1D2J
 
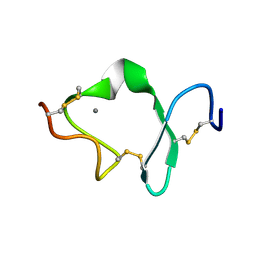 | | LDL RECEPTOR LIGAND-BINDING MODULE 6 | | Descriptor: | CALCIUM ION, LOW-DENSITY LIPOPROTEIN RECEPTOR | | Authors: | North, C.L, Blacklow, S.C. | | Deposit date: | 1999-09-23 | | Release date: | 2000-03-22 | | Last modified: | 2021-11-03 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the sixth LDL-A module of the LDL receptor.
Biochemistry, 39, 2000
|
|
1COZ
 
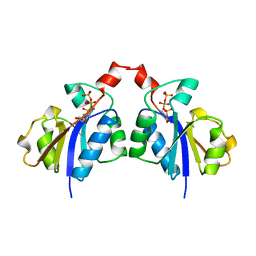 | | CTP:GLYCEROL-3-PHOSPHATE CYTIDYLYLTRANSFERASE FROM BACILLUS SUBTILIS | | Descriptor: | CYTIDINE-5'-TRIPHOSPHATE, PROTEIN (GLYCEROL-3-PHOSPHATE CYTIDYLYLTRANSFERASE) | | Authors: | Weber, C.H, Park, Y.S, Sanker, S, Kent, C, Ludwig, M.L. | | Deposit date: | 1999-05-29 | | Release date: | 1999-10-06 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | A prototypical cytidylyltransferase: CTP:glycerol-3-phosphate cytidylyltransferase from bacillus subtilis.
Structure Fold.Des., 7, 1999
|
|
1CHI
 
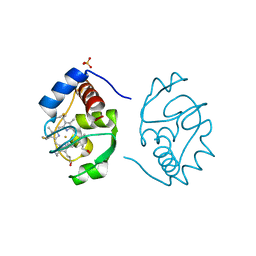 | |
7Q92
 
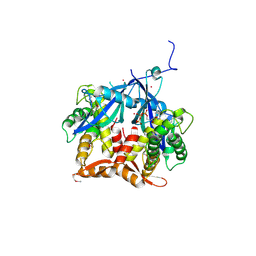 | | Crystal Structure of Agrobacterium tumefaciens NADQ, ATP complex. | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, NADQ transcription factor, POTASSIUM ION, ... | | Authors: | Cianci, M, Minazzato, G, Heroux, A, Raffaelli, N, Sorci, L, Gasparrini, M. | | Deposit date: | 2021-11-11 | | Release date: | 2022-11-09 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (2.18 Å) | | Cite: | Bacterial NadQ (COG4111) is a Nudix-like, ATP-responsive regulator of NAD biosynthesis.
J.Struct.Biol., 214, 2022
|
|
7Q93
 
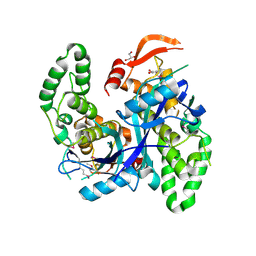 | | Crystal Structure of Agrobacterium tumefaciens NADQ, NAD complex. | | Descriptor: | GLYCEROL, NADQ transcription factor, NICOTINAMIDE-ADENINE-DINUCLEOTIDE, ... | | Authors: | Cianci, M, Minazzato, G, Heroux, A, Raffaelli, N, Sorci, L, Gasparrini, M. | | Deposit date: | 2021-11-11 | | Release date: | 2022-11-23 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (2.19 Å) | | Cite: | Bacterial NadQ (COG4111) is a Nudix-like, ATP-responsive regulator of NAD biosynthesis.
J.Struct.Biol., 214, 2022
|
|
8AYL
 
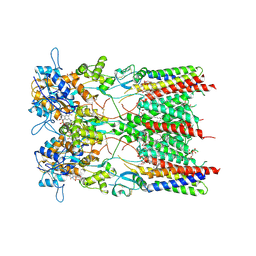 | | Resting state GluA1/A2 AMPA receptor in complex with TARP gamma 8 and ligand JNJ-61432059 | | Descriptor: | (2R)-2,3-dihydroxypropyl (9Z)-octadec-9-enoate, (2S)-3-(hexadecanoyloxy)-2-[(9Z)-octadec-9-enoyloxy]propyl 2-(trimethylammonio)ethyl phosphate, 5-[2-(4-fluorophenyl)-7-(4-oxidanylpiperidin-1-yl)pyrazolo[1,5-c]pyrimidin-3-yl]-1,3-dihydroindol-2-one, ... | | Authors: | Zhang, D, Lape, R, Shaikh, S, Kohegyi, B, Watson, J.F, Cais, O, Nakagawa, T, Greger, I. | | Deposit date: | 2022-09-02 | | Release date: | 2023-04-19 | | Last modified: | 2023-05-24 | | Method: | ELECTRON MICROSCOPY (3.2 Å) | | Cite: | Modulatory mechanisms of TARP gamma 8-selective AMPA receptor therapeutics.
Nat Commun, 14, 2023
|
|
8ARV
 
