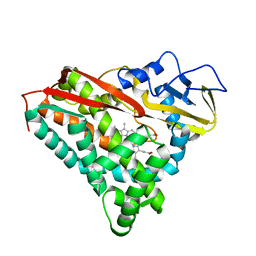
1PHC
 
 | |
1YL0
 
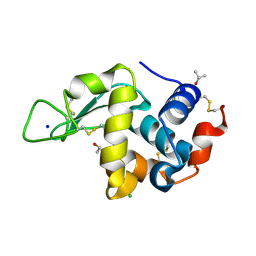 | | Effect of alcohols on protein hydration | | Descriptor: | CHLORIDE ION, ISOPROPYL ALCOHOL, Lysozyme C, ... | | Authors: | Deshpande, A, Nimsadkar, S, Mande, S.C. | | Deposit date: | 2005-01-18 | | Release date: | 2005-07-05 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Effect of alcohols on protein hydration: crystallographic analysis of hen egg-white lysozyme in the presence of alcohols.
Acta Crystallogr.,Sect.D, 61, 2005
|
|
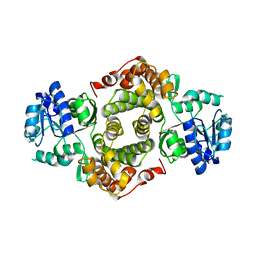
1YJ8
 
 | |
4IID
 
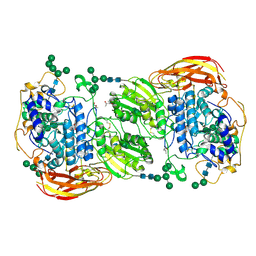 | | Crystal structure of beta-glucosidase 1 from Aspergillus aculeatus in complex with 1-deoxynojirimycin | | Descriptor: | (4R)-2-METHYLPENTANE-2,4-DIOL, (4S)-2-METHYL-2,4-PENTANEDIOL, 1-DEOXYNOJIRIMYCIN, ... | | Authors: | Suzuki, K, Sumitani, J, Kawaguchi, T, Fushinobu, S. | | Deposit date: | 2012-12-20 | | Release date: | 2013-04-10 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal structures of glycoside hydrolase family 3 beta-glucosidase 1 from Aspergillus aculeatus
Biochem.J., 452, 2013
|
|
3WPL
 
 | |
1YKP
 
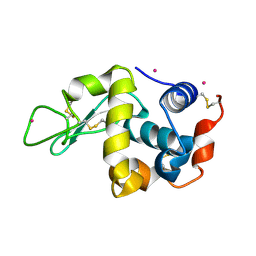
 | | Protocatechuate 3,4-Dioxygenase Y408H mutant bound to DHB | | Descriptor: | 3,4-DIHYDROXYBENZOIC ACID, FE (III) ION, Protocatechuate 3,4-dioxygenase alpha chain, ... | | Authors: | Brown, C.K, Ohlendorf, D.H. | | Deposit date: | 2005-01-18 | | Release date: | 2005-08-16 | | Last modified: | 2021-10-20 | | Method: | X-RAY DIFFRACTION (2.41 Å) | | Cite: | Roles of the equatorial tyrosyl iron ligand of protocatechuate 3,4-dioxygenase in catalysis
Biochemistry, 44, 2005
|
|
3WNV
 
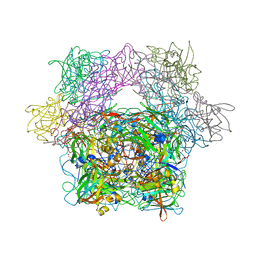
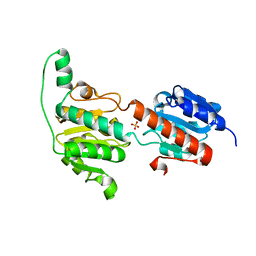 | | Crystal structure of a glyoxylate reductase from Paecilomyes thermophila | | Descriptor: | SULFATE ION, glyoxylate reductase | | Authors: | Duan, X, Hu, S, Zhou, P, Zhou, Y, Jiang, Z. | | Deposit date: | 2013-12-17 | | Release date: | 2014-12-03 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Characterization and crystal structure of a first fungal glyoxylate reductase from Paecilomyes thermophila
Enzyme.Microb.Technol., 60, 2014
|
|
1YL1
 
