5VXG
 
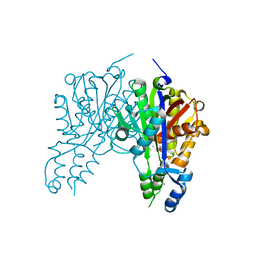 | | Crystal structure of Xanthomonas campestris OleA E117Q bound with Cerulenin | | Descriptor: | (2S, 3R)-3-HYDROXY-4-OXO-7,10-TRANS,TRANS-DODECADIENAMIDE, 3-oxoacyl-[ACP] synthase III, ... | | Authors: | Jensen, M.R, Wilmot, C.M. | | Deposit date: | 2017-05-23 | | Release date: | 2017-10-25 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.07 Å) | | Cite: | OleA Glu117 is key to condensation of two fatty-acyl coenzyme A substrates in long-chain olefin biosynthesis.
Biochem. J., 474, 2017
|
|
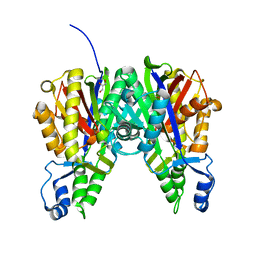
5VXD
 
 | |
5VXI
 
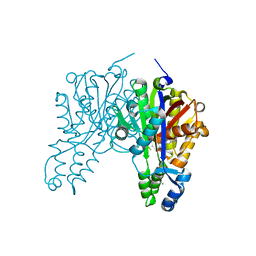 | | Crystal structure of Xanthomonas campestris OleA E117D bound with Cerulenin | | Descriptor: | (2S, 3R)-3-HYDROXY-4-OXO-7,10-TRANS,TRANS-DODECADIENAMIDE, 3-oxoacyl-[ACP] synthase III, ... | | Authors: | Jensen, M.R, Wilmot, C.M. | | Deposit date: | 2017-05-23 | | Release date: | 2017-10-25 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.08 Å) | | Cite: | OleA Glu117 is key to condensation of two fatty-acyl coenzyme A substrates in long-chain olefin biosynthesis.
Biochem. J., 474, 2017
|
|
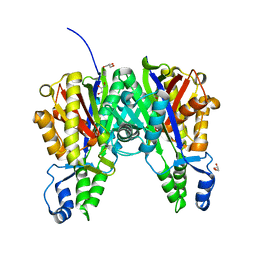
5VXF
 
 | |
5VXH
 
 | | Crystal structure of Xanthomonas campestris OleA E117D | | Descriptor: | 3-oxoacyl-[ACP] synthase III, GLYCEROL, PHOSPHATE ION | | Authors: | Jensen, M.R, Wilmot, C.M. | | Deposit date: | 2017-05-23 | | Release date: | 2017-10-25 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.84 Å) | | Cite: | OleA Glu117 is key to condensation of two fatty-acyl coenzyme A substrates in long-chain olefin biosynthesis.
Biochem. J., 474, 2017
|
|
5VRD
 
 | |
4FB1
 | |
4FAV
 | |
4FAN
 
 | |
4FA1
 
 | |
4FA9
 
 | |
4FA5
 
 | |
4FA4
 | |
4EV2
 
 | |
4EV5
 | |
4FAS
 | |
4KFD
 
 | | Crystal structure of Hansenula polymorpha copper amine oxidase-1 reduced by methylamine at pH 6.0 | | Descriptor: | COPPER (II) ION, GLYCEROL, HYDROGEN PEROXIDE, ... | | Authors: | Johnson, B.J, Yukl, E.T, Klema, V.J, Wilmot, C.M. | | Deposit date: | 2013-04-26 | | Release date: | 2013-08-21 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.69 Å) | | Cite: | Structural evidence for the semiquinone in a copper amine oxidase from Hansenula polymorpha: implications for the catalytic mechanism
J.Biol.Chem., 2013
|
|
4KFE
 | | Crystal structure of Hansenula polymorpha copper amine oxidase-1 reduced by methylamine at pH 7.0 | | Descriptor: | COPPER (II) ION, FORMYL GROUP, GLYCEROL, ... | | Authors: | Johnson, B.J, Yukl, E.T, Klema, V.J, Wilmot, C.M. | | Deposit date: | 2013-04-26 | | Release date: | 2013-08-21 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structural evidence for the semiquinone in a copper amine oxidase from Hansenula polymorpha: implications for the catalytic mechanism
J.Biol.Chem., 2013
|
|
4K3I
 
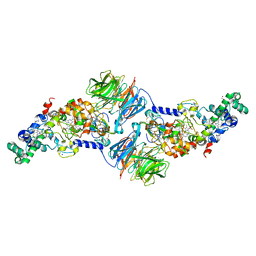 | | Crystal Structure of the Quinol Form of Methylamine Dehydrogenase in Complex with the Diferrous Form of MauG, C2 Space Group | | Descriptor: | 1,2-ETHANEDIOL, ACETATE ION, CALCIUM ION, ... | | Authors: | Yukl, E.Y, Wilmot, C.M. | | Deposit date: | 2013-04-10 | | Release date: | 2013-07-10 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structures of MauG in complex with quinol and quinone MADH.
Acta Crystallogr.,Sect.F, 69, 2013
|
|
4JO0
 
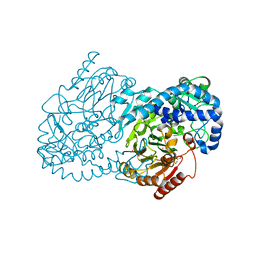 | | Crystal Structure of CmlA, a diiron beta-hydroxylase from Streptomyces venezuelae | | Descriptor: | ACETATE ION, CmlA, FE (III) ION, ... | | Authors: | Knoot, C.J, Makris, T.M, Wilmot, C.M, Lipscomb, J.D. | | Deposit date: | 2013-03-16 | | Release date: | 2013-09-11 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.17 Å) | | Cite: | Structure of a Dinuclear Iron Cluster-Containing beta-Hydroxylase Active in Antibiotic Biosynthesis.
Biochemistry, 52, 2013
|
|
4L3G
 
 | |
4L3H
 | |
4KFF
 
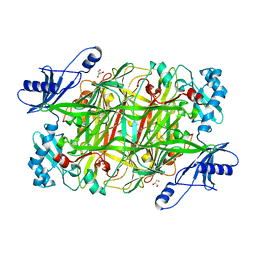 | | Crystal structure of Hansenula polymorpha copper amine oxidase-1 reduced by methylamine at pH 8.5 | | Descriptor: | COPPER (II) ION, GLYCEROL, Peroxisomal primary amine oxidase | | Authors: | Johnson, B.J, Yukl, E.T, Klema, V.J, Wilmot, C.M. | | Deposit date: | 2013-04-26 | | Release date: | 2013-08-28 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Structural evidence for the semiquinone in a copper amine oxidase from Hansenula polymorpha: implications for the catalytic mechanism
J.Biol.Chem., 2013
|
|
4L1Q
 
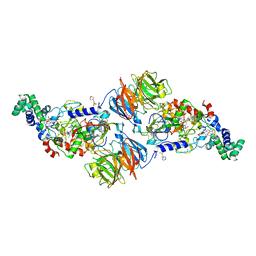 | |
3Q09
 
 | |