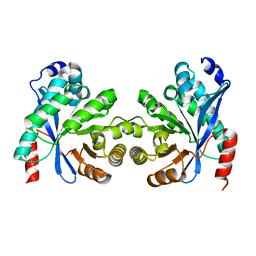 | |
8QLV
 
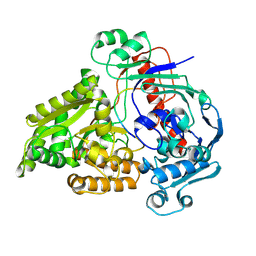 | | Crystal structure of the pneumococcal Substrate-binding protein AliB in complex with Peptide 4 | | Descriptor: | Oligopeptide-binding protein AliB, VAL-MET-VAL-LYS-GLY-PRO-GLY-PRO-GLY-ARG | | Authors: | Alcorlo, M, Abdullah, M.R, Hammerschmidt, S, Hermoso, J. | | Deposit date: | 2023-09-20 | | Release date: | 2024-05-22 | | Last modified: | 2024-07-03 | | Method: | X-RAY DIFFRACTION (1.49 Å) | | Cite: | Molecular and structural basis of oligopeptide recognition by the Ami transporter system in pneumococci.
Plos Pathog., 20, 2024
|
|
6RIA
 
 | |
8ARC
 
 | |
8ARB
 
 | | Heterologous Complex of shortened Aeromonas hydrophila Type III secretion substrate AscX with Yersinia enterocolitica chaperone YscY | | Descriptor: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, AscX, Chaperone protein YscY, ... | | Authors: | Gilzer, D, Flottmann, F, Niemann, H.H. | | Deposit date: | 2022-08-16 | | Release date: | 2023-04-26 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.63 Å) | | Cite: | The type III secretion chaperone SctY may shield the hydrophobic export gate-binding C-terminus of its substrate SctX.
Acta Crystallogr D Struct Biol, 79, 2023
|
|
8ARA
 
 | | Heterologous Complex of Aeromonas hydrophila Type III secretion substrate AscX with Yersinia enterocolitica chaperone YscY | | Descriptor: | ACETATE ION, AscX, CHLORIDE ION, ... | | Authors: | Gilzer, D, Kowal, J.L, Niemann, H.H. | | Deposit date: | 2022-08-16 | | Release date: | 2023-04-26 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | The type III secretion chaperone SctY may shield the hydrophobic export gate-binding C-terminus of its substrate SctX.
Acta Crystallogr D Struct Biol, 79, 2023
|
|
1CSV
 
 | |
1CSU
 
 | |
6RWH
 
 | | Fragment AZ-007 binding at a primary and secondary binding site of the the p53pT387/14-3-3 sigma complex | | Descriptor: | 14-3-3 protein sigma, 5-(1~{H}-imidazol-5-yl)-4-phenyl-thiophene-2-carboximidamide, CALCIUM ION, ... | | Authors: | Leysen, S, Guillory, X, Wolter, M, Genet, S, Somsen, B, Patel, J, Castaldi, P, Ottmann, C. | | Deposit date: | 2019-06-05 | | Release date: | 2020-06-17 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.68 Å) | | Cite: | Fragment-based Differential Targeting of PPI Stabilizer Interfaces.
J.Med.Chem., 63, 2020
|
|
1CSX
 
 | |
1CSW
 
 | |
8Q6P
 
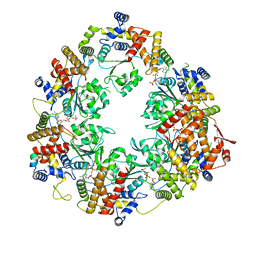 | | X. laevis CMG dimer bound to dimeric DONSON - MCM ATPase | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, DNA replication licensing factor mcm2, DNA replication licensing factor mcm4-B, ... | | Authors: | Butryn, A, Cvetkovic, M.A, Costa, A. | | Deposit date: | 2023-08-14 | | Release date: | 2023-10-18 | | Last modified: | 2023-11-29 | | Method: | ELECTRON MICROSCOPY (3.53 Å) | | Cite: | The structural mechanism of dimeric DONSON in replicative helicase activation.
Mol.Cell, 83, 2023
|
|
1CMF
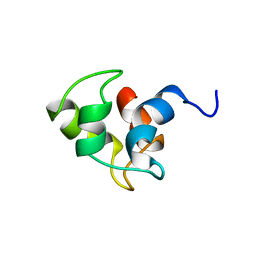 
 | | NMR SOLUTION STRUCTURE OF APO CALMODULIN CARBOXY-TERMINAL DOMAIN | | Descriptor: | CALMODULIN (VERTEBRATE) | | Authors: | Finn, B.E, Evenas, J, Drakenberg, T, Waltho, J.P, Thulin, E, Forsen, S. | | Deposit date: | 1995-07-19 | | Release date: | 1995-12-07 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Calcium-induced structural changes and domain autonomy in calmodulin.
Nat.Struct.Biol., 2, 1995
|
|
1D42
 
