1O9E
 
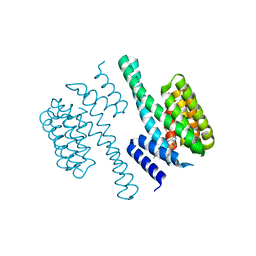 | | Structural view of a fungal toxin acting on a 14-3-3 regulatory complex | | Descriptor: | 14-3-3-LIKE PROTEIN C, CITRATE ANION, FUSICOCCIN | | Authors: | Wurtele, M, Jelich-Ottmann, C, Wittinghofer, A, Oecking, C. | | Deposit date: | 2002-12-12 | | Release date: | 2003-03-06 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structural View of a Fungal Toxin Acting on a 14-3-3 Regulatory Complex
Embo J., 22, 2003
|
|
1O9F
 
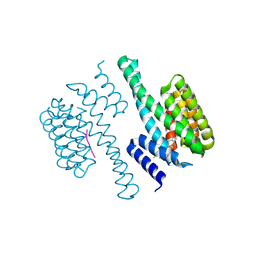 | | Structural view of a fungal toxin acting on a 14-3-3 regulatory complex | | Descriptor: | 14-3-3-LIKE PROTEIN C, FUSICOCCIN, PLASMA MEMBRANE H+ ATPASE | | Authors: | Wurtele, M, Jelich-Ottmann, C, Wittinghofer, A, Oecking, C. | | Deposit date: | 2002-12-12 | | Release date: | 2003-03-06 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structural View of a Fungal Toxin Acting on a 14-3-3 Regulatory Complex
Embo J., 22, 2003
|
|
1O9G
 
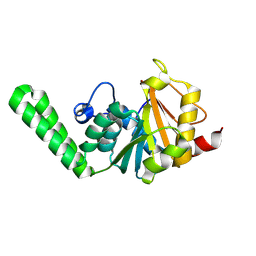 | |
1O9H
 
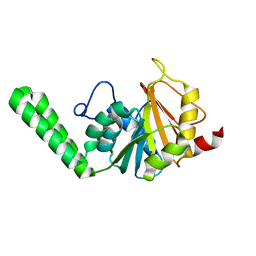 | |
1O9I
 
 | | Crystal structure of the Y42F mutant of manganese catalase from Lactobacillus plantarum at 1.33A resolution | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, CALCIUM ION, MANGANESE (III) ION, ... | | Authors: | Barynin, V.V, Whittaker, M.M, Whittaker, J.W. | | Deposit date: | 2002-12-13 | | Release date: | 2003-12-18 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.33 Å) | | Cite: | Outer Sphere Mutagenesis of Lactobacillus Plantarum Manganese Catalase Disrupts the Cluster Core. Mechanistic Implications.
Eur.J.Biochem., 270, 2003
|
|
1O9J
 
 | | The X-ray crystal structure of eta-crystallin | | Descriptor: | (2R,3S)-1,4-DIMERCAPTOBUTANE-2,3-DIOL, 2,3-DIHYDROXY-1,4-DITHIOBUTANE, ALDEHYDE DEHYDROGENASE, ... | | Authors: | Purkiss, A.G, Van Montfort, R, Wistow, G, Slingsby, C. | | Deposit date: | 2002-12-15 | | Release date: | 2003-04-17 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal Structure of Eta-Crystallin: Adaptation of a Class 1 Aldehyde Dehydrogenase for a New Role in the Eye Lens
Biochemistry, 42, 2003
|
|
1O9K
 
 | | Crystal structure of the retinoblastoma tumour suppressor protein bound to E2F peptide | | Descriptor: | RETINOBLASTOMA-ASSOCIATED PROTEIN, TRANSCRIPTION FACTOR E2F1 | | Authors: | Xiao, B, Spencer, J, Clements, A, Ali-Khan, N, Mittnacht, S, Broceno, C, Burghammer, M, Perrakis, A, Marmorstein, R, Gamblin, S.J. | | Deposit date: | 2002-12-16 | | Release date: | 2003-03-06 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Crystal Structure of the Retinoblastoma Tumor Suppressor Protein Bound to E2F and the Molecular Basis of its Regulation
Proc.Natl.Acad.Sci.USA, 100, 2003
|
|
1O9L
 
 | |
1O9M
 
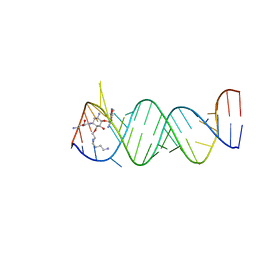 | | The Complex of a novel antibiotic with the Aminoacyl Site of the Bacterial Ribosome Revealed by X-Ray Crystallography. | | Descriptor: | 1-(AMINOETHYL)AMINO-4-AMINOBUTANE, 2,6-diamino-2,6-dideoxy-alpha-D-glucopyranose, 4-AMINO-2-HYDROXYBUTANOIC ACID, ... | | Authors: | Russell, R, Murray, J.B. | | Deposit date: | 2002-12-17 | | Release date: | 2003-03-27 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | The Complex of a Designer Antibiotic with a Model Aminoacyl Site of the 30S Ribosomal Subunit Revealed by X-Ray Crystallography
J.Am.Chem.Soc., 125, 2003
|
|
1O9N
 
