8Q3L
 
 | |
8BPI
 
 | |
7A8Y
 
 | |
8BD0
 
 | |
4C3C
 
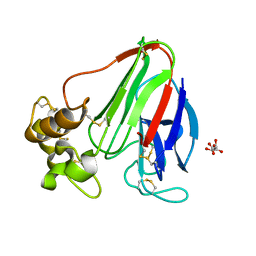 | | Thaumatin refined against hatrx data for time-point 1 | | Descriptor: | L(+)-TARTARIC ACID, THAUMATIN-1 | | Authors: | Yorke, B.A, Beddard, G.S, Owen, R.L, Pearson, A.R. | | Deposit date: | 2013-08-22 | | Release date: | 2014-09-03 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Time-Resolved Crystallography Using the Hadamard Transform.
Nat.Methods, 11, 2014
|
|
4AOK
 
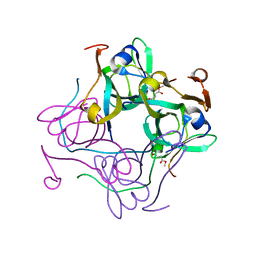 | |
4AON
 
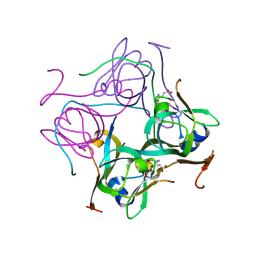 | |
4AZD
 
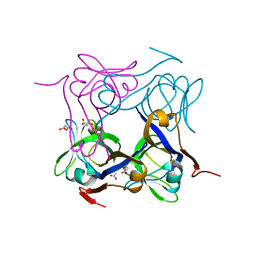 | | T57V mutant of aspartate decarboxylase | | Descriptor: | ASPARTATE 1-DECARBOXYLASE, MALONATE ION | | Authors: | Webb, M.E, Yorke, B.A, Kershaw, T, Lovelock, S, Lobley, C.M.C, Kilkenny, M.L, Smith, A.G, Blundell, T.L, Pearson, A.R, Abell, C. | | Deposit date: | 2012-06-25 | | Release date: | 2012-07-25 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.62 Å) | | Cite: | Threonine 57 is Required for the Post-Translational Activation of Escherichia Coli Aspartate Alpha-Decarboxylase
Acta Crystallogr.,Sect.D, 70, 2014
|
|