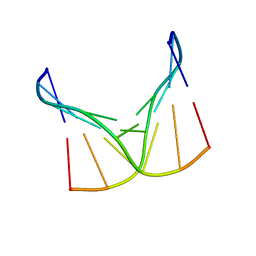 | |
8Q6O
 
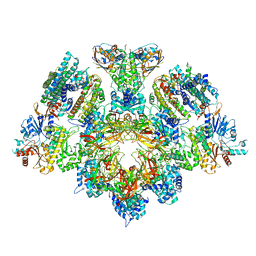 | | X. laevis CMG dimer bound to dimeric DONSON - without ATPase | | Descriptor: | Cell division control protein 45 homolog, DNA replication complex GINS protein PSF1, DNA replication complex GINS protein PSF2, ... | | Authors: | Butryn, A, Cvetkovic, M.A, Costa, A. | | Deposit date: | 2023-08-14 | | Release date: | 2023-10-18 | | Last modified: | 2023-11-29 | | Method: | ELECTRON MICROSCOPY (3.14 Å) | | Cite: | The structural mechanism of dimeric DONSON in replicative helicase activation.
Mol.Cell, 83, 2023
|
|
1CMG
 
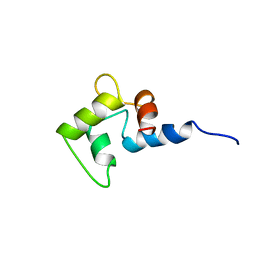 | | NMR SOLUTION STRUCTURE OF CALCIUM-LOADED CALMODULIN CARBOXY-TERMINAL DOMAIN | | Descriptor: | CALMODULIN (VERTEBRATE) | | Authors: | Evenas, J, Finn, B.E, Drakenberg, T, Waltho, J.P, Thulin, E, Forsen, S. | | Deposit date: | 1995-07-19 | | Release date: | 1995-12-07 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Calcium-induced structural changes and domain autonomy in calmodulin.
Nat.Struct.Biol., 2, 1995
|
|
8AZY
 
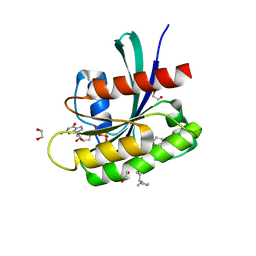 | | KRAS-G12D in complex with BI-2865 | | Descriptor: | (4S)-2-azanyl-4-methyl-4-[3-[4-[(1S)-1-[(2S)-1-methylpyrrolidin-1-ium-2-yl]ethoxy]pyrimidin-2-yl]-1,2,4-oxadiazol-5-yl]-6,7-dihydro-5H-1-benzothiophene-3-carbonitrile, 1,2-ETHANEDIOL, GTPase KRas, ... | | Authors: | Boettcher, J, Herdeis, L. | | Deposit date: | 2022-09-06 | | Release date: | 2023-06-07 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.09 Å) | | Cite: | Pan-KRAS inhibitor disables oncogenic signalling and tumour growth.
Nature, 619, 2023
|
|
5WB5
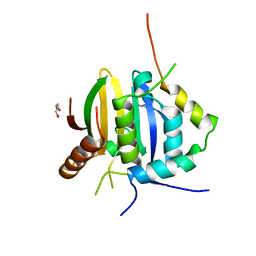 
 | | Leishmania IF4E-1 bound to Leishmania 4E-IP1 | | Descriptor: | DI(HYDROXYETHYL)ETHER, Putative eukaryotic translation initiation factor eIF-4E, Uncharacterized protein | | Authors: | Leger-Abraham, M, Meleppattu, S, Arthanari, H, Zinoviev, A, Boeszoermenyi, A, Wagner, G, Shapira, M. | | Deposit date: | 2017-06-27 | | Release date: | 2018-03-14 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structural basis for LeishIF4E-1 modulation by an interacting protein in the human parasite Leishmania major.
Nucleic Acids Res., 46, 2018
|
|
8AZR
 
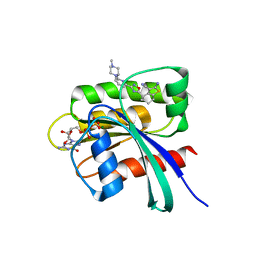 | | KRAS in complex with precursor 1 | | Descriptor: | (4~{S})-2-azanyl-4-[3-[6-[(2~{S})-2,4-dimethylpiperazin-1-yl]pyridin-2-yl]-1,2,4-oxadiazol-5-yl]-4-methyl-6,7-dihydro-5~{H}-1-benzothiophene-3-carbonitrile, 1,2-ETHANEDIOL, GTPase KRas, ... | | Authors: | Boettcher, J, Herdeis, L. | | Deposit date: | 2022-09-06 | | Release date: | 2023-06-07 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Pan-KRAS inhibitor disables oncogenic signalling and tumour growth.
Nature, 619, 2023
|
|
