6CMK
 
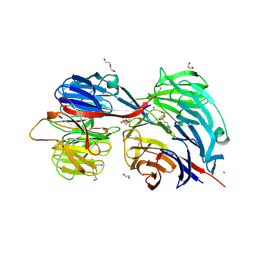 | | Crystal structure of Citrobacter koseri AztD | | Descriptor: | 1,2-ETHANEDIOL, 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, AztD protein, ... | | Authors: | Yukl, E.T. | | Deposit date: | 2018-03-05 | | Release date: | 2019-03-13 | | Last modified: | 2020-01-01 | | Method: | X-RAY DIFFRACTION (1.732 Å) | | Cite: | Crystal structures of AztD provide mechanistic insights into direct zinc transfer between proteins.
Commun Biol, 2, 2019
|
|
7RCJ
 
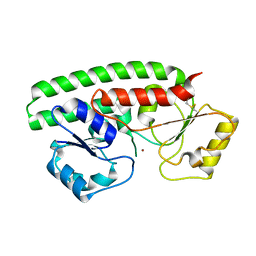 | | Crystal structure of ZnuA from Citrobacter koseri | | Descriptor: | 6-tungstotellurate(VI), High-affinity zinc uptake system protein ZnuA, ZINC ION | | Authors: | Yukl, E.T, Yekwa, E.L. | | Deposit date: | 2021-07-07 | | Release date: | 2022-05-18 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (3.15 Å) | | Cite: | Conformational flexibility in the zinc solute-binding protein ZnuA.
Acta Crystallogr.,Sect.F, 78, 2022
|
|
5KZJ
 
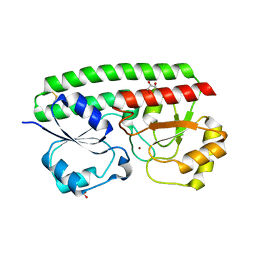 | | Loop Deletion mutant of Paracoccus denitrificans AztC | | Descriptor: | GLYCEROL, Periplasmic solute binding protein, ZINC ION | | Authors: | Yukl, E. | | Deposit date: | 2016-07-25 | | Release date: | 2017-07-05 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Mechanisms of zinc binding to the solute-binding protein AztC and transfer from the metallochaperone AztD.
J. Biol. Chem., 292, 2017
|
|
6XPN
 | | Z-loop Deletion Mutant of AztC from Paracoccus denitrificans | | Descriptor: | Periplasmic solute binding protein, ZINC ION | | Authors: | Yukl, E.T. | | Deposit date: | 2020-07-08 | | Release date: | 2020-08-26 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.26 Å) | | Cite: | Structural Features Mediating Zinc Binding and Transfer in the AztABCD Zinc Transporter System.
Biomolecules, 10, 2020
|
|
6N01
 
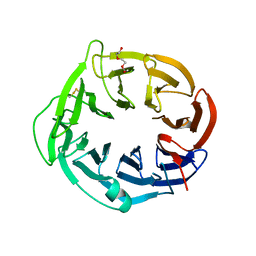 | | Structure of apo AztD from Citrobacter koseri | | Descriptor: | 1,2-ETHANEDIOL, 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, AztD Protein, ... | | Authors: | Yukl, E.T, Neupane, D.P. | | Deposit date: | 2018-11-06 | | Release date: | 2019-09-18 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.98 Å) | | Cite: | Crystal structures of AztD provide mechanistic insights into direct zinc transfer between proteins.
Commun Biol, 2, 2019
|
|
6BYW
 
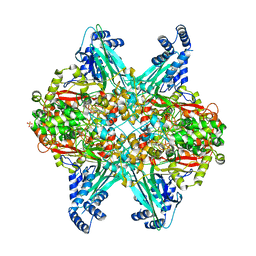 | | Structure of GoxA from Pseudoalteromonas luteoviolacea | | Descriptor: | 1,2-ETHANEDIOL, DI(HYDROXYETHYL)ETHER, GoxA, ... | | Authors: | Yukl, E.T, Avalos, D. | | Deposit date: | 2017-12-21 | | Release date: | 2018-02-14 | | Last modified: | 2018-02-28 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Structure and Enzymatic Properties of an Unusual Cysteine Tryptophylquinone-Dependent Glycine Oxidase from Pseudoalteromonas luteoviolacea.
Biochemistry, 57, 2018
|
|
6CK1
 