 | |
1O9O
 | |
1O9P
 
 | |
1O9Q
 
 | |
1O9R
 
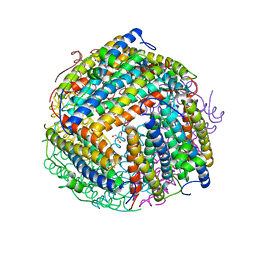 | | The X-ray crystal structure of Agrobacterium tumefaciens Dps, a member of the family that protect DNA without binding | | Descriptor: | 1,2-ETHANEDIOL, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, AGROBACTERIUM TUMEFACIENS DPS, ... | | Authors: | Ilari, A, Ceci, P, Chiancone, E. | | Deposit date: | 2002-12-18 | | Release date: | 2003-05-29 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | The Dps Protein of Agrobacterium Tumefaciens Does not Bind to DNA But Protects It Toward Oxidative Cleavage: X-Ray Crystal Structure, Iron Binding, and Hydroxyl-Radical Scavenging Properties
J.Biol.Chem., 278, 2003
|
|
1O9S
 
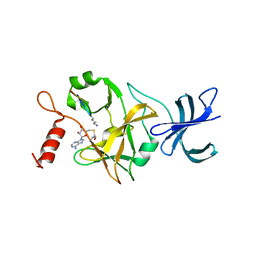 | | Crystal structure of a ternary complex of the human histone methyltransferase SET7/9 | | Descriptor: | GENE FRAGMENT FOR HISTONE H3, HISTONE-LYSINE N-METHYLTRANSFERASE, H3 LYSINE-4 SPECIFIC, ... | | Authors: | Xiao, B, Jing, C, Wilson, J.R, Walker, P.A, Vasisht, N, Kelly, G, Howell, S, Taylor, I.A, Blackburn, G.M, Gamblin, S.J. | | Deposit date: | 2002-12-18 | | Release date: | 2003-02-06 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Structure and Catalytic Mechanism of the Human Histone Methyltransferase Set7/9
Nature, 421, 2003
|
|
1O9T
 
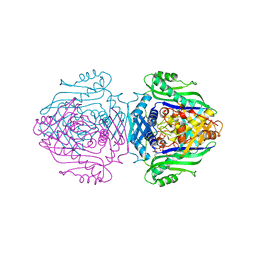 | | Methionine adenosyltransferase complexed with both substrates ATP and methionine | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, MAGNESIUM ION, METHIONINE, ... | | Authors: | Gonzalez, B, Pajares, M.A, Hermoso, J.A, Sanz-Aparicio, J. | | Deposit date: | 2002-12-18 | | Release date: | 2003-08-07 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Crystal Structures of Methionine Adenosyltransferase Complexed with Substrates and Products Reveal the Methionine-ATP Recognition and Give Insights Into the Catalytic Mechanism
J.Mol.Biol., 331, 2003
|
|
1O9U
 
 | |
1O9V
 
 | | F17-aG lectin domain from Escherichia coli in complex with a selenium carbohydrate derivative | | Descriptor: | F17-AG LECTIN DOMAIN, methyl 2-acetamido-2-deoxy-1-seleno-beta-D-glucopyranoside | | Authors: | Buts, L, De Genst, E, Loris, R, Oscarson, S, Lahmann, M, Messens, J, Brosens, E, Wyns, L, Bouckaert, J, De Greve, H. | | Deposit date: | 2002-12-20 | | Release date: | 2003-05-29 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | The Fimbrial Adhesin F17-G of Enterotoxigenic Escherichia Coli Has an Immunoglobulin-Like Lectin Domain that Binds N-Acetylglucosamine
Mol.Microbiol., 49, 2003
|
|
1O9W
 
 | | F17-aG lectin domain from Escherichia coli in complex with N-acetyl-glucosamine | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, F17A-G FIMBRIAL ADHESIN | | Authors: | Buts, L, De Genst, E, Loris, R, Oscarson, S, Lahmann, M, Messens, J, Brosens, E, Wyns, L, Bouckaert, J, De Greve, H. | | Deposit date: | 2002-12-20 | | Release date: | 2003-05-29 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | The Fimbrial Adhesin F17-G of Enterotoxigenic Escherichia Coli Has an Immunoglobulin-Like Lectin Domain that Binds N-Acetylglucosamine
Mol.Microbiol., 49, 2003
|
|
1O9X
 
