2P9U
 
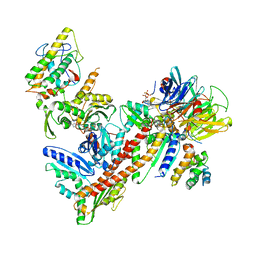 | |
1U2V
 
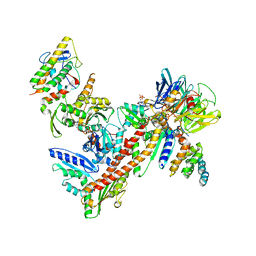 | | Crystal structure of Arp2/3 complex with bound ADP and calcium | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, Actin-Related Protein 2, Actin-Related Protein 3, ... | | Authors: | Nolen, B.J, Littlefield, R.S, Pollard, T.D. | | Deposit date: | 2004-07-20 | | Release date: | 2004-11-09 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Crystal structures of actin-related protein 2/3 complex with bound ATP or ADP
Proc.Natl.Acad.Sci.Usa, 101, 2004
|
|
1TYQ
 
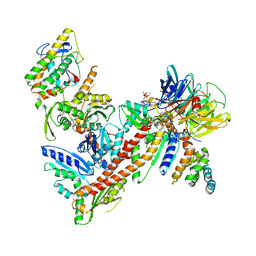 | | Crystal structure of Arp2/3 complex with bound ATP and calcium | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, Actin-related Protein 2, Actin-related protein 3, ... | | Authors: | Nolen, B.J, Littlefield, R.S, Pollard, T.D. | | Deposit date: | 2004-07-08 | | Release date: | 2004-11-09 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Crystal structures of actin-related protein 2/3 complex with bound ATP or ADP
Proc.Natl.Acad.Sci.Usa, 101, 2004
|
|
4I6M
 
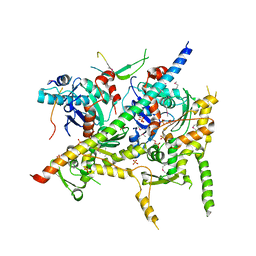 | | Structure of Arp7-Arp9-Snf2(HSA)-RTT102 subcomplex of SWI/SNF chromatin remodeler. | | Descriptor: | Actin-like protein ARP9, Actin-related protein 7, PHOSPHATE ION, ... | | Authors: | Schubert, H.L, Cairns, B.R, Hill, C.P. | | Deposit date: | 2012-11-29 | | Release date: | 2013-02-13 | | Last modified: | 2017-08-16 | | Method: | X-RAY DIFFRACTION (2.801 Å) | | Cite: | Structure of an actin-related subcomplex of the SWI/SNF chromatin remodeler.
Proc.Natl.Acad.Sci.USA, 110, 2013
|
|
5AEY
 
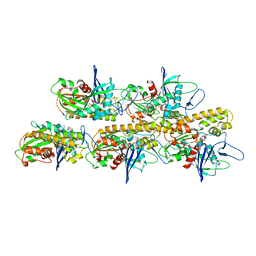 | | actin-like ParM protein bound to AMPPNP | | Descriptor: | PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER, PLASMID SEGREGATION PROTEIN PARM | | Authors: | Bharat, T.A.M, Murshudov, G.N, Sachse, C, Lowe, J. | | Deposit date: | 2015-01-12 | | Release date: | 2015-04-22 | | Last modified: | 2024-05-08 | | Method: | ELECTRON MICROSCOPY (4.3 Å) | | Cite: | Structures of Actin-Like Parm Filaments Show Architecture of Plasmid-Segregating Spindles.
Nature, 523, 2015
|
|
7M0G
 
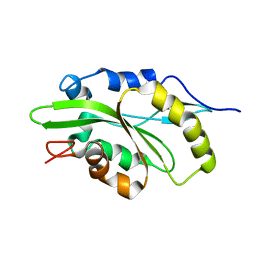 | |
6SL3
 
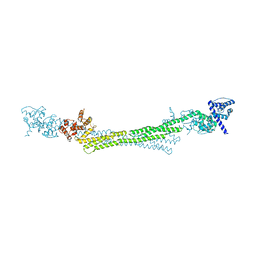 | |
5WJE
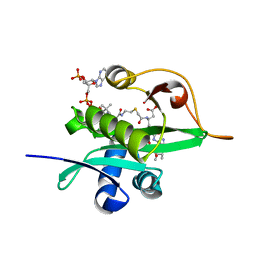 
 | | Crystal structure of Naa80 bound to a bisubstrate analogue | | Descriptor: | Actin N-terminus peptide, CARBOXYMETHYL COENZYME *A, CG8481, ... | | Authors: | Goris, M, Magin, R.S, Marmorstein, R, Arnesen, T. | | Deposit date: | 2017-07-21 | | Release date: | 2018-03-28 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.765 Å) | | Cite: | Structural determinants and cellular environment define processed actin as the sole substrate of the N-terminal acetyltransferase NAA80.
Proc. Natl. Acad. Sci. U.S.A., 115, 2018
|
|
1UJS
 