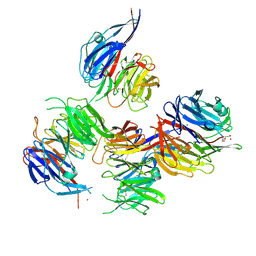 | | Crystal structure of Paracoccus denitrificans AztD | | Descriptor: | A1B2F4 protein, DI(HYDROXYETHYL)ETHER, ZINC ION | | Authors: | Yukl, E.T. | | Deposit date: | 2018-02-27 | | Release date: | 2019-03-06 | | Last modified: | 2020-01-01 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Crystal structures of AztD provide mechanistic insights into direct zinc transfer between proteins.
Commun Biol, 2, 2019
|
|
6EER
 
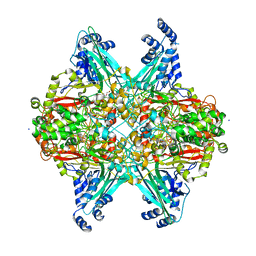 | | Structure of glycine-bound GoxA from Pseudoalteromonas luteoviolacea | | Descriptor: | 1,2-ETHANEDIOL, DI(HYDROXYETHYL)ETHER, GoxA, ... | | Authors: | Yukl, E.T, Avalos, D. | | Deposit date: | 2018-08-15 | | Release date: | 2019-01-16 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.82 Å) | | Cite: | Structural and Spectroscopic Characterization of a Product Schiff Base Intermediate in the Reaction of the Quinoprotein Glycine Oxidase, GoxA.
Biochemistry, 58, 2019
|
|
3SLE
 | |
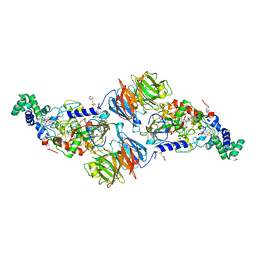
4O1Q
 
 | |
4L1Q
 
 | |
4K3I
 
 | | Crystal Structure of the Quinol Form of Methylamine Dehydrogenase in Complex with the Diferrous Form of MauG, C2 Space Group | | Descriptor: | 1,2-ETHANEDIOL, ACETATE ION, CALCIUM ION, ... | | Authors: | Yukl, E.Y, Wilmot, C.M. | | Deposit date: | 2013-04-10 | | Release date: | 2013-07-10 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structures of MauG in complex with quinol and quinone MADH.
Acta Crystallogr.,Sect.F, 69, 2013
|
|
3PXS
 
 | |
3PXT
 
 | | Crystal Structure of Ferrous CO Adduct of MauG in Complex with Pre-Methylamine Dehydrogenase | | Descriptor: | 1-(2-METHOXY-ETHOXY)-2-{2-[2-(2-METHOXY-ETHOXY]-ETHOXY}-ETHANE, ACETATE ION, CALCIUM ION, ... | | Authors: | Yukl, E.T, Goblirsch, B.R, Wilmot, C.M. | | Deposit date: | 2010-12-10 | | Release date: | 2011-03-23 | | Last modified: | 2018-01-24 | | Method: | X-RAY DIFFRACTION (2.16 Å) | | Cite: | Crystal Structures of CO and NO Adducts of MauG in Complex with Pre-Methylamine Dehydrogenase: Implications for the Mechanism of Dioxygen Activation.
Biochemistry, 50, 2011
|
|
3PXW
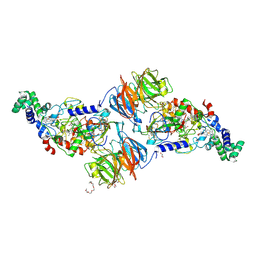 
 | | Crystal Structure of Ferrous NO Adduct of MauG in Complex with Pre-Methylamine Dehydrogenase | | Descriptor: | 1,2-ETHANEDIOL, 1-(2-METHOXY-ETHOXY)-2-{2-[2-(2-METHOXY-ETHOXY]-ETHOXY}-ETHANE, ACETATE ION, ... | | Authors: | Yukl, E.T, Goblirsch, B.R, Wilmot, C.M. | | Deposit date: | 2010-12-10 | | Release date: | 2011-03-23 | | Last modified: | 2018-01-24 | | Method: | X-RAY DIFFRACTION (2.11 Å) | | Cite: | Crystal Structures of CO and NO Adducts of MauG in Complex with Pre-Methylamine Dehydrogenase: Implications for the Mechanism of Dioxygen Activation.
Biochemistry, 50, 2011
|
|
3RLM
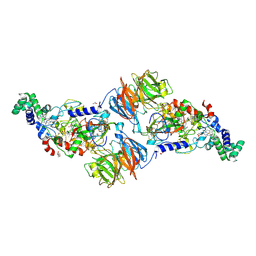 
 | |
4L3G
 
