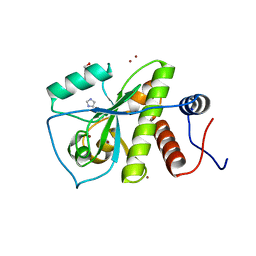
3LAT
 
 | |
6ELU
 | |
8Q7C
 
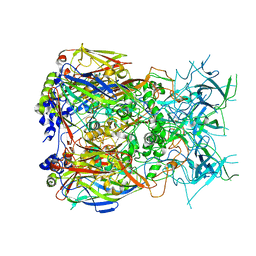 | | Cryo-EM structure of Adenovirus C5 hexon | | Descriptor: | Hexon protein | | Authors: | Zoll, S, Dhillon, A. | | Deposit date: | 2023-08-16 | | Release date: | 2023-08-30 | | Last modified: | 2024-03-27 | | Method: | ELECTRON MICROSCOPY (2.9 Å) | | Cite: | Structural insights into the interaction between adenovirus C5 hexon and human lactoferrin.
J.Virol., 98, 2024
|
|
4QO2
 
 | | Crystal structure of rhomboid intramembrane protease GlpG in complex with peptide derived inhibitor Ac-IATA-cmk | | Descriptor: | 6-AMINO-2-METHYL-1,7-DIHYDRO-8H-IMIDAZO[4,5-G]QUINAZOLIN-8-ONE, CHLORIDE ION, Rhomboid protease GlpG, ... | | Authors: | Zoll, S, Strisovsky, K. | | Deposit date: | 2014-06-19 | | Release date: | 2014-09-24 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Substrate binding and specificity of rhomboid intramembrane protease revealed by substrate-peptide complex structures.
Embo J., 33, 2014
|
|
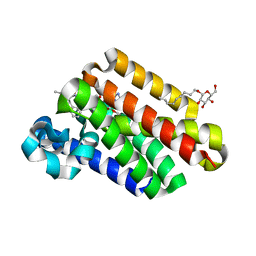
4QNZ
 
 | |
4QO0
 
 | |
4EPC
 
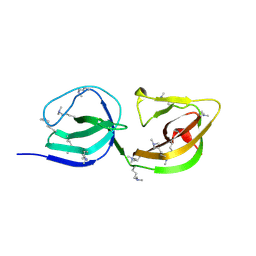 | |
7ZGJ
 
 | |
7ZGK
 
 | |
4KNK
 
 | | Crystal structure of Staphylococcus aureus hydrolase AmiA | | Descriptor: | 1,2-ETHANEDIOL, Bifunctional autolysin, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Buettner, F.M, Zoll, S, Stehle, T. | | Deposit date: | 2013-05-10 | | Release date: | 2014-03-12 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.124 Å) | | Cite: | Structure-function analysis of Staphylococcus aureus amidase reveals the determinants of peptidoglycan recognition and cleavage.
J.Biol.Chem., 289, 2014
|
|
