3DIR
 
 | |
3DIM
 
 | |
3DJ2
 
 | |
3DIL
 
 | |
3DIS
 
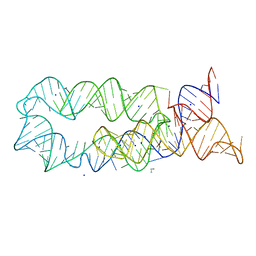 | |
3DJ0
 
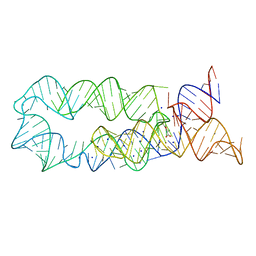 | |
3PDR
 
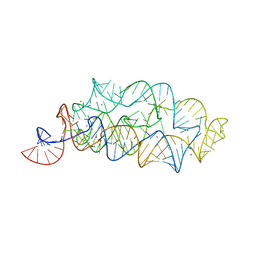 | | Crystal structure of manganese bound M-box RNA | | 分子名称: | M-box Riboswitch RNA, MANGANESE (II) ION, POTASSIUM ION | | 著者 | Ramesh, A, Winkler, W.C. | | 登録日 | 2010-10-23 | | 公開日 | 2011-02-23 | | 最終更新日 | 2024-02-21 | | 実験手法 | X-RAY DIFFRACTION (1.853 Å) | | 主引用文献 | Insights into Metalloregulation by M-box Riboswitch RNAs via Structural Analysis of Manganese-Bound Complexes.
J.Mol.Biol., 407, 2011
|
|
1Y26
 
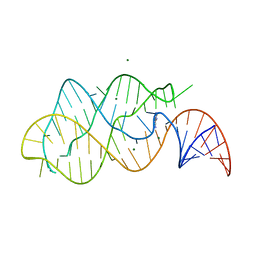 | | A-riboswitch-adenine complex | | 分子名称: | ADENINE, MAGNESIUM ION, Vibrio vulnificus A-riboswitch | | 著者 | Serganov, A, Yuan, Y.R, Patel, D.J. | | 登録日 | 2004-11-20 | | 公開日 | 2004-12-28 | | 最終更新日 | 2024-02-14 | | 実験手法 | X-RAY DIFFRACTION (2.1 Å) | | 主引用文献 | Structural Basis for Discriminative Regulation of Gene Expression by Adenine- and Guanine-Sensing mRNAs
Chem.Biol., 11, 2004
|
|
5C45
 
 | | Selective Small Molecule Inhibition of the FMN Riboswitch | | 分子名称: | (6M)-2-[(3S)-1-{[2-(methylamino)pyrimidin-5-yl]methyl}piperidin-3-yl]-6-(thiophen-2-yl)pyrimidin-4-ol, FMN Riboswitch, MAGNESIUM ION, ... | | 著者 | Fischmann, T.O. | | 登録日 | 2015-06-17 | | 公開日 | 2015-10-07 | | 最終更新日 | 2023-10-25 | | 実験手法 | X-RAY DIFFRACTION (2.93 Å) | | 主引用文献 | Selective small-molecule inhibition of an RNA structural element.
Nature, 526, 2015
|
|
3DS7
 
 | |
8F3C
 
 | | Cryo-EM consensus structure of Escherichia coli que-PEC (paused elongation complex) RNA Polymerase minus preQ1 ligand | | 分子名称: | DNA (38-MER), DNA (39-MER), DNA-directed RNA polymerase subunit alpha, ... | | 著者 | Porta, J.C, Chauvier, A, Deb, I, Ellinger, E, Frank, A.T, Meze, K, Ohi, M.D, Walter, N.G. | | 登録日 | 2022-11-09 | | 公開日 | 2023-06-21 | | 最終更新日 | 2024-06-19 | | 実験手法 | ELECTRON MICROSCOPY (3.4 Å) | | 主引用文献 | Structural basis for control of bacterial RNA polymerase pausing by a riboswitch and its ligand.
Nat.Struct.Mol.Biol., 30, 2023
|
|
8G00
 