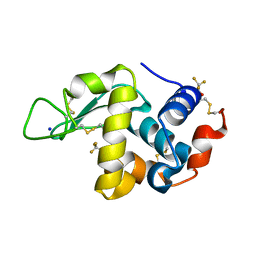 | | Effect of alcohols on protein hydration | | Descriptor: | Lysozyme C, SODIUM ION, TRIFLUOROETHANOL | | Authors: | Deshpande, A.A, Nimsadkar, S, Mande, S.C. | | Deposit date: | 2005-01-18 | | Release date: | 2005-07-05 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Effect of alcohols on protein hydration: crystallographic analysis of hen egg-white lysozyme in the presence of alcohols.
Acta Crystallogr.,Sect.D, 61, 2005
|
|
1PHG
 
 | |
4IIG
 
 | | Crystal structure of beta-glucosidase 1 from Aspergillus aculeatus in complex with D-glucose | | Descriptor: | (4R)-2-METHYLPENTANE-2,4-DIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Suzuki, K, Sumitani, J, Kawaguchi, T, Fushinobu, S. | | Deposit date: | 2012-12-20 | | Release date: | 2013-04-10 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal structures of glycoside hydrolase family 3 beta-glucosidase 1 from Aspergillus aculeatus
Biochem.J., 452, 2013
|
|
1YL6
 
 | | crystal structure of Mycobacterium tuberculosis dihydrodipicolinate reductase (Rv2773c) (crystal form B) | | Descriptor: | Dihydrodipicolinate reductase, MAGNESIUM ION | | Authors: | Janowski, R, Kefala, G, Weiss, M.S, TB Structural Genomics Consortium (TBSGC) | | Deposit date: | 2005-01-19 | | Release date: | 2006-01-17 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | The structure of dihydrodipicolinate reductase (DapB) from Mycobacterium tuberculosis in three crystal forms.
Acta Crystallogr.,Sect.D, 66, 2010
|
|
3WQ5
 
 | |
1YL5
 
 | | Crystal structure of Mycobacterium tuberculosis dihydrodipicolinate reductase (RV2773C) (crystal form A) | | Descriptor: | Dihydrodipicolinate reductase, MAGNESIUM ION | | Authors: | Janowski, R, Kefala, G, Weiss, M.S, TB Structural Genomics Consortium (TBSGC) | | Deposit date: | 2005-01-19 | | Release date: | 2006-01-17 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | The structure of dihydrodipicolinate reductase (DapB) from Mycobacterium tuberculosis in three crystal forms.
Acta Crystallogr.,Sect.D, 66, 2010
|
|
1YJK
 
 | | Reduced Peptidylglycine Alpha-Hydroxylating Monooxygenase (PHM) in a New Crystal Form | | Descriptor: | COPPER (II) ION, GLYCEROL, Peptidyl-glycine alpha-amidating monooxygenase | | Authors: | Siebert, X, Eipper, B.A, Mains, R.E, Prigge, S.T, Blackburn, N.J, Amzel, L.M. | | Deposit date: | 2005-01-14 | | Release date: | 2005-11-15 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The Catalytic Copper of Peptidylglycine alpha-Hydroxylating Monooxygenase also Plays a Critical Structural Role.
Biophys.J., 89, 2005
|
|
3WRE
 
 | | The crystal structure of native HypBA1 from Bifidobacterium longum JCM 1217 | | Descriptor: | Non-reducing end beta-L-arabinofuranosidase, ZINC ION | | Authors: | Huang, C.H, Zhu, Z, Cheng, Y.S, Chan, H.C, Ko, T.P, Chen, C.C, Wang, I, Ho, M.R, Hsu, S.T, Zeng, Y.F, Huang, Y.N, Liu, J.R, Guo, R.T. | | Deposit date: | 2014-02-25 | | Release date: | 2014-09-03 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.78 Å) | | Cite: | Structure and Catalytic Mechanism of a Glycoside Hydrolase Family-127 beta-L-Arabinofuranosidase (HypBA1)
J BIOPROCESS BIOTECH, 4, 2014
|
|
4C76
 