 | |
2DJ7
 
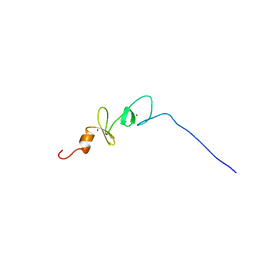 | | Solution Structure of 3rd LIM Domain of Actin-binding LIM Protein 3 | | Descriptor: | Actin-binding LIM protein 3, ZINC ION | | Authors: | Niraula, T.N, Sasagawa, A, Koshiba, S, Inoue, M, Kigawa, T, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2006-03-31 | | Release date: | 2006-10-01 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of 3rd LIM Domain of Actin-binding LIM Protein 3
To be Published
|
|
2KRH
 
 | |
2CT4
 
 | | Solution Strutcure of the SH3 domain of the Cdc42-interacting protein 4 | | Descriptor: | Cdc42-interacting protein 4 | | Authors: | Miyamoto, K, Tomizawa, T, Koshiba, S, Inoue, M, Kigawa, T, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2005-05-23 | | Release date: | 2005-11-23 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution Strutcure of the SH3 domain of the Cdc42-interacting protein 4
To be Published
|
|
1V6G
 
 | | Solution Structure of the LIM Domain of the Human Actin Binding LIM Protein 2 | | Descriptor: | Actin Binding LIM Protein 2, ZINC ION | | Authors: | Miyamoto, K, Tomizawa, T, Koshiba, S, Inoue, M, Kigawa, T, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2003-11-29 | | Release date: | 2004-06-01 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of the LIM Domain of the Human Actin Binding LIM Protein 2
To be Published
|
|
1WLX
 
 | |
3E35
 
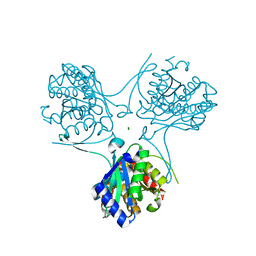 | | Actinobacteria-specific protein of unknown function, SCO1997 | | Descriptor: | MAGNESIUM ION, Uncharacterized protein SCO1997 | | Authors: | Gao, B, Gupta, R.S, Sugiman-Marangos, S, Junop, M.S. | | Deposit date: | 2008-08-06 | | Release date: | 2009-06-23 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structural and phylogenetic analysis of a conserved actinobacteria-specific protein (ASP1; SCO1997) from Streptomyces coelicolor.
Bmc Struct.Biol., 9, 2009
|
|
5DZR
 
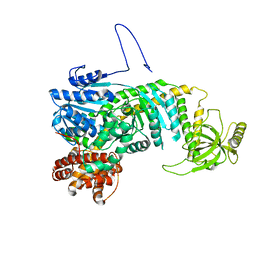 | |
8FIU
 
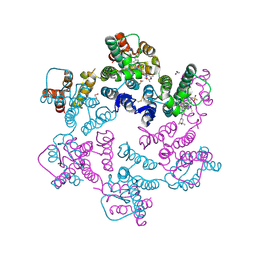 | | Potent long-acting inhibitors targeting HIV-1 capsid based on a versatile quinazolin-4-one scaffold | | Descriptor: | 1,2-ETHANEDIOL, HIV-1 capsid, N-[(1S)-1-{(3P)-3-{4-chloro-3-[(methanesulfonyl)amino]-1-methyl-1H-indazol-7-yl}-7-[(2R,6S)-2,6-dimethylmorpholin-4-yl]-4-oxo-3,4-dihydroquinazolin-2-yl}-2-(3,5-difluorophenyl)ethyl]-2-[(3bS,4aR)-3-(difluoromethyl)-5,5-difluoro-3b,4,4a,5-tetrahydro-1H-cyclopropa[3,4]cyclopenta[1,2-c]pyrazol-1-yl]acetamide | | Authors: | Nolte, R.T. | | Deposit date: | 2022-12-16 | | Release date: | 2023-02-15 | | Method: | X-RAY DIFFRACTION (1.56 Å) | | Cite: | Potent Long-Acting Inhibitors Targeting the HIV-1 Capsid Based on a Versatile Quinazolin-4-one Scaffold.
J.Med.Chem., 66, 2023
|
|
4RFX
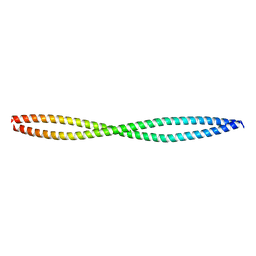 
 | |
4IK2
 
