6A0S
 
 | | Homoserine dehydrogenase from Thermus thermophilus HB8 complexed with HSE and NADPH | | Descriptor: | 3-CYCLOHEXYL-1-PROPYLSULFONIC ACID, FORMIC ACID, GLYCEROL, ... | | Authors: | Akai, S, Ikushiro, H, Sawai, T, Yano, T, Kamiya, N, Miyahara, I. | | Deposit date: | 2018-06-06 | | Release date: | 2018-11-28 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The crystal structure of homoserine dehydrogenase complexed with l-homoserine and NADPH in a closed form
J. Biochem., 165, 2019
|
|
6A0U
 
 | | Homoserine dehydrogenase K195A mutant from Thermus thermophilus HB8 complexed with HSE and NADP+ | | Descriptor: | 3-CYCLOHEXYL-1-PROPYLSULFONIC ACID, FORMIC ACID, GLYCEROL, ... | | Authors: | Akai, S, Ikushiro, H, Sawai, T, Yano, T, Kamiya, N, Miyahara, I. | | Deposit date: | 2018-06-06 | | Release date: | 2018-11-28 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.93 Å) | | Cite: | The crystal structure of homoserine dehydrogenase complexed with l-homoserine and NADPH in a closed form
J. Biochem., 165, 2019
|
|
6A0T
 
 | | Homoserine dehydrogenase K99A mutant from Thermus thermophilus HB8 complexed with HSE and NADP+ | | Descriptor: | 3-CYCLOHEXYL-1-PROPYLSULFONIC ACID, FORMIC ACID, GLYCEROL, ... | | Authors: | Akai, S, Ikushiro, H, Sawai, T, Yano, T, Kamiya, N, Miyahara, I. | | Deposit date: | 2018-06-06 | | Release date: | 2018-11-28 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.87 Å) | | Cite: | The crystal structure of homoserine dehydrogenase complexed with l-homoserine and NADPH in a closed form
J. Biochem., 165, 2019
|
|
5X9Y
 
 | |
5Z41
 
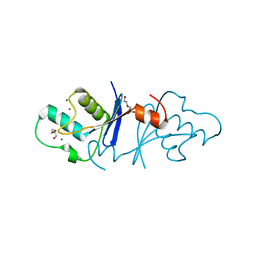 | | Aquifex aeolicus MutL endonuclease domain with a single zinc ion. | | Descriptor: | DI(HYDROXYETHYL)ETHER, DNA mismatch repair protein MutL, MAGNESIUM ION, ... | | Authors: | Fukui, K, Yano, T. | | Deposit date: | 2018-01-10 | | Release date: | 2018-04-25 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Multiple zinc ions maintain the open conformation of the catalytic site in the DNA mismatch repair endonuclease MutL from Aquifex aeolicus
FEBS Lett., 592, 2018
|
|
5Z42
 
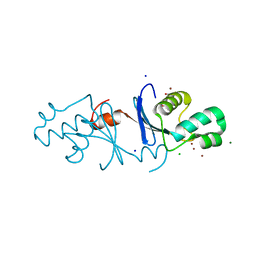 | | Aquifex aeolicus MutL endonuclease domain with three zinc ions. | | Descriptor: | CHLORIDE ION, DI(HYDROXYETHYL)ETHER, DNA mismatch repair protein MutL, ... | | Authors: | Fukui, K, Yano, T. | | Deposit date: | 2018-01-10 | | Release date: | 2018-04-25 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Multiple zinc ions maintain the open conformation of the catalytic site in the DNA mismatch repair endonuclease MutL from Aquifex aeolicus
FEBS Lett., 592, 2018
|
|
5YTQ
 
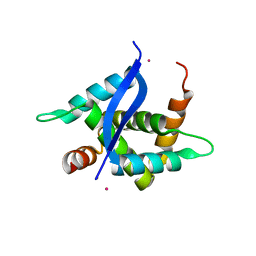 | | Crystal Structure of TTHA0139 L34A with Lanthanum from Thermus thermophilus HB8 | | Descriptor: | LANTHANUM (III) ION, TTHA0139 | | Authors: | Takao, K, Inoue, M, Fukui, K, Yano, T, Masui, R. | | Deposit date: | 2017-11-19 | | Release date: | 2018-11-21 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (2.393 Å) | | Cite: | Crystal Structure of TTHA0139 L34A with Lanthanum from Thermus thermophilus HB8
To Be Published
|
|
5YTP
 
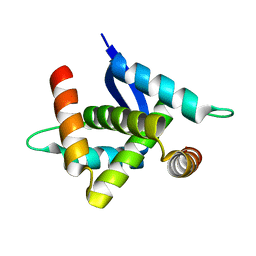 | |
1K5O
 
 | | CPI-17(35-120) deletion mutant | | Descriptor: | CPI-17 | | Authors: | Ohki, S, Eto, M, Kariya, E, Hayano, T, Hayashi, Y, Yazawa, M, Brautigan, D, Kainosho, M. | | Deposit date: | 2001-10-11 | | Release date: | 2002-10-11 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution NMR Structure of the Myosin Phosphatase Inhibitor Protein CPI-17 Shows Phosphorylation-induced Conformational Changes Responsible for Activation
J.Mol.Biol., 314, 2001
|
|