 | |
4L3H
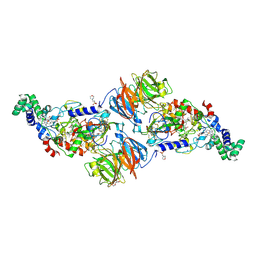 
 | |
6UBR
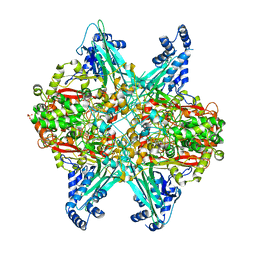 
 | | Crystal structure of D678A GoxA bound to glycine at pH 7.5 | | Descriptor: | DI(HYDROXYETHYL)ETHER, GLYCINE, MAGNESIUM ION, ... | | Authors: | Yukl, E.T, Avalos, D. | | Deposit date: | 2019-09-12 | | Release date: | 2019-10-23 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.96 Å) | | Cite: | Kinetic and structural evidence that Asp-678 plays multiple roles in catalysis by the quinoprotein glycine oxidase.
J.Biol.Chem., 294, 2019
|
|
6UBN
 | | Crystal structure of D678E GoxA bound to glycine | | Descriptor: | MAGNESIUM ION, Quinoprotein glycine oxidase, SODIUM ION | | Authors: | Yukl, E.T. | | Deposit date: | 2019-09-12 | | Release date: | 2019-10-23 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Kinetic and structural evidence that Asp-678 plays multiple roles in catalysis by the quinoprotein glycine oxidase.
J.Biol.Chem., 294, 2019
|
|
6UFQ
 
 | | Crystal structure of D678N GoxA bound to glycine | | Descriptor: | GLYCINE, Glycine Oxidase GoxA, MAGNESIUM ION | | Authors: | Yukl, E.T. | | Deposit date: | 2019-09-24 | | Release date: | 2019-10-23 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.51 Å) | | Cite: | Kinetic and structural evidence that Asp-678 plays multiple roles in catalysis by the quinoprotein glycine oxidase.
J.Biol.Chem., 294, 2019
|
|
6UBZ
 | | Crystal structure of D678A GoxA bound to glycine at pH 5.5 | | Descriptor: | GLYCINE, MAGNESIUM ION, Uncharacterized protein GoxA | | Authors: | Yukl, E.T. | | Deposit date: | 2019-09-13 | | Release date: | 2019-10-23 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.83 Å) | | Cite: | Kinetic and structural evidence that Asp-678 plays multiple roles in catalysis by the quinoprotein glycine oxidase.
J.Biol.Chem., 294, 2019
|
|
6UC1
 | | Crystal structure of D678A GoxA soaked in glycine at pH 7.5 | | Descriptor: | GLYCINE, MAGNESIUM ION, SULFATE ION, ... | | Authors: | Yukl, E.T. | | Deposit date: | 2019-09-13 | | Release date: | 2019-10-23 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.19 Å) | | Cite: | Kinetic and structural evidence that Asp-678 plays multiple roles in catalysis by the quinoprotein glycine oxidase.
J.Biol.Chem., 294, 2019
|
|
4FAV
 | |
4FA1
 
 | |