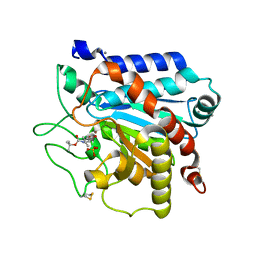 | | G215S, A251G, T257A, D260G, T262D mutant of carboxypeptidase T from Thermoactinomyces vulgaris with N-BOC-L-Leu | | Descriptor: | CALCIUM ION, Carboxypeptidase T, N-(tert-butoxycarbonyl)-L-leucine, ... | | Authors: | Timofeev, V.I, Kuznetsov, S.A, Akparov, V.K, Kuranova, I.P. | | Deposit date: | 2012-12-25 | | Release date: | 2013-12-25 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | G215S, A251G, T257A, D260G, T262D mutant of carboxypeptidase T from Thermoactinomyces vulgaris with N-BOC-L-Leu
To be published
|
|
4IHM
 
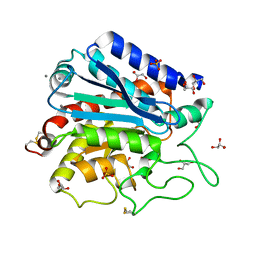 | | G215S, A251G, T257A, D260G, T262D mutant of carboxypeptidase T from Thermoactinomyces vulgaris | | Descriptor: | CALCIUM ION, Carboxypeptidase T, GLYCEROL, ... | | Authors: | Timofeev, V.I, Kuznetsov, S.A, Akparov, V.K, Kuranova, I.P. | | Deposit date: | 2012-12-19 | | Release date: | 2013-12-25 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.29 Å) | | Cite: | G215S, A251G, T257A, D260G, T262D mutant of carboxypeptidase T from Thermoactinomyces vulgaris
To be published
|
|
1SZZ
 
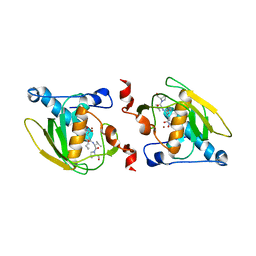 | | Crystal structure of peptide deformylase from Leptospira Interrogans complexed with inhibitor actinonin | | Descriptor: | ACTINONIN, Peptide deformylase, ZINC ION | | Authors: | Zhou, Z, Song, X, Li, Y, Gong, W. | | Deposit date: | 2004-04-06 | | Release date: | 2005-08-16 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | Novel conformational states of peptide deformylase from pathogenic bacterium Leptospira interrogans: implications for population shift
J.Biol.Chem., 280, 2005
|
|
4U10
 
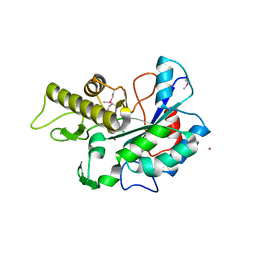 | |
1WGL
 
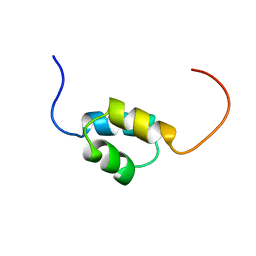 | | Solution Structure of CUE domain in the C-terminal of Human Toll-interacting Protein (Tollip) | | Descriptor: | Toll-interacting protein | | Authors: | Zhao, C, Koshiba, S, Inoue, M, Kigawa, T, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2004-05-28 | | Release date: | 2004-11-28 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of CUE domain in the C-terminal of Human Toll-interacting Protein (Tollip)
To be Published
|
|
7XMK
 
 | | Crystal structure of human RIPK1 kinase domain in complex with compound SKLB923 | | Descriptor: | 5-[2-(cyclopropylcarbonylamino)-[1,2,4]triazolo[1,5-a]pyridin-7-yl]-N-[(1S)-1-(3-fluorophenyl)ethyl]-1-methyl-indole-3-carboxamide, IODIDE ION, Receptor-interacting serine/threonine-protein kinase 1 | | Authors: | Zhang, L, Wang, Y, Li, Y, Yang, S. | | Deposit date: | 2022-04-26 | | Release date: | 2023-04-26 | | Last modified: | 2024-05-15 | | Method: | X-RAY DIFFRACTION (2.376 Å) | | Cite: | From Hit to Lead: Structure-Based Optimization of Novel Selective Inhibitors of Receptor-Interacting Protein Kinase 1 (RIPK1) for the Treatment of Inflammatory Diseases.
J.Med.Chem., 67, 2024
|
|
6JPD
 
 | | Mouse receptor-interacting protein kinase 3 (RIP3) amyloid structure by solid-state NMR | | Descriptor: | Receptor-interacting serine/threonine-protein kinase 3 | | Authors: | Wu, X.L, Hu, H, Zhang, J, Dong, X.Q, Wang, J, Schwieters, C, Wang, H.Y, Lu, J.X. | | Deposit date: | 2019-03-26 | | Release date: | 2020-10-28 | | Last modified: | 2024-05-15 | | Method: | SOLID-STATE NMR | | Cite: | The amyloid structure of mouse RIPK3 (receptor interacting protein kinase 3) in cell necroptosis.
Nat Commun, 12, 2021
|
|