 | |
3WRJ
 
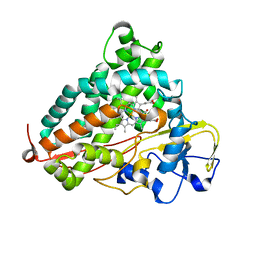 | | Crystal structure of P450cam | | Descriptor: | CAMPHOR, Camphor 5-monooxygenase, POTASSIUM ION, ... | | Authors: | Kishimoto, A, Takagi, K, Amano, A, Sakurai, K, Mizushima, T, Shimada, H. | | Deposit date: | 2014-02-25 | | Release date: | 2015-03-18 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Structure of P450cam intermedite
to be published
|
|
1YJL
 
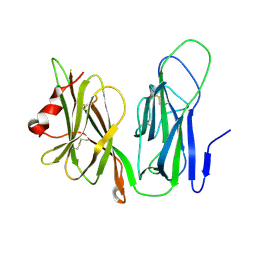 | | Reduced Peptidylglycine alpha-Hydroxylating Monooxygenase in a new crystal form | | Descriptor: | Peptidyl-glycine alpha-amidating monooxygenase | | Authors: | Siebert, X, Eipper, B.A, Mains, R.E, Prigge, S.T, Blackburn, N.J, Amzel, L.M. | | Deposit date: | 2005-01-14 | | Release date: | 2005-11-15 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | The catalytic copper of Peptidylglycine alpha-Hydroxylating Monooxygenase also plays a critical structural role.
Biophys.J., 89, 2005
|
|
4CEU
 
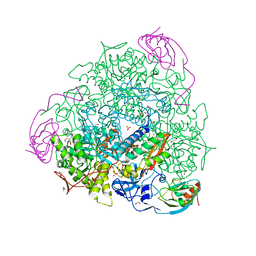 | | 1.58 A resolution native Sporosarcina pasteurii urease | | Descriptor: | 1,2-ETHANEDIOL, HYDROXIDE ION, NICKEL (II) ION, ... | | Authors: | Benini, S, Cianci, M, Ciurli, S. | | Deposit date: | 2013-11-12 | | Release date: | 2014-08-27 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.58 Å) | | Cite: | Fluoride Inhibition of Sporosarcina Pasteurii Urease: Structure and Thermodynamics.
J.Biol.Inorg.Chem., 19, 2014
|
|
4C7W
 
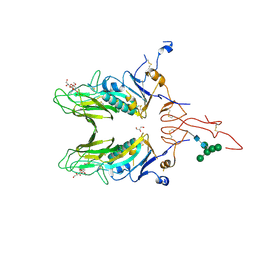 | |
1PBB
 
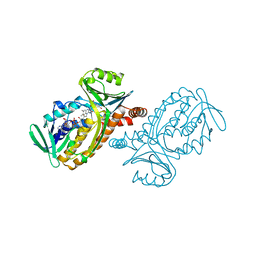 | | CRYSTAL STRUCTURES OF WILD-TYPE P-HYDROXYBENZOATE HYDROXYLASE COMPLEXED WITH 4-AMINOBENZOATE, 2,4-DIHYDROXYBENZOATE AND 2-HYDROXY-4-AMINOBENZOATE AND OF THE TRY222ALA MUTANT, COMPLEXED WITH 2-HYDROXY-4-AMINOBENZOATE. EVIDENCE FOR A PROTON CHANNEL AND A NEW BINDING MODE OF THE FLAVIN RING | | Descriptor: | 2,4-DIHYDROXYBENZOIC ACID, FLAVIN-ADENINE DINUCLEOTIDE, P-HYDROXYBENZOATE HYDROXYLASE | | Authors: | Schreuder, H.A, Mattevi, A, Hol, W.G.J. | | Deposit date: | 1994-07-06 | | Release date: | 1994-09-30 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal structures of wild-type p-hydroxybenzoate hydroxylase complexed with 4-aminobenzoate,2,4-dihydroxybenzoate, and 2-hydroxy-4-aminobenzoate and of the Tyr222Ala mutant complexed with 2-hydroxy-4-aminobenzoate. Evidence for a proton channel and a new binding mode of the flavin ring
Biochemistry, 33, 1994
|
|
2Y0P
 
