphatase provide insight towards the structure and function of mammalian and yeast alkaline phosphatases.
J.Mol.Biol., 253, 1995
|
|
1ANI
 
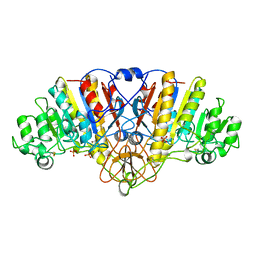 | | ALKALINE PHOSPHATASE (D153H, K328H) | | Descriptor: | ALKALINE PHOSPHATASE, PHOSPHATE ION, ZINC ION | | Authors: | Murphy, J.E, Tibbitts, T.T, Kantrowitz, E.R. | | Deposit date: | 1995-09-06 | | Release date: | 1996-01-29 | | Last modified: | 2021-11-03 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Mutations at positions 153 and 328 in Escherichia coli alkaline phosphatase provide insight towards the structure and function of mammalian and yeast alkaline phosphatases.
J.Mol.Biol., 253, 1995
|
|
1GBD
 
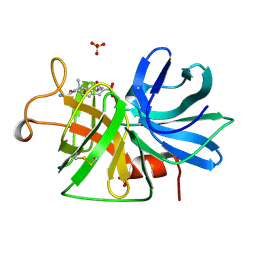 | |
1GBF
 
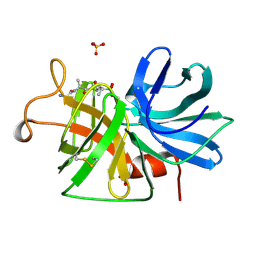 | |
1CWD
 
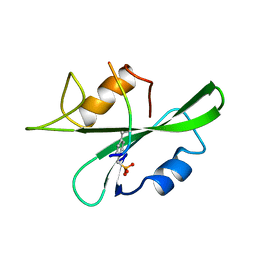 | | HUMAN P56LCK TYROSINE KINASE COMPLEXED WITH PHOSPHOPEPTIDE | | Descriptor: | (PHOSPHONOMETHYL)PHENYLALANINE-CONTAINING PEPTIDE PRO-GLU-GLY-ASP-PM3-GLU-GLU-VAL-LEU, P56LCK TYROSINE KINASE | | Authors: | Mikol, V. | | Deposit date: | 1995-09-06 | | Release date: | 1996-12-07 | | Last modified: | 2024-06-05 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | The crystal structures of the SH2 domain of p56lck complexed with two phosphopeptides suggest a gated peptide binding site.
J.Mol.Biol., 246, 1995
|
|
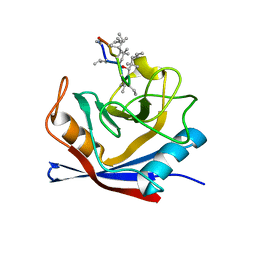
1CWC
 | |
1CWE
 
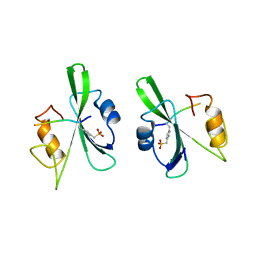 | | HUMAN P56LCK TYROSINE KINASE COMPLEXED WITH PHOSPHOPEPTIDE | | Descriptor: | P56LCK TYROSINE KINASE, PHOSPHOPEPTIDE ACQ-PMP-GLU-GLU-ILE-PRO | | Authors: | Mikol, V. | | Deposit date: | 1995-09-06 | | Release date: | 1996-12-07 | | Last modified: | 2024-06-05 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | The crystal structures of the SH2 domain of p56lck complexed with two phosphopeptides suggest a gated peptide binding site.
J.Mol.Biol., 246, 1995
|
|
1CWA
 
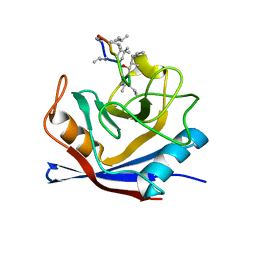 | |
1GBL
 
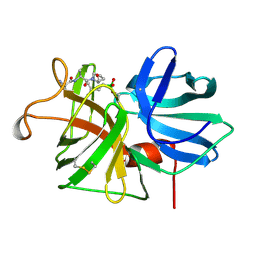 | |
1GBI
 
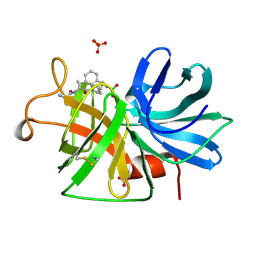 | |
1GBJ
 
 | | ALPHA-LYTIC PROTEASE WITH MET 190 REPLACED BY ALA | | Descriptor: | ALPHA-LYTIC PROTEASE, SULFATE ION | | Authors: | Mace, J.E, Agard, D.A. | | Deposit date: | 1995-09-06 | | Release date: | 1996-01-29 | | Last modified: | 2021-11-03 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Kinetic and structural characterization of mutations of glycine 216 in alpha-lytic protease: a new target for engineering substrate specificity.
J.Mol.Biol., 254, 1995
|
|
1GBC
 
 | |
1GBH
 
 | |
1GBE
 
 | |
1FVL
 
 | |
2ANH
 
 | | ALKALINE PHOSPHATASE (D153H) | | Descriptor: | ALKALINE PHOSPHATASE, PHOSPHATE ION, ZINC ION | | Authors: | Murphy, J.E, Tibbitts, T.T, Kantrowitz, E.R. | | Deposit date: | 1995-09-06 | | Release date: | 1996-01-29 | | Last modified: | 2021-11-03 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Mutations at positions 153 and 328 in Escherichia coli alkaline phosphatase provide insight towards the structure and function of mammalian and yeast alkaline phosphatases.
J.Mol.Biol., 253, 1995
|
|
1GBK
 
 | |
1GBB
 
 | |
1GBM
 
 | |
1CWB
 
 | |
1GBA
 
 | |