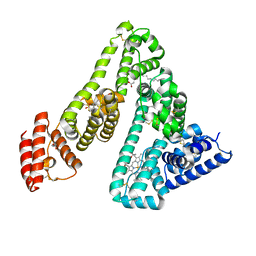 | | HUMAN SERUM ALBUMIN COMPLEXED WITH TETRADECANOIC ACID (MYRISTIC ACID) AND HEMIN | | Descriptor: | MYRISTIC ACID, PROTOPORPHYRIN IX CONTAINING FE, SERUM ALBUMIN | | Authors: | Zunszain, P.A, Ghuman, J, Komatsu, T, Tsuchida, E, Curry, S. | | Deposit date: | 2002-12-20 | | Release date: | 2003-03-13 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Crystal Structure Analysis of Human Serum Albumin Complexed with Hemin and Fatty Acid
Bmc Struct.Biol., 3, 2003
|
|
1O9Y
 
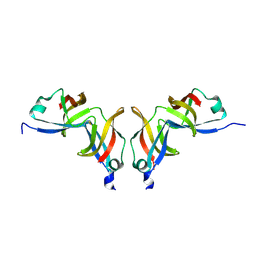 | |
1O9Z
 
 | | F17-aG lectin domain from Escherichia coli (ligand free) | | Descriptor: | F17-AG LECTIN DOMAIN | | Authors: | Buts, L, De Genst, E, Loris, R, Oscarson, S, Lahmann, M, Messens, J, Brosens, E, Wyns, L, Bouckaert, J, De Greve, H. | | Deposit date: | 2002-12-23 | | Release date: | 2003-05-29 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | The Fimbrial Adhesin F17-G of Enterotoxigenic Escherichia Coli Has an Immunoglobulin-Like Lectin Domain that Binds N-Acetylglucosamine
Mol.Microbiol., 49, 2003
|
|
1OA0
 
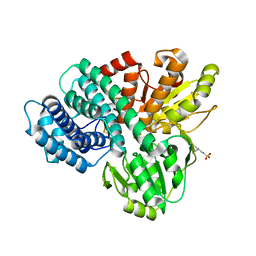 | | REDUCED HYBRID CLUSTER PROTEIN FROM DESULFOVIBRIO DESULFURICANS X-RAY STRUCTURE AT 1.25A RESOLUTION | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, FE4-S3 CLUSTER, IRON/SULFUR CLUSTER, ... | | Authors: | Macedo, S, Aragao, D, Mitchell, E.P, Romao, C.V, Liu, M.Y, Frazao, C, Saraiva, L.M, Xavier, A.V, Legall, J, Van Dongen, W.M.A.M, Hagen, W.R, Teixeira, M, Carrondo, M.A, Lindley, P.F. | | Deposit date: | 2002-12-23 | | Release date: | 2003-04-08 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.25 Å) | | Cite: | Reduced hybrid cluster proteins (HCP) from Desulfovibrio desulfuricans ATCC 27774 and Desulfovibrio vulgaris (Hildenborough): X-ray structures at high resolution using synchrotron radiation.
J. Biol. Inorg. Chem., 8, 2003
|
|
1OA1
 
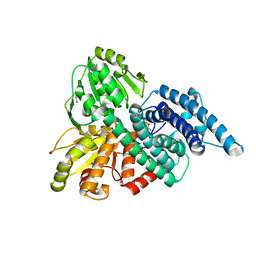 | | REDUCED HYBRID CLUSTER PROTEIN (HCP) FROM DESULFOVIBRIO VULGARIS HILDENBOROUGH STRUCTURE AT 1.55A RESOLUTION USING SYNCHROTRON RADIATION. | | Descriptor: | FE4-S3 CLUSTER, GLYCEROL, HYDROXYLAMINE REDUCTASE, ... | | Authors: | Aragao, D, Macedo, S, Mitchell, E.P, Romao, C.V, Liu, M.Y, Frazao, C, Saraiva, L.M, Xavier, A.V, Legall, J, Van Dongen, W.M.A.M, Hagen, W.R, Teixeira, M, Carrondo, M.A, Lindley, P.F. | | Deposit date: | 2002-12-23 | | Release date: | 2003-04-08 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Reduced Hybrid Cluster Proteins (Hcp) from Desulfovibrio Desulfuricans Atcc 27774 and Desulfovibrio Vulgaris (Hildenborough): X-Ray Structures at High Resolution Using Synchrotron Radiation
J.Biol.Inorg.Chem., 8, 2003
|
|
1OA2
 
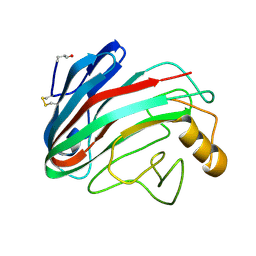 | | Comparison of Family 12 Glycoside Hydrolases and Recruited Substitutions Important for Thermal Stability | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, ENDO-BETA-1,4-GLUCANASE | | Authors: | Sandgren, M, Gualfetti, P.J, Shaw, A, Gross, L.S, Saldajeno, M, Day, A.G, Jones, T.A, Mitchinson, C. | | Deposit date: | 2002-12-28 | | Release date: | 2003-03-27 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Comparison of Family 12 Glycoside Hydrolases and Recruited Substitutions Important for Thermal Stability
Protein Sci., 12, 2003
|
|