 | | Cryo-EM structure of 3DVA component 0 of Escherichia coli que-PEC (paused elongation complex) RNA Polymerase minus preQ1 ligand | | 分子名称: | DNA (31-MER), DNA (39-mer), DNA-directed RNA polymerase subunit alpha, ... | | 著者 | Porta, J.C, Chauvier, A, Deb, I, Ellinger, E, Frank, A.T, Meze, K, Ohi, M.D, Walter, N.G. | | 登録日 | 2023-01-31 | | 公開日 | 2023-06-21 | | 最終更新日 | 2024-06-19 | | 実験手法 | ELECTRON MICROSCOPY (3.4 Å) | | 主引用文献 | Structural basis for control of bacterial RNA polymerase pausing by a riboswitch and its ligand.
Nat.Struct.Mol.Biol., 30, 2023
|
|
7KVV
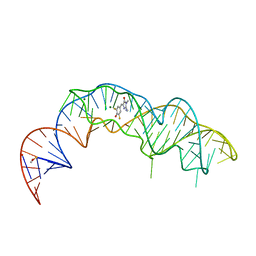 
 | |
7KVT
 
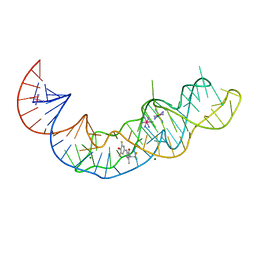 | | Crystal structure of Squash RNA aptamer in complex with DFHBI-1T with iridium (III) ions | | 分子名称: | (5Z)-5-(3,5-difluoro-4-hydroxybenzylidene)-2-methyl-3-(2,2,2-trifluoroethyl)-3,5-dihydro-4H-imidazol-4-one, IRIDIUM HEXAMMINE ION, MAGNESIUM ION, ... | | 著者 | Truong, L, Ferre-D'Amare, A.R. | | 登録日 | 2020-11-28 | | 公開日 | 2022-01-19 | | 最終更新日 | 2024-05-22 | | 実験手法 | X-RAY DIFFRACTION (2.73 Å) | | 主引用文献 | The fluorescent aptamer Squash extensively repurposes the adenine riboswitch fold.
Nat.Chem.Biol., 18, 2022
|
|
7KVU
 
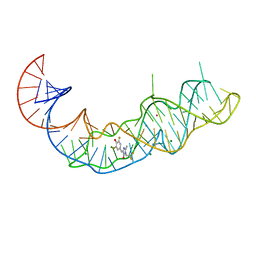 | | Crystal structure of Squash RNA aptamer in complex with DFHBI-1T | | 分子名称: | (5Z)-5-(3,5-difluoro-4-hydroxybenzylidene)-2-methyl-3-(2,2,2-trifluoroethyl)-3,5-dihydro-4H-imidazol-4-one, MAGNESIUM ION, POTASSIUM ION, ... | | 著者 | Truong, L, Ferre-D'Amare, A.R. | | 登録日 | 2020-11-28 | | 公開日 | 2022-01-19 | | 最終更新日 | 2023-10-18 | | 実験手法 | X-RAY DIFFRACTION (2.68 Å) | | 主引用文献 | The fluorescent aptamer Squash extensively repurposes the adenine riboswitch fold.
Nat.Chem.Biol., 18, 2022
|
|
4B5R
 
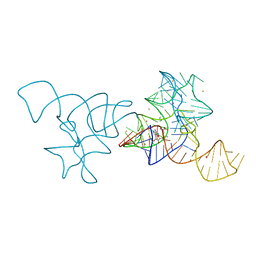 | |
5FK4
 
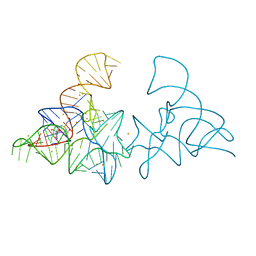 | |
5FK1
 
 | |
5FK5
 
 | |
5FK3
 
 | |
5FK2
 
 | |
5FKF
 
 | |
5FKD
 
 | |
5FKG
 
 | |
5FKH
 
 | |