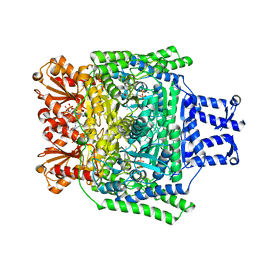 | | Crystal structure of the SucA domain of Mycobacterium smegmatis alpha- ketoglutarate decarboxylase in complex with the enamine-ThDP intermediate and acetyl-CoA | | Descriptor: | (4E)-4-{3-[(4-amino-2-methylpyrimidin-5-yl)methyl]-5-(2-{[(S)-hydroxy(phosphonooxy)phosphoryl]oxy}ethyl)-4-methyl-1,3-thiazol-2(3H)-ylidene}-4-hydroxybutanoic acid, 2-OXOGLUTARATE DECARBOXYLASE, ACETYL COENZYME *A, ... | | Authors: | Wagner, T, Bellinzoni, M, Wehenkel, A.M, O'Hare, H.M, Alzari, P.M. | | Deposit date: | 2010-12-07 | | Release date: | 2011-06-15 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Functional Plasticity and Allosteric Regulation of Alpha-Ketoglutarate Decarboxylase in Central Mycobacterial Metabolism.
Chem.Biol., 18, 2011
|
|
4BHX
 
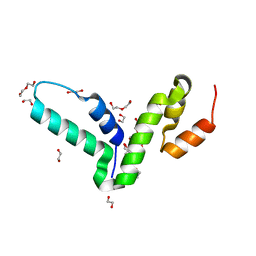 | | Crystal structure of the SCAN domain from human paternally expressed gene 3 protein | | Descriptor: | 1,2-ETHANEDIOL, DI(HYDROXYETHYL)ETHER, PATERNALLY-EXPRESSED GENE 3 PROTEIN, ... | | Authors: | Rimsa, V, Eadsforth, T.C, Hunter, W.N. | | Deposit date: | 2013-04-09 | | Release date: | 2013-04-17 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Structure of the Scan Domain of Human Paternally Expressed Gene 3 Protein.
Plos One, 8, 2013
|
|
3WTC
 
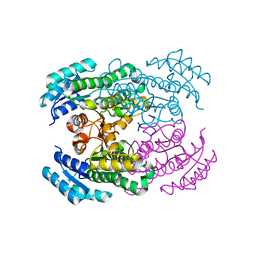 | | Crystal structure of Gox2036 | | Descriptor: | Putative oxidoreductase | | Authors: | Yuan, Y.A, Lin, J.P. | | Deposit date: | 2014-04-09 | | Release date: | 2015-04-15 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Crystal structure of Gox0525
To be Published
|
|
1PIG
 
 | | PIG PANCREATIC ALPHA-AMYLASE COMPLEXED WITH THE OLIGOSACCHARIDE V-1532 | | Descriptor: | 4-amino-4,6-dideoxy-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose, 4-amino-4,6-dideoxy-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose-(1-4)-beta-D-glucopyranose, 5-HYDROXYMETHYL-CHONDURITOL, ... | | Authors: | Machius, M, Vertesy, L, Huber, R, Wiegand, G. | | Deposit date: | 1996-06-15 | | Release date: | 1996-12-07 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Carbohydrate and protein-based inhibitors of porcine pancreatic alpha-amylase: structure analysis and comparison of their binding characteristics.
J.Mol.Biol., 260, 1996
|
|
